External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G18550 201 / 1e-65
HD
HB-2, ATHB21
homeobox-2, homeobox protein 21 (.1)
AT4G36740 200 / 3e-65
HD
HB-5, ATHB40
homeobox protein 40 (.1)
AT5G66700 142 / 3e-42
HD
ATHB53, HB53, HB-8
HOMEOBOX-8, ARABIDOPSIS THALIANA HOMEOBOX 53, homeobox 53 (.1)
AT3G01470 83 / 4e-19
HD
HD-ZIP-1, HAT5, ATHB1, ATHB-1
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
AT4G40060 77 / 6e-17
HD
ATHB16 ,ATHB-16
homeobox protein 16 (.1)
AT2G22430 77 / 7e-17
HD
ATHB6
homeobox protein 6 (.1)
AT5G65310 77 / 1e-16
HD
ATHB-5, ATHB5
homeobox protein 5 (.1.2)
AT1G69780 74 / 1e-15
HD
ATHB13
Homeobox-leucine zipper protein family (.1)
AT5G03790 72 / 3e-15
HD
LMI1, ATHB51
LATE MERISTEM IDENTITY1, homeobox 51 (.1)
AT5G15150 70 / 3e-14
HD
ATHB3, HAT7, ATHB-3
HOMEOBOX FROM ARABIDOPSIS THALIANA 7, homeobox 3 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041719
156 / 1e-47
AT4G36740 192 / 9e-62
homeobox protein 40 (.1)
Lus10024031
149 / 6e-45
AT4G36740 189 / 2e-60
homeobox protein 40 (.1)
Lus10037218
82 / 2e-18
AT3G01470 140 / 7e-40
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10042180
82 / 2e-18
AT3G01470 159 / 6e-47
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10022225
81 / 3e-18
AT3G01470 270 / 2e-90
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10035583
79 / 9e-18
AT5G03790 146 / 2e-43
LATE MERISTEM IDENTITY1, homeobox 51 (.1)
Lus10036674
80 / 1e-17
AT3G01470 153 / 8e-45
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10030800
79 / 1e-17
AT3G01470 135 / 4e-38
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10008795
79 / 2e-17
AT3G01470 261 / 3e-87
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10033102
78 / 3e-17
AT3G01470 151 / 4e-44
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00046
Homeodomain
Homeodomain
Representative CDS sequence
>Potri.005G126100.1 pacid=42804851 polypeptide=Potri.005G126100.1.p locus=Potri.005G126100 ID=Potri.005G126100.1.v4.1 annot-version=v4.1
ATGAATCACAAGGTTGAAGATCACATAGCACTCATATCTCAATTGTACCCCGGTTTATACACTCAAATGGTTCCAAAACAAGGTGAATCTAAACCACGAC
GAAGGCGAAAGAAGAACACAGGAGAAGGTGTTAGTGGCGCAAGGAAGAGGAAACTTAGTGCTGAGCAAGTTAATTTCCTAGAAATGAATTTTGGTGATGA
ACATAAGTTGGAAACCGAGAGGAAGGACAAGCTTGCTTCAGATTTGGGACTTGATCCTCGTCAAGTTGCGGTTTGGTTTCAAAACCGAAGGGCAAGGTGG
AAGAACAAGAAGCTAGAGGAGGAATACACAAAGTTGAAGACAGCCCATGAGAGCATAGTTGTCCAGAAATGCCAGCTTGAATCCGAGGTTTTAAAGCTCA
AGGAACAACTCTCTAGAACAGAGAAGGAGATCCAAAGGCTATCAGACCGTGCTGATAGGGTTTCCACCAACAGTCCAAGTTCATCCCTGTCAATGGCCAT
TGAGCCCCCATTTCTGGGAGAATTTGCAGTACTGGAAGGATATGGAGATGCTTTCTACATGCCACCAGAAAACAATTACATTCCCGGAATGGAATGGATT
AGTCAGTATATGTGA
AA sequence
>Potri.005G126100.1 pacid=42804851 polypeptide=Potri.005G126100.1.p locus=Potri.005G126100 ID=Potri.005G126100.1.v4.1 annot-version=v4.1
MNHKVEDHIALISQLYPGLYTQMVPKQGESKPRRRRKKNTGEGVSGARKRKLSAEQVNFLEMNFGDEHKLETERKDKLASDLGLDPRQVAVWFQNRRARW
KNKKLEEEYTKLKTAHESIVVQKCQLESEVLKLKEQLSRTEKEIQRLSDRADRVSTNSPSSSLSMAIEPPFLGEFAVLEGYGDAFYMPPENNYIPGMEWI
SQYM
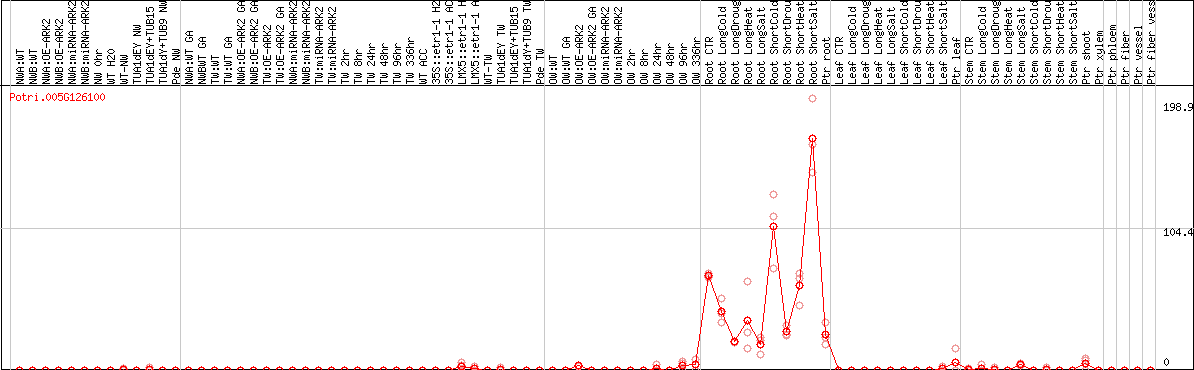
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G126100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.