Potri.005G129800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G129800.2 pacid=42802484 polypeptide=Potri.005G129800.2.p locus=Potri.005G129800 ID=Potri.005G129800.2.v4.1 annot-version=v4.1
ATGGATGATGGCAGAGTGAATGACTTATTGCTAAATGTAGAAGAGTCTCGTATAGAAAGACAATGTGAGGGACTAGGGACAGTAGACAAATTGCATATTT
CTGAAGGTGGAACTTCATATTCTGATTGTAAGGTAGAGAGCCAAAGGTTATCCTGTGATTCTCAGGATTTTGGAGAAGATGACATAAACGTACAGAATTA
TTACACAGAACCCAATGCAGCCTCTGAAAACTCTAATTTAATTGTTGACACCATTGAAAGCGAACCAAACAGCTGTAGGTATGGAGAGCCATCTCTTTTG
GAGCCTAACTGGCTAGAACACGATGAATCTGTGGCACTATGGGTCAAGTGGAGAGGCAAGTGGCAGGCTGGAATCAGATGTGCACGGGCAGACTGGCCAT
TGTCAACTTTGAGAGCCAAACCAACTCATGATCGAAAGCAGTATTTTGTGATATTTTTTCCTCACACCAGAAATTATTCCTGGGCAGATATGCTGCTTGT
CCAACCAATCAATGGATTTCCAGAGCCTATTGCATATAAGACTCACAAAATTGGGCTAAAAATGGTTAAAGATATGAGTGTTGCACGGCGGTTTATAATG
AAAAAGCTGGCTGTTGCCATGGTGAATATTGTCGATCAGTTTCATTCTGAGGCTTTAGTAGACCCAGCTCGTGATGTGATGGTCTGGAAGGAATTTGCCA
TGGAGGCTTCACGATGTAGTGCTTATTCTGATCTTGGAAGAATGCTTCTGAAACTTCAAAATATGATTTTGCAACAGTACATAAGTTCTGATTGGCTGCA
GAATTCTTTTCAATCCTGGGTACAGCAGTGTCAAGTTGCATGTAGTGCTGAATCCATTGAATTGCTTAGGGAGGAATTGTACAATTCTATTCTATGGAAT
GAAGTCGATTCTCTCCATGATGCACCAGTGCAATCGACATTGGGATCCGAGTGGAAAACCTGGAAGCATGAAGCTATGAAGTGGTTCTCAACCTCCCAAC
CTGTGACAAGTGGTGGAGATATGGAGCAGCAGAACTGTGACAATCTCTCCCCCTCAACCATCAGTCTTCAAGCTACCAGAAAGAGGCCAAAGCTTGAAGT
TCGCCGTGCAGAGACACATGCTTCCCAGGTTGAGTCCAGCAGTCCACTTCAAACTACGAATGTCGAAATTGACTCTGAGTTTTTCAGTAATCGAGACACT
GTCAATGCTCATACTTTAGAATCAGAGCTTTCTAAGGAGGATGGTTTTGGCGAGGTAGCTGCACCTTTAGAGTCACCTTGTAGCATGGCTGATAGATGGG
ATGGGATTGTAGTTGAAGCAGGAAATCCTGAGCTTGTACAGAACAAAGGTGTAGAAATGACACCTGTTAATGAAGTTTTGGCTAAGGAAAGCATAGAACC
TGGGAGTAAGAATCGCCAGTGTACAGCTTTTATTGAATCTAAGGGAAGGCAGTGTGTGAGGTGGGCTAATGATGGTGATGTTTATTGTTGTGTGCATCTG
GCCTCTCGTTTTGCTGGCAGCTCCACTAGAGGAGAGGCAAGTCCTGTTCATAGTCCTATGTGTGAAGGTACCACTGTTCTTGGTACTCGATGCAAGCATC
GGTCTCTACCAGGTACCACATTCTGCAAGAAACACAGACCCTGGCCTGATGCAGAGAAGACCTCAAATTTGCCAGAGAATCCCCTTAAGAGAAAACATGA
AGAGATCTTTCCCAGTTCAGACACCACATATTGCAAAGAGATGGTATTGTCTGGACAAGTTGAAAATCCCCTTCGAGTGCAACCTGTCTCAGCCATGGAT
GGTGATGCCTTTCATGGAAGGAAAAGCTTACCCGAGAAACTTGAGCATCCTGGCCATGATTGCAATAGCTCCAAGATGCTGCACTGTATTGGCTCAAGCT
CCCTTGATAGTAGTATCCTGTGTCCTGAGAGTCCAAAGCGCTATTCGTTGTACTGTGACAAACACATTCCTAGTTGGCTCAAGCGTGCTAGAAATGGTAG
AAGTAGGATAATATCAAAAGAAGTGTTTATAGATCTTTTGAAGGATTGTCGCTCACCGCAGCAGAAATTGCATTTACATCAAGCATGTGAGCTTTTCTAC
AAGCTTTTCAAAAGCATTTTCTCTCTAAGAAACCCAGTTCCTATGGAGGTTCAACTTCAGTGGGCTCTCTCTGAAGCTTCTAAAGATTTTAATGTTGGGG
AACTCTTGTTGAAGTTAGTTTTCACTGAAAAGGAGAGACTCAAAAAGTTATGGGGCTTTGCTGTGGAGGAAGATTTACAAGTTTCCTCTGTCATGGAAGA
ACCAGCAATTTTTCCATTGGCTGTCAACTGTAGCCATGATGATGAAAATTCTATTAGGTGCAAAATATGCTCAAAGGAGTTTCTCGATGATAAAGAGCTT
GGAAACCACTGGATGGACAACCATAAAAAGGAAGCACAGTGGCATTTCAGAGGTCATGCTTGTGCCATATGCCTGGATTCTTTTACTGACAGAAAAAGTT
TGGAAACCCATGTGCAGGAGAGACATCATGTGGAATTTGTTGAACAGTGCATGCTGTTCCAGTGCATTCCTTGTGCCAGCCACTTTGGGAACACTGACCA
GTTATGGTTGCATGTCCTCTCTGTTCACCCTGCTGATTTCAGACTGCCAAAAGGTGCTCAACAACTTAATCCGTCTATGGGTGAAGAGAAGGAGGATTCC
TTGCAAAAACTTGAGCTGCAAAATGCAGCATCAATGGAAAATCACACTGAGAATTTAGGTGGTGTAAGAAAGTACATTTGCAAGTTCTGTGGCCTGAAGT
TTGATTTACTACCTGATCTTGGTCGCCACCATCAAGCTGCTCATATGGGACCAAATCTATTCAGCTCTCGCCCCCCAAAGAGAGGTGTTCGATATTATGC
TTATAGATTAAAATCTGGAAGACTTAGCCGCCCGAAATTTAAGAAAGGTCTGGGGGCAGCAACATATAGCAGCATCAGGAACAGGATGACTTCTGGTCTG
AAGAAACGTATCCAGGCTTCAAAATCACTTAGCAGCCAGGGACTAAGCATACAGTCTAATTTGACTGAGGCAGGGGCTTTAGGTAGATTAGCAGAATCTC
AATGCTCAGCGGTTGCAAAGATATTGTTTTCTGAGGTTCAGAAAACAAAACCTCGGCCTAATAACCTTGATATCTTAGCTATTGCACGCTCTGCTTGCTG
CAAGGTGAGCCTAAAAGCATCACTGGAGGGGAAATATGGAGTGTTGCCAGAACGCTTTTATTTGAAGGCAGCCAAGCTCTGCAGTGAGCATAATATTCAA
GTACAATGGCACCAGGAGGAGTTTAGTTGTTCCAGAGGATGTAAGAGTTTCAAGGATCCAGGATTATTCTCGCCCTTGATGGCTCTTCCAAATGGTTTTA
AGGGGAAACAAATGATACATTCATCTGATCATACAAACAGTGAGTGTGAAGTGGATGAGTGCCACTATATTATAGATGTGCATGATGTTACAGAGGGGCC
TAAGCAAAAGGCTACTGTCTTGTGTACTGACATAAGCTTTGGGAAGGAGACAATACCAGTGGCATGTGTTGTAGATGAAGATCTCCTGGATTCCCTCCAT
GTCTTGGCAGATGGTTACGATGGTCAAATTAGTAAATTCCCCAAGCCCTGGGATACTTTTACCTATGTAACAGGGCCAGTTCATGATCAATGTGATAGTC
TGGACATTGAGGGTTTGCAATTGAGGTGTTCTTGCCAATATTCAATGTGCTGTCCTGAAACATGTGATCATGTTTATCTCTTTGATAACGACTATGAAGA
TGCAAAGGACATATATGGGAAATCCATGCTTGGCAGATTTCCGTATGACTATAAAGGGCGGCTTGTTCTGGAGGAAGGTTACCTTGTTTACGAGTGCAAT
AGCATGTGCAACTGCAATAAAACATGCCCAAATAGAGTTTTGCAGAATGGAATACGAGTGAAACTGGAAGTCTTCAATACAGATAATAAAGGTTGGGCTG
TCAGGGCAGGTGAACCAATCCTACGTGGTACATTTATATGTGAGTACACTGGAGAGATTTTAAATGAACAGGAGGCAAGCAATAGGCGTGACAGGTATGG
CAAAGAAGTTTGCAGCTATATGTATAAAATTGATGCTCATACCAATGATATGAGCAGAATGGTTGAAGGACAGGCCCATTATTTTATTGATGCTACAAAG
TATGGGAATGTTTCACGGTTCATCAATCACAGCTGCATGCCAAATCTTGTGAACCACCAAGTTCTGGTTGACAGCATGGATTCTCAGCGTGCTCATATCG
GTCTTTATGCAAGTCAGGATATAGCTTTTGGTGAAGAACTGACATACAACTATCGGTATGAGCTGCTGCCTGGAGAAGGATATCCTTGCCATTGTGGAGC
TTCAAAGTGCCGGGGCCGCCTGTATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G129800.2 pacid=42802484 polypeptide=Potri.005G129800.2.p locus=Potri.005G129800 ID=Potri.005G129800.2.v4.1 annot-version=v4.1
MDDGRVNDLLLNVEESRIERQCEGLGTVDKLHISEGGTSYSDCKVESQRLSCDSQDFGEDDINVQNYYTEPNAASENSNLIVDTIESEPNSCRYGEPSLL
EPNWLEHDESVALWVKWRGKWQAGIRCARADWPLSTLRAKPTHDRKQYFVIFFPHTRNYSWADMLLVQPINGFPEPIAYKTHKIGLKMVKDMSVARRFIM
KKLAVAMVNIVDQFHSEALVDPARDVMVWKEFAMEASRCSAYSDLGRMLLKLQNMILQQYISSDWLQNSFQSWVQQCQVACSAESIELLREELYNSILWN
EVDSLHDAPVQSTLGSEWKTWKHEAMKWFSTSQPVTSGGDMEQQNCDNLSPSTISLQATRKRPKLEVRRAETHASQVESSSPLQTTNVEIDSEFFSNRDT
VNAHTLESELSKEDGFGEVAAPLESPCSMADRWDGIVVEAGNPELVQNKGVEMTPVNEVLAKESIEPGSKNRQCTAFIESKGRQCVRWANDGDVYCCVHL
ASRFAGSSTRGEASPVHSPMCEGTTVLGTRCKHRSLPGTTFCKKHRPWPDAEKTSNLPENPLKRKHEEIFPSSDTTYCKEMVLSGQVENPLRVQPVSAMD
GDAFHGRKSLPEKLEHPGHDCNSSKMLHCIGSSSLDSSILCPESPKRYSLYCDKHIPSWLKRARNGRSRIISKEVFIDLLKDCRSPQQKLHLHQACELFY
KLFKSIFSLRNPVPMEVQLQWALSEASKDFNVGELLLKLVFTEKERLKKLWGFAVEEDLQVSSVMEEPAIFPLAVNCSHDDENSIRCKICSKEFLDDKEL
GNHWMDNHKKEAQWHFRGHACAICLDSFTDRKSLETHVQERHHVEFVEQCMLFQCIPCASHFGNTDQLWLHVLSVHPADFRLPKGAQQLNPSMGEEKEDS
LQKLELQNAASMENHTENLGGVRKYICKFCGLKFDLLPDLGRHHQAAHMGPNLFSSRPPKRGVRYYAYRLKSGRLSRPKFKKGLGAATYSSIRNRMTSGL
KKRIQASKSLSSQGLSIQSNLTEAGALGRLAESQCSAVAKILFSEVQKTKPRPNNLDILAIARSACCKVSLKASLEGKYGVLPERFYLKAAKLCSEHNIQ
VQWHQEEFSCSRGCKSFKDPGLFSPLMALPNGFKGKQMIHSSDHTNSECEVDECHYIIDVHDVTEGPKQKATVLCTDISFGKETIPVACVVDEDLLDSLH
VLADGYDGQISKFPKPWDTFTYVTGPVHDQCDSLDIEGLQLRCSCQYSMCCPETCDHVYLFDNDYEDAKDIYGKSMLGRFPYDYKGRLVLEEGYLVYECN
SMCNCNKTCPNRVLQNGIRVKLEVFNTDNKGWAVRAGEPILRGTFICEYTGEILNEQEASNRRDRYGKEVCSYMYKIDAHTNDMSRMVEGQAHYFIDATK
YGNVSRFINHSCMPNLVNHQVLVDSMDSQRAHIGLYASQDIAFGEELTYNYRYELLPGEGYPCHCGASKCRGRLY
|
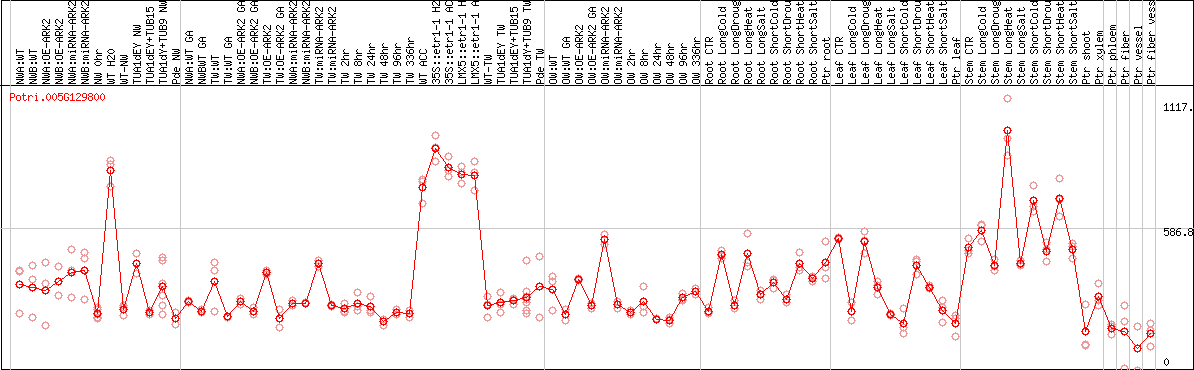
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G129800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
