Potri.005G170066 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G170066.2 pacid=42803226 polypeptide=Potri.005G170066.2.p locus=Potri.005G170066 ID=Potri.005G170066.2.v4.1 annot-version=v4.1
ATGTCGAGGATTACTAGGTGGAAGAATGAGAAAACTAAAGTGAAAGTGGTCTTTCGGCTGCAGTTCCATGCCACACATATTCCTCATCCAGGATGGGATA
AACTTTTTATATCTTTCATTCCTGCTGATTCTGGAAAAGCAACTGCAAAAACAACTAAAGCCAATGTAAGAAACGGGACCTGCAAATGGGCAGATCCTAT
CTACGAGACGACAAGGCTTCTTCAAGACGTTAAAACCAAGCGATATGATGAAAAGCTCTATAAGCTTGTCATTTCGATGGGTTCATCACGGTCTAGTGTC
CTTGGTGAGGCTACCATCAACCTTGCTGATTATGCTGATGCGTTAAAACCTTCTGTTGTTGCCCTGCCTCTTCATGGATCTGATTCTGGAACTGCTTTGC
ATGTCACTGTACAACTGCTCACTTCAAAAACTGGTTTCCGAGAGTTTGAGCAGCAGAGGGAACTCAGAGAGAGGGGGTTGCAGACTGACCAAAATAGCCC
TGATGAATCCTCTAGTGGAAAAGTATCATCTTCTGAAGGGATTATCAATGATCAGATAGATAAGGTAAATATCAGAGTTAGATTTAAAGAAAAATCTAAA
GACCTTGCTTCACTCGAAGAAGAAGTGGGGCCAAATGAAGAATATGCAGACTCAGCTGTTGGATTTGATGGCTCTTCCAATACATCAGAGAGTTTATACG
CAGAGAAGCATGATACTTCCAGCACACATGAAATTGATAGCCTTAAGAGTACCGTCTCTGGTGACTTAGCTGGACTTTCCCTCAGCCAAAGTCCTCAGCT
GGAAAAAGGGGATCCTTCTGATCATCGATTTTTGGCACAGGGGACGAATGATTGGGTTCATGCCTGGAGTTCAGACTATTCTGCAGATAATGACTTGGCA
GCTGCTTATGAAGAGAATGGTAGGCTTAGAGGAAGCTTGGAAGTGGCTGAGTCATCTGTTCTTGAGCTTAAACAGGAGGTAAGCTCCTTACAGGGTCATG
CTGATGAAATAGGTTATGAAGCCCAAACATTTGCTAAGCAACTTGCCTCTGAGATTGCTTCTGGAGAAGAGATGGTGAAAGAAGTGTCTGTACTGAAGTC
GGAGTGTTCAAAGTTGAAAGCTGATCTTGAACAGTTAAAGGTTTCTCAATTATGCCCTCCATTTAGCAGCAGAAATGCTGCAGAACCCCTACAGGATCAC
AGATTTCAAGAATTGAAGCTCAGATGGATAAAGGGGCTTTTATCCATGGAAGATAAGATAAAAGAGCTTCAAAATAATGCATGCTTGGGATACCATGAAA
GTGACTTCAGATTCCTTCACTCAGATGTAGAGGAGTTGATTGGTGTGCTGCAGGATCTTAAGCAAGGAACAGGGCTGCCAATTTCCAGTAACCACTTGGT
ACCTTGTGAAGGATCTAGCCTGAAGGAGATCAGAGAAATGAGTTTACATAAAAATAGCCAGTTTGTAAGTGAATCTGGCTTTGATGTGGACTTATATCAA
CCAGAACTAGGCATGCTTCATTGTCTTAACATTCCTGGCTTGGTTTCTCATGAGCCCGACTCTATAGATACTACCAATGCAATGAAAGGAAAAATATTTG
AACTTCTGAGGGAGTTGGATGAATCAAAAGCTGAACGGGAGAGCCTTGCAAAGAAAATGGACCAGATGGAATGCTACTATGAAGCTCTTGTCCAGGAGCT
TGAGGAACATCAGAGGCAGATGCTGGGGGAGTTGCAGAATCTCAGAAATGAGCATGCTACTTGCTTGTATGCAGCTGCAACCACGAAGCAAGAGATGGAG
ACAATGCGCCAAGATCTGAATGGCCAGCTATTAAGAGTTGTTGAAGAAAAACGTGATTTGGATTCTCTGAACAAGGAATTTGAAAGAAGGACTGTTACTG
CTGAGGCAGCACTCAAACGGGCACGCCTGAATTATTCAATTGCTGTCGACCAGTTACAAAAGGATCTGGAACTGCTCTCAGTCCAGGTTTTGTCCATGTT
TGAGACTAATGAGAATCTCATCCGGCAAGCTTTTGTAGATTCTTCACAATCATGCTTTGAAGGAAACCCAATAACCACAGAGAGCCAGAGATCTGGTTCA
AGTGAAGTTCGCATGGGGAAACTCTTTCAGTTCCAAAATCAGTTTGTCGGGAAGAAAAAACAGCAGTTGGGTGGTGATATTCTTTTAGATGACTTGAAAA
GATCTCTACATTTGCAGGAAGGGCTTTACAGAAAGGTTGAAGAAGAAGCTTGTGAAATGCATTTTGATAATTTATACTTGGATGTACTTTCAAATGTTCT
ACAGGAAACTTTGCTTGAGGCCAGTGACAATGTCAAATGTATGAGAGAGAAAATAAACAAACTTATGCAGCAGCTGGAGCTTTCAACTGAGTCCAAGGAG
TTGTTGTCGCAAAAGCTGCACTCTGCACTGAATGATGTTCATGCCTTAAATGAGCATAGGGCAACTTGCGTTGCCAAGTGCAGTGATATGGCTCAGCAAA
ACCAAGTTTTGGAAGCAAATTTGCAAAATGTCACATGTGAAAATCACCTTCTTTTGCAGAAGATTGCTGAATGGGAGTCTCTAGTGATGCACTACAGAAG
TTACGAGAGCATGTATAAGGCTTCTGCTGCAGAGAACACGGAGTTGGCATGTTTATTGGAAAAGAAAACCCTAGAAAATTGTGATCTTCAAAATGAGATC
TTCTCTTTGCAGGAAGAGCTGAAAACTTTCAGAAATGAATTTGATGATCTGGCTTCTGTGAAGGAAAAGCTACAGGATCTAGTCAATTTCATGGAGAGTA
AGTTGCAGAACTTGCTGGCGTCCTATGACAAAAGTATAAATGGATTGCCTCCTAGTGAATCTGGTGATCATGATTTAAAGCCCCAGGATTTAATTGGTGT
CATGATGCAATTGGAAGAGCTTCAGCATAATTCATGTGAAAGGATTCTCCTGCTCATGGAAGAGAAGAAAGGTCTGGTTCACGAGAGAGATATCGCTCAA
GTGTCCATCACTGCAGCTAAATCAGAGATCGCATTGGTGAAACAGAAGTTTGAACGTGATATACTGAACATGGTGGATAAATTCAATGTGTCAAATGCTT
TGGTTGAGCAGCTTCAGTTGGACGTTGAGGGTATTGCTTACAAACTCAAGGTTAGCTCTGAAGCTGAAGAAAAATATGCACAGCTACACAATGAGCTATT
TTCTGATCTTGATCGTTTGGAAGCTCAGTTGAAAGAACTTATTTCTATGAACCAGGACATTGGTCATGAAATCTTGGCATTAGATACTGTTGCTAGTGAA
CTTGATAAGACTAAACTGGCTGCAGCAGAACTTGTGATAGAAAACCAAGCTTTGATGGCATCTATACAGGATAAAAATGAGGTGTCCTCAGGGATTGCAT
CAGAACTCGAGAGTTTGAAAGGAAGTTTGCAATCTCTGCATGATGAAAACCAAGCTTTGATGGCATCTTCACAAGATAAGAAGGAGTCGTCTGCTCAATT
GGCTTCAGAGTTGAGCAACTTGAAAGATAGCATACAATCTCTGCATGATGAAAACCAAGCTCTGATGGAAATTTTACGGAATAAAACTGAGGAAGCTGGC
AACCTTGCATCAGAGCTAAATAGTTTAAAAGAAAATTTGCGATTTTTGCGTGATGAAAATCAAGCTTTGATGGCATCTTCACAGGATAAAGAGGAGGAAC
ATGCCAAGCTTGCAATGGAACTAAACTGTTTGAAAGAATGTTTGCAGACTCTGCATGATGAAAATCAAGCTCAGATGACATCGGCAATGGATGCGAAGGA
GGAATCTACCAAGCTTGTGTCAGAGATAAACAGTTTGAAAGGAAGCTTGCAATCTCTGCATGGTGAAAAACAAGCTCTGATGATATCTACGAGGGATAAA
ACTGAGGAATCTTCCAAGCTTGCATCAGAGCTGAACATTTTGAAAGAGAGTTCACAATCTTTGCATTGTGAAAACCAAGTTTTGATGGCAGGTTTGCAGG
ATAAGACAGAGGAATCTGCCAGGCTTGCATCAGAACTAAACAGTTTGAGAGAATGCCTGCATACTTTGCAACATGAAAAGCAAGCTTTGATGGTGTTCTT
ACAGGATAAAACTGAGGAGTCTGCCCATCTTGCTTCAGATTTGATCAGTTTGAGAGAGAGTTTGCAATCTCTGCATGATGAGCTGCATGATGAGAGAAGC
TTGCGAGAGGGGTTACAGAGTACAATAGTGGATCTAACTTCACAATTAAACGAGAAGCAGTGCCAGTTGCTCCAATTTGATCACCACAAGTCTGAATTGG
CTCATCTCAAACACCTGGTATCAGATCTAGAATCTGAGAAAGCAAGAGTTTGCCATCTTCTGTTGCAGTCTGAAGAATGTCTTAATAATGCTCGTGAAGA
AGCTTCCACTGTTTCTGCACTAAAAACTCAGTTGTCTGAGATGCATGAACCTCTAATAGCTGCTGATGTTCGATTCATTTTTGCAAAGACTCAATATGAC
AGCGGATTTGAAGTGCTTCTTCATCAACTACACTCCACAGATAGGCTGCTTGCCCAACTTCAGAAGAAGCATATTGACATGGAAACCACCCTTAATCGGT
GTCTTGCAAGTGAAACCCAATATGCTGAAGAGAATGCGAGGTTGTTGACAAATCTCAACTCTGTTCTGTCTGAGCTAGAGGCTTCTATTGCTGAAAACAG
GTTACTTGTCGAGAAAAACAGAGTTGTAAGAGCTGAGCTTGAGGAATTCAAACACAACTCTCAAAATGTTGTGCTTGGTTATATGGAGGATAAAACTCAA
CATTCTCTAGAGGTTGAAAAGCTGAAGTGCATGTTGGTGACATCTGAAGAAGAGATTGATAATCTGGTGTTCTCCAAGGTTGAGCTGGAAGTAAAAGTTC
TAGTACTTGAAGCCAAATTGGATGAACAGCAAGCTCAGATCATCACGCTTGAAGGATACTACGATGAATTAGTGATGGTGCAGAAACACTGCAATGAGCT
TAACCAGAGGCTTTCTGATCAGATTTTGAAGACAGAAGAATTCAGAAACTTGTCTGTCCATTTGAAAGAGCTCAAAGACAAAGCTGATGCTGAGTGTATC
CAGGCTCGTGAAAAAAGAGAACCTGAAGGACCATCTGTGGCCATGCAGGAGTCCTTAAGAATTGCATTTATCAAAGAGCAGTACGAAACAAGGCTGCAAG
AACTGAAACAGCAGTTGTCCATCTCCAAAAAGCACAGTGAGGAAATGCTATGGAAATTACAAGATGCGATTGATGAGATTGAGAATAGGAAGAAATCTGA
AGCTTCTCATCTGAAGAAAAATGAAGAACTGGGAATGAAGATATTGGAATTGGAGGCTGAGTTACAATCTGTAGTTTCTGACAAACGTGAAAAAGTGAAG
GCTTATGACCTGATGAAGGCTGAAATGGAATGCTCATTAATAAGTCTCGAGTGCTGCAAGGAAGAAAAACAAAAACTCGAAGCTTCTTTGGAAGAATGCA
ATGAGGAGAAGTCAAAAATTGCTGTTGAACACACTTTGATGAAAGAATTGCTAGAGAATTCCAAATCACCAGGGAACATGCAGGAAGAACAAAATGATGT
ATCTTGCGAAGTGGATTGCTTGATTGTGGACGCTTCGAACTATGGAATAAAAAGAGCACACACAGTCCCTTTGAATCGTCCATCCAGGAATCCTAACCAG
AAGTGCTTGGGAAGAGATGGTTTGAGGAATTGTGAAGAAGCAGAACTTGCATTCCCTGCTTCTGTTGACAGAGTTGATCATTTGAACACACTTATGCATG
AGCAACCTGAGCAGGATGTTCTTGCGTCTTGTGGGATGAATGGGCTTAAAAGTTCTGCACTTATAAATCAAGACAGGTTGCTGCATAGTGACATGAAGCA
TTTAGCTATCATCAATGATCATTTCAGAGCTGAAAGTTTAAAGTCTAGCATGGACCACTTAAGTAATGAGTTAGAAAGGATGAAAAACGAGAATTCACTT
TTGTTGCAAGATGATCATGATTTTGACCAAAAGTTTCCAGGCTTACAAAGTGAATTTATGAAGCTACAGAAGGCCAATGAAGAACTAGGAAGCATGTTCC
CCTTGTTTAATGAATTCTCAGGGAGTGGAAACGCATTAGAGAGGGTGCTTGCATTAGAAATTGAGCTTGCTGAAGCATTGCAAGCAAAGAAGAGATCAAG
CATCCTTTTTCAGAGTTCATTCTTTAAACAACACAGTGACGAGGAAGCTGTATTCAAAAGCTTTAGAGATATCAATGAGTTGATCAAGGACATGCTGGAA
CTAAAGGGAAGGTATACAACTGTGGAGACTCAACTAAAAGAGATGCATGACCGCTATTCTCAGCTAAGCCTTCAATTTGCAGAGGTTGAAGGTGAGAGGC
AGAAGCTCACGATGACTCTCAAGAATGTTCGGGCATCAAAGAAAGCCCTATGCGTAAACCGGTCCTCATCAGCTTCCCTTGGGGACCATTCCTCCTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G170066.2 pacid=42803226 polypeptide=Potri.005G170066.2.p locus=Potri.005G170066 ID=Potri.005G170066.2.v4.1 annot-version=v4.1
MSRITRWKNEKTKVKVVFRLQFHATHIPHPGWDKLFISFIPADSGKATAKTTKANVRNGTCKWADPIYETTRLLQDVKTKRYDEKLYKLVISMGSSRSSV
LGEATINLADYADALKPSVVALPLHGSDSGTALHVTVQLLTSKTGFREFEQQRELRERGLQTDQNSPDESSSGKVSSSEGIINDQIDKVNIRVRFKEKSK
DLASLEEEVGPNEEYADSAVGFDGSSNTSESLYAEKHDTSSTHEIDSLKSTVSGDLAGLSLSQSPQLEKGDPSDHRFLAQGTNDWVHAWSSDYSADNDLA
AAYEENGRLRGSLEVAESSVLELKQEVSSLQGHADEIGYEAQTFAKQLASEIASGEEMVKEVSVLKSECSKLKADLEQLKVSQLCPPFSSRNAAEPLQDH
RFQELKLRWIKGLLSMEDKIKELQNNACLGYHESDFRFLHSDVEELIGVLQDLKQGTGLPISSNHLVPCEGSSLKEIREMSLHKNSQFVSESGFDVDLYQ
PELGMLHCLNIPGLVSHEPDSIDTTNAMKGKIFELLRELDESKAERESLAKKMDQMECYYEALVQELEEHQRQMLGELQNLRNEHATCLYAAATTKQEME
TMRQDLNGQLLRVVEEKRDLDSLNKEFERRTVTAEAALKRARLNYSIAVDQLQKDLELLSVQVLSMFETNENLIRQAFVDSSQSCFEGNPITTESQRSGS
SEVRMGKLFQFQNQFVGKKKQQLGGDILLDDLKRSLHLQEGLYRKVEEEACEMHFDNLYLDVLSNVLQETLLEASDNVKCMREKINKLMQQLELSTESKE
LLSQKLHSALNDVHALNEHRATCVAKCSDMAQQNQVLEANLQNVTCENHLLLQKIAEWESLVMHYRSYESMYKASAAENTELACLLEKKTLENCDLQNEI
FSLQEELKTFRNEFDDLASVKEKLQDLVNFMESKLQNLLASYDKSINGLPPSESGDHDLKPQDLIGVMMQLEELQHNSCERILLLMEEKKGLVHERDIAQ
VSITAAKSEIALVKQKFERDILNMVDKFNVSNALVEQLQLDVEGIAYKLKVSSEAEEKYAQLHNELFSDLDRLEAQLKELISMNQDIGHEILALDTVASE
LDKTKLAAAELVIENQALMASIQDKNEVSSGIASELESLKGSLQSLHDENQALMASSQDKKESSAQLASELSNLKDSIQSLHDENQALMEILRNKTEEAG
NLASELNSLKENLRFLRDENQALMASSQDKEEEHAKLAMELNCLKECLQTLHDENQAQMTSAMDAKEESTKLVSEINSLKGSLQSLHGEKQALMISTRDK
TEESSKLASELNILKESSQSLHCENQVLMAGLQDKTEESARLASELNSLRECLHTLQHEKQALMVFLQDKTEESAHLASDLISLRESLQSLHDELHDERS
LREGLQSTIVDLTSQLNEKQCQLLQFDHHKSELAHLKHLVSDLESEKARVCHLLLQSEECLNNAREEASTVSALKTQLSEMHEPLIAADVRFIFAKTQYD
SGFEVLLHQLHSTDRLLAQLQKKHIDMETTLNRCLASETQYAEENARLLTNLNSVLSELEASIAENRLLVEKNRVVRAELEEFKHNSQNVVLGYMEDKTQ
HSLEVEKLKCMLVTSEEEIDNLVFSKVELEVKVLVLEAKLDEQQAQIITLEGYYDELVMVQKHCNELNQRLSDQILKTEEFRNLSVHLKELKDKADAECI
QAREKREPEGPSVAMQESLRIAFIKEQYETRLQELKQQLSISKKHSEEMLWKLQDAIDEIENRKKSEASHLKKNEELGMKILELEAELQSVVSDKREKVK
AYDLMKAEMECSLISLECCKEEKQKLEASLEECNEEKSKIAVEHTLMKELLENSKSPGNMQEEQNDVSCEVDCLIVDASNYGIKRAHTVPLNRPSRNPNQ
KCLGRDGLRNCEEAELAFPASVDRVDHLNTLMHEQPEQDVLASCGMNGLKSSALINQDRLLHSDMKHLAIINDHFRAESLKSSMDHLSNELERMKNENSL
LLQDDHDFDQKFPGLQSEFMKLQKANEELGSMFPLFNEFSGSGNALERVLALEIELAEALQAKKRSSILFQSSFFKQHSDEEAVFKSFRDINELIKDMLE
LKGRYTTVETQLKEMHDRYSQLSLQFAEVEGERQKLTMTLKNVRASKKALCVNRSSSASLGDHSS
|
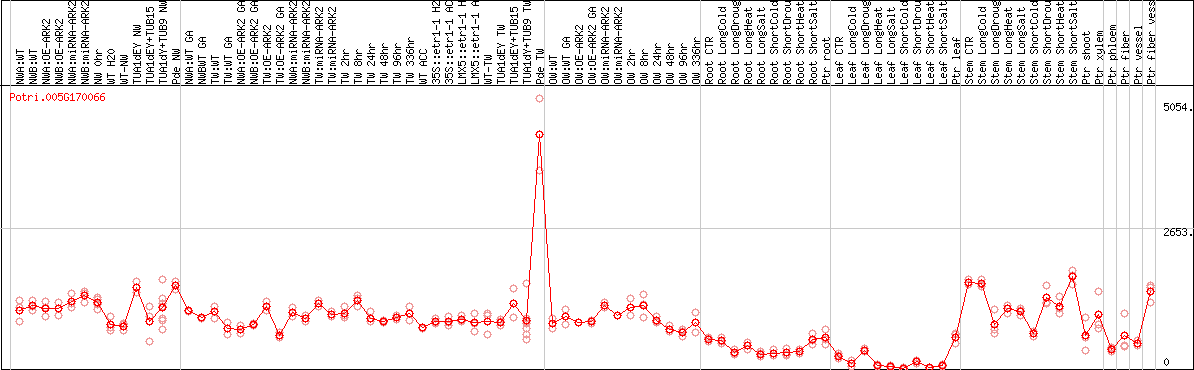
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G170066 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
