External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G58850 82 / 4e-21
HLH2, PAR2
phy rapidly regulated 2 (.1)
AT2G42870 80 / 3e-20
PAR1 ,HLH1
HELIX-LOOP-HELIX 1, phy rapidly regulated 1 (.1)
AT3G17100 46 / 1e-06
bHLH
bHLH147, AIF3
sequence-specific DNA binding transcription factors (.1.2)
AT5G57780 45 / 2e-06
P1R1
P1R1, unknown protein
AT4G30410 45 / 2e-06
sequence-specific DNA binding transcription factors (.1.2)
AT1G09250 45 / 3e-06
bHLH
bHLH149, AIF4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G05800 42 / 3e-05
bHLH
bHLH150, AIF1
AtBS1(activation-tagged BRI1 suppressor 1)-interacting factor 1 (.1)
AT3G06590 42 / 3e-05
bHLH
bHLH148, AIF2
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT2G47270 40 / 5e-05
bHLH
bHLH151, UPB1
UPBEAT1, sequence-specific DNA binding transcription factors;transcription regulators (.1)
AT2G43060 39 / 0.0002
bHLH
AtIBH1, bHLH158
ILI1 binding bHLH 1 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026730
50 / 2e-08
AT2G43060 97 / 4e-26
ILI1 binding bHLH 1 (.1)
Lus10001447
45 / 7e-07
AT2G47270 69 / 2e-16
UPBEAT1, sequence-specific DNA binding transcription factors;transcription regulators (.1)
Lus10015500
46 / 1e-06
AT4G30410 114 / 6e-32
sequence-specific DNA binding transcription factors (.1.2)
Lus10002419
45 / 1e-06
AT2G47270 67 / 4e-16
UPBEAT1, sequence-specific DNA binding transcription factors;transcription regulators (.1)
Lus10025511
45 / 2e-06
AT2G43060 98 / 9e-27
ILI1 binding bHLH 1 (.1)
Lus10037783
44 / 8e-06
AT3G06590 172 / 9e-54
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10017065
44 / 8e-06
AT3G06590 171 / 1e-53
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10030641
42 / 8e-06
AT1G23965 42 / 3e-06
unknown protein
Lus10030844
39 / 0.0001
AT1G23965 48 / 2e-08
unknown protein
Lus10038432
39 / 0.0009
AT1G30670 108 / 4e-28
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Potri.005G201500.1 pacid=42805531 polypeptide=Potri.005G201500.1.p locus=Potri.005G201500 ID=Potri.005G201500.1.v4.1 annot-version=v4.1
ATGGAGAATAGTTTAAATCAAGAAATAACACCCACATTTCATAAAAGAAAAGAAAGTAAAACAACCCACAAAAAACCTCATGAGATGGGTAATGCTAAGC
ATGAATCTTTAGTTAGTTTGACAAGGAGGAAGAGGGCAGATAATGTTCATAAGAGAAACAGTAGAAAGGGAAATGAAATATTGAAGGCTCAAGAAAAGGA
TCAAGCGTCGACGGTCGGTGTTGAAGACGATGGGAAGGCAGAAGTGGAGAGAAAGATTGTGGCATTGCAAAGGATTGTCCCTGGGGGTGAGTTGTTCGGG
GTAGACAAGCTGTTTGAAGAGACTGCCGATTATATAATGGCTTTGCAGTGCCAAATTAAAGCCATGAGAGTTCTTGCTGGTTTTCTTGAAGGGCTAGAAA
AGGAAAAGAGGAAGTTTGGAGGTTAA
AA sequence
>Potri.005G201500.1 pacid=42805531 polypeptide=Potri.005G201500.1.p locus=Potri.005G201500 ID=Potri.005G201500.1.v4.1 annot-version=v4.1
MENSLNQEITPTFHKRKESKTTHKKPHEMGNAKHESLVSLTRRKRADNVHKRNSRKGNEILKAQEKDQASTVGVEDDGKAEVERKIVALQRIVPGGELFG
VDKLFEETADYIMALQCQIKAMRVLAGFLEGLEKEKRKFGG
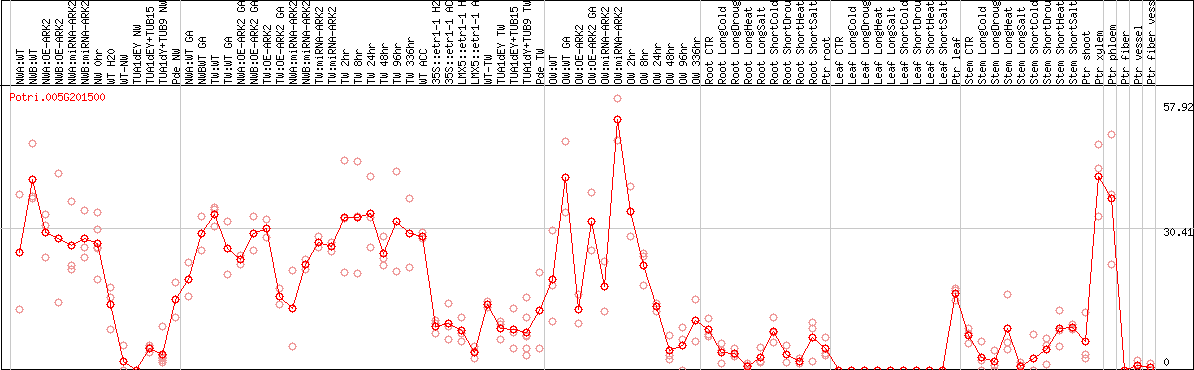
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G201500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.