External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G06080 87 / 8e-22
TBL10
TRICHOME BIREFRINGENCE-LIKE 10, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
AT5G19160 83 / 1e-20
TBL11
TRICHOME BIREFRINGENCE-LIKE 11 (.1)
AT1G78710 62 / 4e-13
TBL42
TRICHOME BIREFRINGENCE-LIKE 42 (.1.2)
AT1G60790 62 / 6e-13
TBL2
TRICHOME BIREFRINGENCE-LIKE 2, Plant protein of unknown function (DUF828) (.1)
AT5G06700 61 / 1e-12
TBR
TRICHOME BIREFRINGENCE, Plant protein of unknown function (DUF828) (.1)
AT5G20590 60 / 2e-12
TBL5
TRICHOME BIREFRINGENCE-LIKE 5 (.1)
AT5G58600 60 / 3e-12
TBL44, PMR5
TRICHOME BIREFRINGENCE-LIKE 44, POWDERY MILDEW RESISTANT 5, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
AT2G30900 60 / 3e-12
TBL43
TRICHOME BIREFRINGENCE-LIKE 43 (.1)
AT5G49340 58 / 1e-11
TBL4
TRICHOME BIREFRINGENCE-LIKE 4 (.1)
AT5G06230 57 / 2e-11
TBL9
TRICHOME BIREFRINGENCE-LIKE 9 (.1.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028025
88 / 4e-22
AT3G06080 520 / 0.0
TRICHOME BIREFRINGENCE-LIKE 10, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Lus10003734
86 / 2e-21
AT3G06080 540 / 0.0
TRICHOME BIREFRINGENCE-LIKE 10, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Lus10034041
79 / 6e-19
AT5G19160 605 / 0.0
TRICHOME BIREFRINGENCE-LIKE 11 (.1)
Lus10041219
75 / 1e-17
AT5G19160 514 / 0.0
TRICHOME BIREFRINGENCE-LIKE 11 (.1)
Lus10034039
69 / 2e-15
AT3G06080 553 / 0.0
TRICHOME BIREFRINGENCE-LIKE 10, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Lus10030590
64 / 4e-14
AT1G60790 226 / 7e-71
TRICHOME BIREFRINGENCE-LIKE 2, Plant protein of unknown function (DUF828) (.1)
Lus10030900
61 / 1e-12
AT1G60790 371 / 2e-127
TRICHOME BIREFRINGENCE-LIKE 2, Plant protein of unknown function (DUF828) (.1)
Lus10037726
61 / 1e-12
AT5G49340 435 / 3e-150
TRICHOME BIREFRINGENCE-LIKE 4 (.1)
Lus10011932
61 / 2e-12
AT5G20590 543 / 0.0
TRICHOME BIREFRINGENCE-LIKE 5 (.1)
Lus10004159
60 / 2e-12
AT5G06700 552 / 0.0
TRICHOME BIREFRINGENCE, Plant protein of unknown function (DUF828) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0264
SGNH_hydrolase
PF13839
PC-Esterase
GDSL/SGNH-like Acyl-Esterase family found in Pmr5 and Cas1p
Representative CDS sequence
>Potri.005G208301.1 pacid=42805596 polypeptide=Potri.005G208301.1.p locus=Potri.005G208301 ID=Potri.005G208301.1.v4.1 annot-version=v4.1
ATGTCAGCACGGAGAAAGGATGGCCACCCATCCGTGTACTATCTAGGATCAAGTATAGGTCCACCCTTCCTACACCGTCAAGACTGCAGCCATTGGTACC
TGCCTGGTGTACCTGATTCCTGGAATGAGCTTCTTTATACGCTTCTTCTCAAGTGGGAGTCTGTTCATGCACAAAATCTAACAGAATCTTCTCAAGCTAC
TCCGTGA
AA sequence
>Potri.005G208301.1 pacid=42805596 polypeptide=Potri.005G208301.1.p locus=Potri.005G208301 ID=Potri.005G208301.1.v4.1 annot-version=v4.1
MSARRKDGHPSVYYLGSSIGPPFLHRQDCSHWYLPGVPDSWNELLYTLLLKWESVHAQNLTESSQATP
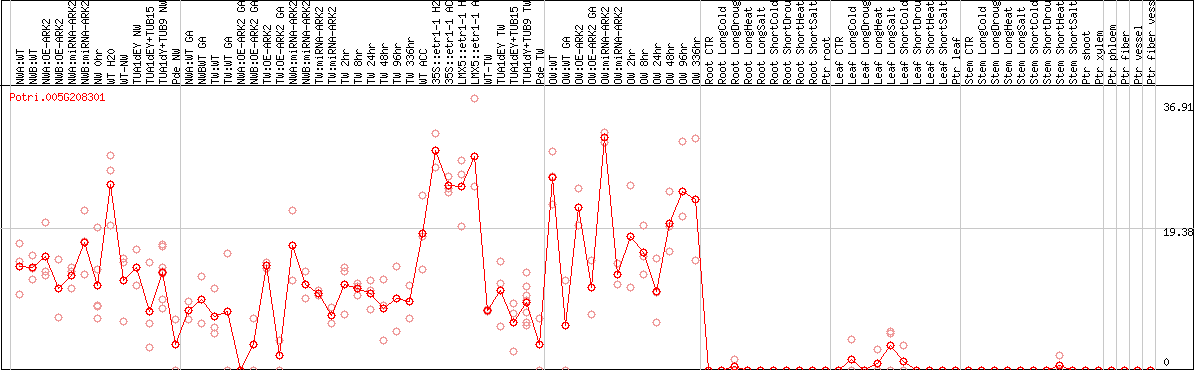
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G208301 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.