External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G48540 52 / 1e-08
receptor-like protein kinase-related family protein (.1)
AT4G23140 46 / 2e-06
RLK5, CRK6
cysteine-rich RLK (RECEPTOR-like protein kinase) 6 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 6 (.2)
AT2G31620 44 / 1e-05
Receptor-like protein kinase-related family protein (.1)
AT4G05200 44 / 2e-05
CRK25
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
AT3G45860 41 / 9e-05
CRK4
cysteine-rich RLK (RECEPTOR-like protein kinase) 4 (.1)
AT3G22060 40 / 0.0001
Receptor-like protein kinase-related family protein (.1)
AT3G58310 40 / 0.0002
Domain of unknown function (DUF26) (.1)
AT4G23280 40 / 0.0002
CRK20
cysteine-rich RLK (RECEPTOR-like protein kinase) 20 (.1)
AT4G23170 40 / 0.0002
CRK9, EP1
CYSTEINE-RICH RLK \(RECEPTOR-LIKE PROTEIN KINASE\) 9, receptor-like protein kinase-related family protein (.1)
AT1G70690 40 / 0.0003
HWI1, PDLP5
PLASMODESMATA-LOCATED PROTEIN 5, HOPW1-1-INDUCED GENE1, Receptor-like protein kinase-related family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G030650
57 / 2e-10
AT4G05200 476 / 1e-159
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Potri.004G023700
53 / 7e-09
AT4G05200 474 / 3e-159
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Potri.004G024140
50 / 9e-08
AT4G23180 497 / 3e-168
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Potri.004G023876
50 / 1e-07
AT4G23180 489 / 5e-165
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Potri.004G024000
49 / 2e-07
AT4G23180 489 / 4e-165
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Potri.004G024900
47 / 6e-07
AT4G23180 506 / 1e-171
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Potri.004G025900
47 / 6e-07
AT4G23130 573 / 0.0
RECEPTOR-LIKE PROTEIN KINASE 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 5 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 5 (.2)
Potri.007G120600
47 / 6e-07
AT3G22060 214 / 2e-69
Receptor-like protein kinase-related family protein (.1)
Potri.007G120601
45 / 2e-06
AT3G22060 199 / 9e-65
Receptor-like protein kinase-related family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028026
146 / 6e-46
AT4G23290 44 / 1e-05
cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.2)
Lus10003735
130 / 1e-39
AT4G23210 43 / 3e-05
cysteine-rich RLK (RECEPTOR-like protein kinase) 13 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 13 (.2), cysteine-rich RLK (RECEPTOR-like protein kinase) 13 (.3)
Lus10011894
61 / 2e-12
AT3G60720 38 / 9e-04
plasmodesmata-located protein 8 (.1)
Lus10022828
60 / 6e-12
AT2G01660 43 / 2e-05
plasmodesmata-located protein 6 (.1.2)
Lus10007631
57 / 3e-10
AT4G23180 257 / 1e-76
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Lus10007504
56 / 3e-10
AT3G22040 42 / 3e-05
Domain of unknown function (DUF26) (.1)
Lus10026169
55 / 4e-10
AT4G05200 51 / 4e-08
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Lus10008656
53 / 1e-09
AT4G05200 44 / 8e-06
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Lus10028980
54 / 2e-09
AT3G21945 39 / 9e-04
Receptor-like protein kinase-related family protein (.1)
Lus10026168
53 / 3e-09
ND 37 / 0.005
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01657
Stress-antifung
Salt stress response/antifungal
Representative CDS sequence
>Potri.005G208400.1 pacid=42803219 polypeptide=Potri.005G208400.1.p locus=Potri.005G208400 ID=Potri.005G208400.1.v4.1 annot-version=v4.1
ATGAAGAGAACAAGAAACATGGGTTTTCCTCCAAAACTGGCCCTGGTTTTCATCGGATTCTTTGGATGGTGCGTCATTGCTACAAGTGTTCCTGATACCA
ACGTGACAACTGTTCTCTGCAACTCAGGGGTGTACTCAAAGGGTGATCCTTTTGGTACTAGTCTAGCTTATGTCCTTGCAGAAATTGAGGCTGTAACCCC
AACGAGCAAGAATTATGACTACTTCAACATCTCTCCTTATCCTAATGCTGTTGCCTATGGTCATGCTGCTTGTAATCAGAACCTCACCAGCTCCGACTGC
TCATCATGTCTTGGTACGGCTAAAACAGCCATGTTTGCGAGTTGTCAAAGCCGAATAGGGGCTCGTTCGGTGCTCCATGATTGTACGATTAGGTATGAGC
AATACCCATTTTCTGATTAA
AA sequence
>Potri.005G208400.1 pacid=42803219 polypeptide=Potri.005G208400.1.p locus=Potri.005G208400 ID=Potri.005G208400.1.v4.1 annot-version=v4.1
MKRTRNMGFPPKLALVFIGFFGWCVIATSVPDTNVTTVLCNSGVYSKGDPFGTSLAYVLAEIEAVTPTSKNYDYFNISPYPNAVAYGHAACNQNLTSSDC
SSCLGTAKTAMFASCQSRIGARSVLHDCTIRYEQYPFSD
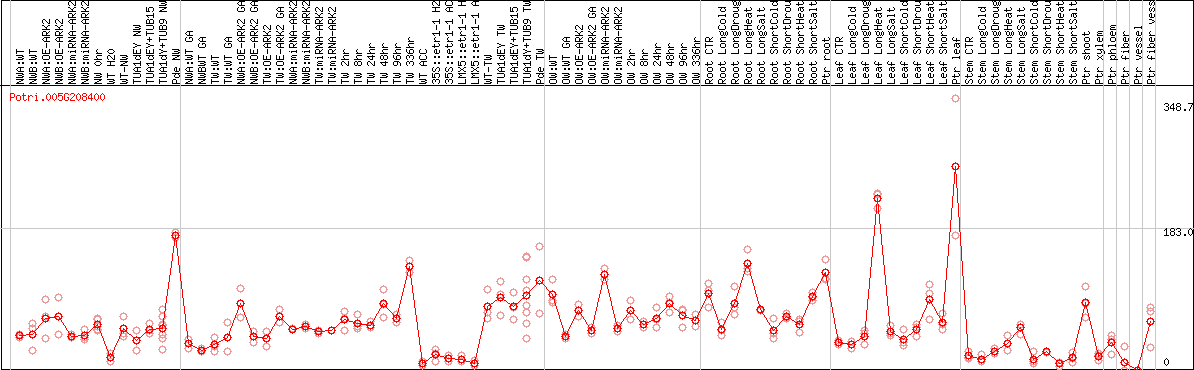
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G208400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.