External link
Symbol
BET11.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G58170 206 / 4e-70
ATBET11, ATBS14A
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14A, BET1P/SFT1P-like protein 14A (.1)
AT4G14455 150 / 6e-48
ATBS14B ,ATBET12
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14BB, Target SNARE coiled-coil domain protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G041500
235 / 1e-81
AT3G58170 202 / 7e-69
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14A, BET1P/SFT1P-like protein 14A (.1)
Potri.010G074100
156 / 2e-50
AT4G14455 147 / 1e-46
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14BB, Target SNARE coiled-coil domain protein (.1)
Potri.011G059700
39 / 0.0004
AT1G28490 310 / 2e-107
syntaxin of plants 61 (.1.2)
Potri.008G164600
36 / 0.0007
ND /
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033868
201 / 5e-68
AT3G58170 187 / 1e-62
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14A, BET1P/SFT1P-like protein 14A (.1)
Lus10024876
208 / 6e-64
AT3G12770 886 / 0.0
mitochondrial editing factor 22 (.1)
Lus10041158
144 / 2e-45
AT4G14455 164 / 6e-53
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14BB, Target SNARE coiled-coil domain protein (.1)
Lus10000706
154 / 2e-44
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Lus10021879
99 / 9e-28
AT4G14455 110 / 4e-32
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14BB, Target SNARE coiled-coil domain protein (.1)
Lus10014756
97 / 2e-24
AT5G11010 440 / 2e-152
Pre-mRNA cleavage complex II protein family (.1.2.3)
Lus10009926
87 / 1e-23
AT3G58170 81 / 6e-21
ARABIDOPSIS THALIANA BET1P/SFT1P-LIKE PROTEIN 14A, BET1P/SFT1P-like protein 14A (.1)
Lus10018642
38 / 0.0008
AT1G28490 334 / 7e-117
syntaxin of plants 61 (.1.2)
PFAM info
Representative CDS sequence
>Potri.005G221600.5 pacid=42804653 polypeptide=Potri.005G221600.5.p locus=Potri.005G221600 ID=Potri.005G221600.5.v4.1 annot-version=v4.1
ATGAATTCGAGGAGGGACATCCGTAACAATAGAGCTGCTCTTTTTGATGGAATTGAGGAGGGTGGCATCAGAGCCTCATCCTCTTACTCCTCCCATGAGA
TTGATGAGCAAGATAATGAAAGAGCTTTGGAAGGATTGCAAGATAGAGTCATTCTGCTGAAGAGACTGTCAGGTGATATAAATGAAGAGGTGGACAACCA
TAATCTTATGCTAGACAGAATGGGCAATGATATGGATTCTTCCAGGGGAGTGCTGTCAGGAACCATGGATCGCTTTAAGATGGTATTTGAAACCAAATCA
AGCCGTAGAATGTTTACACTCGTGGCATCATTTGTGGTCCTATTCCTAATTATATACTATCTCACTAGGTAA
AA sequence
>Potri.005G221600.5 pacid=42804653 polypeptide=Potri.005G221600.5.p locus=Potri.005G221600 ID=Potri.005G221600.5.v4.1 annot-version=v4.1
MNSRRDIRNNRAALFDGIEEGGIRASSSYSSHEIDEQDNERALEGLQDRVILLKRLSGDINEEVDNHNLMLDRMGNDMDSSRGVLSGTMDRFKMVFETKS
SRRMFTLVASFVVLFLIIYYLTR
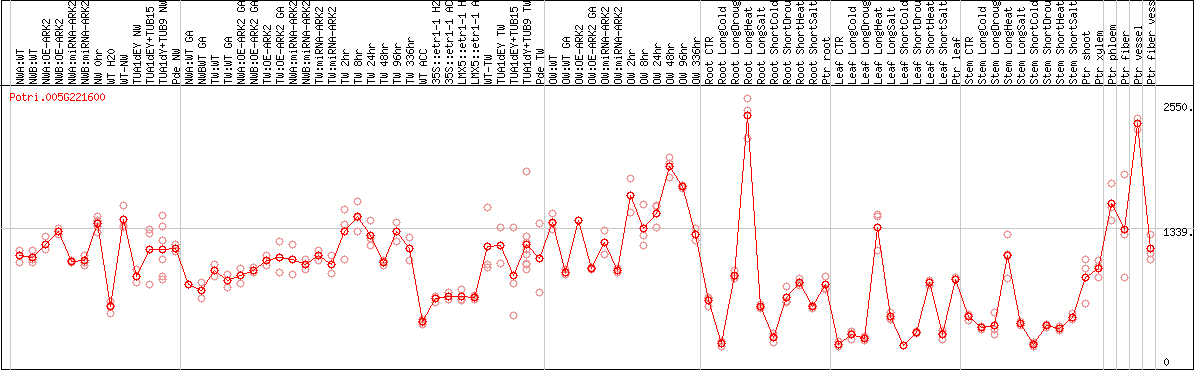
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G221600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.