External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G28530 271 / 6e-90
NAC
ANAC074
NAC domain containing protein 74 (.1.2)
AT1G56010 202 / 3e-63
NAC
NAC1, ANAC021, ANAC022
Arabidopsis NAC domain containing protein 22, Arabidopsis NAC domain containing protein 21, NAC domain containing protein 1 (.1.2)
AT1G76420 201 / 5e-63
NAC
NAC368, CUC3, ANAC031
CUP SHAPED COTYLEDON3, Arabidopsis NAC domain containing protein 31, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT2G24430 201 / 7e-63
NAC
ANAC039, ANAC038
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
AT3G12977 197 / 3e-62
NAC
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G53950 192 / 7e-59
NAC
ATCUC2, CUC2, ANAC098
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G07680 189 / 2e-58
NAC
ANAC079, ATNAC4, ANAC080
Arabidopsis NAC domain containing protein 79, NAC domain containing protein 80 (.1.2)
AT5G61430 189 / 6e-58
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
AT3G18400 188 / 6e-58
NAC
ANAC058
NAC domain containing protein 58 (.1)
AT3G15170 183 / 3e-56
NAC
ATNAC1, CUC1, ANAC054
CUP-SHAPED COTYLEDON1, Arabidopsis NAC domain containing protein 54, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G037100
470 / 1e-169
AT4G28530 276 / 6e-92
NAC domain containing protein 74 (.1.2)
Potri.007G065400
209 / 1e-66
AT1G56010 337 / 4e-116
Arabidopsis NAC domain containing protein 22, Arabidopsis NAC domain containing protein 21, NAC domain containing protein 1 (.1.2)
Potri.005G098200
204 / 2e-64
AT1G56010 332 / 6e-114
Arabidopsis NAC domain containing protein 22, Arabidopsis NAC domain containing protein 21, NAC domain containing protein 1 (.1.2)
Potri.002G005800
201 / 4e-62
AT1G76420 326 / 7e-110
CUP SHAPED COTYLEDON3, Arabidopsis NAC domain containing protein 31, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.005G255900
200 / 6e-62
AT1G76420 295 / 4e-98
CUP SHAPED COTYLEDON3, Arabidopsis NAC domain containing protein 31, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.006G277000
194 / 4e-60
AT2G24430 340 / 1e-116
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Potri.018G003800
194 / 9e-60
AT2G24430 350 / 1e-120
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Potri.015G046800
190 / 1e-58
AT3G18400 326 / 1e-111
NAC domain containing protein 58 (.1)
Potri.012G001400
190 / 2e-58
AT5G61430 448 / 1e-158
NAC domain containing protein 100 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022915
355 / 1e-123
AT4G28530 274 / 7e-91
NAC domain containing protein 74 (.1.2)
Lus10033905
326 / 5e-112
AT4G28530 271 / 2e-89
NAC domain containing protein 74 (.1.2)
Lus10024908
247 / 1e-81
AT4G28530 200 / 2e-62
NAC domain containing protein 74 (.1.2)
Lus10042466
209 / 2e-65
AT1G56010 273 / 9e-90
Arabidopsis NAC domain containing protein 22, Arabidopsis NAC domain containing protein 21, NAC domain containing protein 1 (.1.2)
Lus10003548
207 / 7e-61
AT1G08620 455 / 4e-142
Transcription factor jumonji (jmj) family protein / zinc finger (C5HC2 type) family protein (.1), Transcription factor jumonji (jmj) family protein / zinc finger (C5HC2 type) family protein (.2)
Lus10005537
192 / 1e-59
AT5G53950 317 / 3e-107
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10009029
189 / 7e-58
AT3G18400 311 / 3e-105
NAC domain containing protein 58 (.1)
Lus10009669
189 / 8e-58
AT3G18400 308 / 3e-104
NAC domain containing protein 58 (.1)
Lus10041924
181 / 1e-56
AT5G53950 281 / 9e-95
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10020165
185 / 7e-56
AT2G24430 306 / 2e-102
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.005G225800.1 pacid=42803432 polypeptide=Potri.005G225800.1.p locus=Potri.005G225800 ID=Potri.005G225800.1.v4.1 annot-version=v4.1
ATGGGGCTAAGAGACATTGGAGCAACACTCCCTCCTGGTTTTAGGTTCTATCCTAGTGATGAAGAGCTCGTTTGCCATTACCTCTACAAAAAGATCACAA
ATGAACAAGTCTTGAAGGGTACTTTGGTGGAAGTTGACTTGCATACTTGTGAGCCATGGCAACTTCCTGAGGTGGCTAAGCTCAACGCAACTGAGTGGTA
TTTCTTCAGTTTCCGGGATCGAAAGTACGCCACCGGATTCAGAACCAATCGTGCTACAACAACTGGATACTGGAAGGCTACAGGGAAGGATCGTACCGTT
TTGGACCCTACTACTCGTGAAGTTGTAGGGATGAGAAAGACATTGGTTTTCTATAGGAACAGAGCTCCCAATGGAATCAAAACTGGTTGGATCATGCATG
AGTTTCGTCTTGAGACCCCACATATGCACCCTAAGGAGGACTGGGTGTTGTGCAGAGTATTCCACAAATGCAAAGCAGAAGATTACAACGCAAAATTCAG
CCCACAGCTCATGTTCGAGACCACAACTAGTAGTGGCACTGTAAACTTAGCAGCATCACCTTTAAGTACTGATCATCAAACTTTGCCTTGTGGGTACCAG
CAAATGACATCATTATCATCAAACCCACCCCATCAAAACCAAGACCAAAGCTCTTTGCTAAACCTTCTTCAGCTATCTCAGGACAGGAACGCCAACCCCA
ATATTAGCACCGACATCAGCTCCAAAGCGGTTGATGATTATGCCTTCTTGTGGGATGACATGAACTTGGAAGAAACTAGCTTGGGAGATGGGGTGCTGGC
TTCAAACCTGGAAGACATGAGATTTGAGATTGATCATAACAGCATGGTCTTCATATGA
AA sequence
>Potri.005G225800.1 pacid=42803432 polypeptide=Potri.005G225800.1.p locus=Potri.005G225800 ID=Potri.005G225800.1.v4.1 annot-version=v4.1
MGLRDIGATLPPGFRFYPSDEELVCHYLYKKITNEQVLKGTLVEVDLHTCEPWQLPEVAKLNATEWYFFSFRDRKYATGFRTNRATTTGYWKATGKDRTV
LDPTTREVVGMRKTLVFYRNRAPNGIKTGWIMHEFRLETPHMHPKEDWVLCRVFHKCKAEDYNAKFSPQLMFETTTSSGTVNLAASPLSTDHQTLPCGYQ
QMTSLSSNPPHQNQDQSSLLNLLQLSQDRNANPNISTDISSKAVDDYAFLWDDMNLEETSLGDGVLASNLEDMRFEIDHNSMVFI
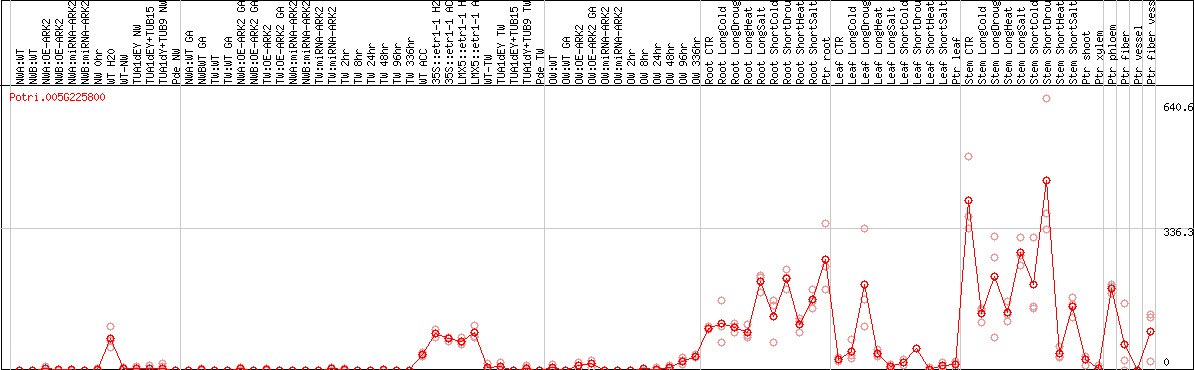
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G225800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.