External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G51030 162 / 4e-53
ATTRX1, ATTRXH1
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
AT1G45145 153 / 1e-49
LIV1, ATTRX5, ATH5
LOCUS OF INSENSITIVITY TO VICTORIN 1, thioredoxin H-type 5 (.1)
AT1G19730 148 / 1e-47
ATTRX4, ATH4
thioredoxin H-type 4, Thioredoxin superfamily protein (.1)
AT5G42980 140 / 2e-44
ATTRXH3, ATTRX3, ATH3
THIOREDOXIN H3, thioredoxin H-type 3, thioredoxin 3 (.1)
AT5G39950 110 / 2e-32
ATTRXH2, ATTRX2, ATH2
Arabidopsis thioredoxin h2, thioredoxin 2 (.1)
AT3G56420 104 / 1e-29
Thioredoxin superfamily protein (.1)
AT3G17880 108 / 2e-29
ATHIP2, ATTDX
HSC-70 INTERACTING PROTEIN, ARABIDOPSIS THALIANA HSP70-INTERACTING PROTEIN 2, tetraticopeptide domain-containing thioredoxin (.1.2)
AT1G59730 95 / 2e-26
ATH7
thioredoxin H-type 7 (.1)
AT2G40790 94 / 9e-26
ATCXXS2
C-terminal cysteine residue is changed to a serine 2 (.1)
AT3G08710 94 / 1e-25
TRXH9, ATH9
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G030000
201 / 2e-68
AT3G51030 160 / 3e-52
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Potri.007G018000
153 / 2e-49
AT3G51030 186 / 2e-62
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Potri.012G045000
124 / 1e-35
AT3G17880 379 / 4e-131
HSC-70 INTERACTING PROTEIN, ARABIDOPSIS THALIANA HSP70-INTERACTING PROTEIN 2, tetraticopeptide domain-containing thioredoxin (.1.2)
Potri.015G036000
119 / 2e-33
AT3G17880 396 / 3e-137
HSC-70 INTERACTING PROTEIN, ARABIDOPSIS THALIANA HSP70-INTERACTING PROTEIN 2, tetraticopeptide domain-containing thioredoxin (.1.2)
Potri.017G076700
109 / 5e-32
AT5G39950 168 / 7e-55
Arabidopsis thioredoxin h2, thioredoxin 2 (.1)
Potri.008G194100
105 / 5e-30
AT1G59730 119 / 3e-35
thioredoxin H-type 7 (.1)
Potri.006G110100
100 / 3e-28
AT3G08710 178 / 1e-58
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
Potri.016G138800
98 / 3e-27
AT3G08710 208 / 1e-70
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
Potri.019G062000
96 / 3e-26
AT3G08710 164 / 6e-53
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024293
153 / 2e-49
AT3G51030 162 / 4e-53
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10000802
152 / 5e-49
AT3G51030 165 / 5e-54
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10041799
149 / 1e-47
AT3G51030 182 / 8e-61
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10014277
147 / 6e-47
AT3G51030 185 / 4e-62
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10028349
144 / 6e-46
AT3G51030 179 / 1e-59
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10025979
128 / 2e-39
AT3G51030 167 / 3e-55
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10005258
113 / 2e-33
AT5G39950 189 / 4e-63
Arabidopsis thioredoxin h2, thioredoxin 2 (.1)
Lus10030666
113 / 2e-33
AT5G39950 189 / 3e-63
Arabidopsis thioredoxin h2, thioredoxin 2 (.1)
Lus10014186
113 / 3e-33
AT3G08710 192 / 3e-64
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
Lus10022727
108 / 3e-31
AT3G08710 189 / 6e-63
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF13098
Thioredoxin_2
Thioredoxin-like domain
Representative CDS sequence
>Potri.005G232700.1 pacid=42802704 polypeptide=Potri.005G232700.1.p locus=Potri.005G232700 ID=Potri.005G232700.1.v4.1 annot-version=v4.1
ATGGCCGAAGAAGGACAAGTTATTGCCTGCCACACAGTGGATACCTGGAAAGAGCATTTCGAGAAGGGAAAAGGGTCTCAGAAACTGATTGTCGTGGATT
TTACTGCTTCATGGTGTCCACCATGTAAAATGATTGCTCCAATCTTCGCCGAGTTGGCGAAGAAGTTTCCCAATGTCACATTCTTGAAGGTGGATGTGGA
TGAATTGAAGGCTGTTGCTGAGGAGTGGAATGTGGAGGCAATGCCAACTTTTATTTTCCTGAAAGATGGAAAATTAGTGGACAAAACTGTGGGTGCTGAT
AAAGATGGCCTGCCAACACTGGTTGCAAAGCACGCAACTGCATAA
AA sequence
>Potri.005G232700.1 pacid=42802704 polypeptide=Potri.005G232700.1.p locus=Potri.005G232700 ID=Potri.005G232700.1.v4.1 annot-version=v4.1
MAEEGQVIACHTVDTWKEHFEKGKGSQKLIVVDFTASWCPPCKMIAPIFAELAKKFPNVTFLKVDVDELKAVAEEWNVEAMPTFIFLKDGKLVDKTVGAD
KDGLPTLVAKHATA
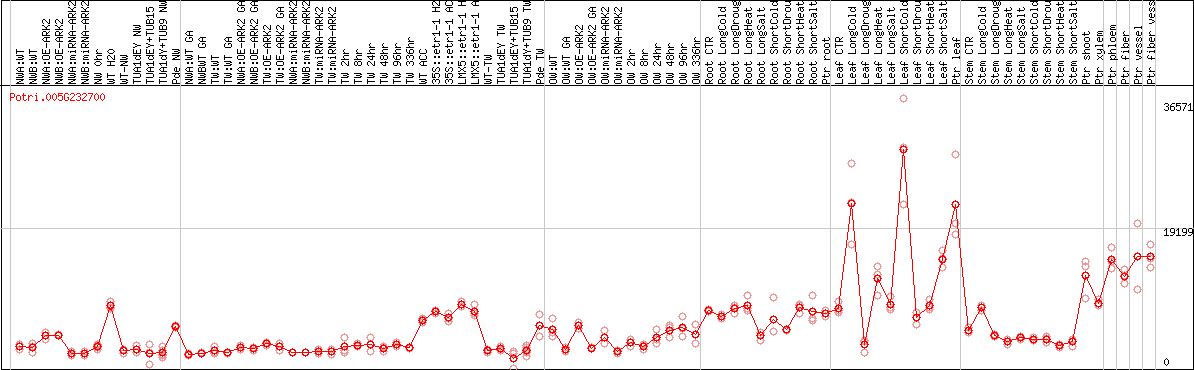
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G232700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.