External link
Symbol
Pt-COL6.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G75540 185 / 2e-56
CO
LHUS, AtBBX21, STH2
long hypocotyl under shade, B-box domain protein 21, B BOX 21, salt tolerance homolog2 (.1)
AT4G39070 166 / 5e-50
CO
B-box zinc finger family protein (.1)
AT1G06040 138 / 5e-40
CO
BBX24, STO
SALT TOLERANCE, B-box domain protein 24, B-box zinc finger family protein (.1.2)
AT1G78600 137 / 4e-38
CO
BBX22, DBB3, STH3, LZF1
SALT TOLERANCE HOMOLOG 3, DOUBLE B-BOX 3, B-box domain protein 22, light-regulated zinc finger protein 1 (.1.2)
AT2G31380 129 / 6e-36
CO
STH
salt tolerance homologue (.1)
AT4G10240 115 / 2e-31
CO
B-box zinc finger family protein (.1)
AT4G38960 100 / 1e-25
CO
B-box type zinc finger family protein (.1.2.3)
AT2G21320 99 / 5e-25
CO
B-box zinc finger family protein (.1)
AT2G24790 81 / 6e-18
CO
COL3, ATCOL3
CONSTANS-like 3 (.1.2)
AT5G15850 81 / 3e-17
CO
COL1, ATCOL1
CONSTANS-like 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G028200
436 / 4e-155
AT1G75540 252 / 2e-82
long hypocotyl under shade, B-box domain protein 21, B BOX 21, salt tolerance homolog2 (.1)
Potri.004G161000
189 / 1e-59
AT4G39070 192 / 6e-62
B-box zinc finger family protein (.1)
Potri.009G122000
183 / 5e-57
AT4G39070 202 / 1e-65
B-box zinc finger family protein (.1)
Potri.007G130100
138 / 2e-39
AT1G06040 264 / 1e-89
SALT TOLERANCE, B-box domain protein 24, B-box zinc finger family protein (.1.2)
Potri.017G028301
137 / 4e-39
AT1G06040 274 / 3e-93
SALT TOLERANCE, B-box domain protein 24, B-box zinc finger family protein (.1.2)
Potri.017G028300
137 / 4e-39
AT2G31380 273 / 4e-93
salt tolerance homologue (.1)
Potri.011G105400
139 / 5e-39
AT1G78600 284 / 1e-95
SALT TOLERANCE HOMOLOG 3, DOUBLE B-BOX 3, B-box domain protein 22, light-regulated zinc finger protein 1 (.1.2)
Potri.001G384000
137 / 2e-38
AT1G78600 284 / 1e-95
SALT TOLERANCE HOMOLOG 3, DOUBLE B-BOX 3, B-box domain protein 22, light-regulated zinc finger protein 1 (.1.2)
Potri.007G015200
97 / 2e-24
AT4G38960 197 / 5e-65
B-box type zinc finger family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028791
182 / 3e-56
AT1G75540 206 / 9e-66
long hypocotyl under shade, B-box domain protein 21, B BOX 21, salt tolerance homolog2 (.1)
Lus10017495
181 / 7e-56
AT1G75540 202 / 2e-64
long hypocotyl under shade, B-box domain protein 21, B BOX 21, salt tolerance homolog2 (.1)
Lus10042498
139 / 9e-39
AT1G78600 274 / 1e-91
SALT TOLERANCE HOMOLOG 3, DOUBLE B-BOX 3, B-box domain protein 22, light-regulated zinc finger protein 1 (.1.2)
Lus10038147
138 / 1e-38
AT1G78600 281 / 2e-94
SALT TOLERANCE HOMOLOG 3, DOUBLE B-BOX 3, B-box domain protein 22, light-regulated zinc finger protein 1 (.1.2)
Lus10025579
127 / 4e-35
AT1G06040 297 / 2e-102
SALT TOLERANCE, B-box domain protein 24, B-box zinc finger family protein (.1.2)
Lus10027041
127 / 5e-35
AT1G06040 298 / 1e-102
SALT TOLERANCE, B-box domain protein 24, B-box zinc finger family protein (.1.2)
Lus10033184
112 / 3e-29
AT1G75540 122 / 7e-33
long hypocotyl under shade, B-box domain protein 21, B BOX 21, salt tolerance homolog2 (.1)
Lus10031087
97 / 1e-23
AT4G38960 229 / 5e-77
B-box type zinc finger family protein (.1.2.3)
Lus10035472
96 / 1e-23
AT4G38960 229 / 4e-77
B-box type zinc finger family protein (.1.2.3)
Lus10018076
93 / 1e-22
AT4G38960 215 / 6e-72
B-box type zinc finger family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00643
zf-B_box
B-box zinc finger
Representative CDS sequence
>Potri.005G234500.1 pacid=42803421 polypeptide=Potri.005G234500.1.p locus=Potri.005G234500 ID=Potri.005G234500.1.v4.1 annot-version=v4.1
ATGAAGATCCAGTGTGACGTTTGTAGCAAAGAAGAGGCCTCGGTGTTTTGCACCGCTGACGAGGCAGCCCTTTGTGACACTTGTGACCACCGTGTCCATC
ATGCCAACAAGCTTGCTTCAAAGCATCAGCGTTTTTCCCTTCTCCATCCTTCCTCAAAAAACTTCCCCATCTGTGATATCTGCCAGGACAAACGGGCTTT
CTTGTTCTGTCAACAAGACAGGGCGATTTTATGCAGAGATTGTGACGGTCCAATACACACAGCAAACGAGCATACCCAGAAGCATAATAGGTTTCTTCTC
ACAGGAGTCAAGCTCTCTGCTACATCTGCTGTTTATATGTCTTCCTCTTCCTCCGTTACCAGTAGTGGCGATCTTGTTCCTGACTCCAAGTCTCAAAAAC
AGCAGCAGCAACAATTAATCAAGAAACCTGTGTCTGTTGCTCCAGTGAATTCCAATCCACCTGCAGTTCCCAGTACCTTGTCAGCAAACACAGTAATAAA
CAAGGACGGGGATAATTTGGTAACAAGTGAAGGGTTTGGTTCGACAACAAGTAGTACTATATCAGAGTACTTGATGGAGACTCTTCCTGGCTGGCATGTT
GAAGAATTTCTTGATTCCTCTTCCACCACTCCATTTGGTTTCAGTAAGATTGATGATGGTCTCTTGCCGTATATGGACACTCATGATCTTGAACGCAATA
TGAGTTCTTTCTCGTCAGAAAGTTTGGGGCTTTGGGTCCCCCAAGCACCAACTCCTCCTCTATGCACATCTCAACAATATTACTATCCACAATTGGTAGG
GCAGAGTGGATTCAAGGAGACAAAAGAGAGCACAAACATGAAAGCTAACAGAAGGTTAACAGATGATGCCTTCACTGTTCCACAGATAAGCCCTCCATCA
AATATTGGCTCCAAGAGATCTAGGCCCTTGTGGTAG
AA sequence
>Potri.005G234500.1 pacid=42803421 polypeptide=Potri.005G234500.1.p locus=Potri.005G234500 ID=Potri.005G234500.1.v4.1 annot-version=v4.1
MKIQCDVCSKEEASVFCTADEAALCDTCDHRVHHANKLASKHQRFSLLHPSSKNFPICDICQDKRAFLFCQQDRAILCRDCDGPIHTANEHTQKHNRFLL
TGVKLSATSAVYMSSSSSVTSSGDLVPDSKSQKQQQQQLIKKPVSVAPVNSNPPAVPSTLSANTVINKDGDNLVTSEGFGSTTSSTISEYLMETLPGWHV
EEFLDSSSTTPFGFSKIDDGLLPYMDTHDLERNMSSFSSESLGLWVPQAPTPPLCTSQQYYYPQLVGQSGFKETKESTNMKANRRLTDDAFTVPQISPPS
NIGSKRSRPLW
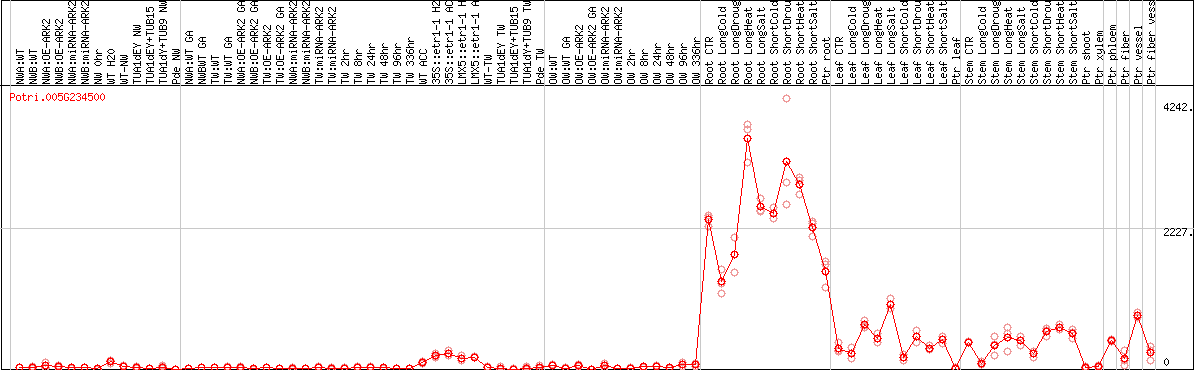
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G234500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.