Potri.006G001400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G001400.1 pacid=42769429 polypeptide=Potri.006G001400.1.p locus=Potri.006G001400 ID=Potri.006G001400.1.v4.1 annot-version=v4.1
ATGGCTTCACCTGGTATTCCACATCAACAAGGAGGTGGGGGGGTAATAGGAGGAGGAGGAGGAGGTTTTGATATGAATAAACTGCTTAAACCCTCTTCTG
CAAATCCTATGAGTATAATGCAACCCCCAAACCAACAACAACAACAGGGTCCTTCTCCCAATAGTAACACTATCACCACCGAGAATCTCACTACCTCTCC
TTCCTTCCCTCTCCCTTCCTCTTCTCCACCTTATCTAACTCCATCTTCTTCTTATCCACCTCCTACTGGCCCTTATCACCCTTTTCACCACCCTCACTAT
CTCTCCCCTTACCCTCCTCCTCCTCCTCCTTTTCAGCAACTTCACAATCAGTTCCTTACTAATACTAATATCCATCACCAAAATAGACCACAGCCCATCT
CTTCTTTTGCACCTCCTCCTCCTCTCTCACCTAGTAATTCAGGTGGTGCTGTTTTGATGGACATCTTGACCAATCAAAACCAACAGCAGCCACCTCTCTC
CAGTAATCTTAGTGGGCCTTTCCCTTCTTATGCTTCCTCCGCTGTATCCACCGCCACCAGTGCTCCTCCTGTCCCCTCTGCCCCTCCTGTTAGTTTAGCA
TCTCCGACGCAGCAATGCTGCCCTCCTCCAGTGAGGATGCTCAGCACCAAGCTCCCTAAGGGCCGGCATTTGAATGGAAACCATGTCGTGTATGATATTG
ATGTGAGATTGCAAGGTGAGGTGCAGCCTCAACTTGAAGTCACTCCCATCACTAAATATGTCTCCGATCCTGGCCTTGTTCTCGGCCGTCAGATTGCTGT
TAATAGGAATTACATTTGCTATGGGCTCAAGCCTGGTGCTATTCGTATCCTCAATATTAATACTGCTTTGAGATCATTGCTTCGCGGTCATAACCAGAAA
GTAACAGACATGGCTTTCTTCGCCGAGGATGTTCACCTTCTGGCCAGTGCATGTGTTGATGGATGTGTTTTCATACGGAAGATCAATGAGGGCCCAGATG
AGGAAGAAAAACCTCAAATTTTTGAAAGGATACTTCTTGCCCTTCACATAATAGCAGATGGAGAACTGGTCCACCCACGAGTGTGCTGGCATCCACACAA
ACAAGAAATTTTGGTCGTTGCAATTGGAAACCTCATCCTGAAAATCGATACCAATAAAGTTGGAAAAGGCGCGGGATTTTCAGCTGAGCAACCTCTCGCT
TGTCCTGTTGATAAGCTGATTGAAGGTGTCCAGCTTGTTGGTAAGCACGATGGCGAAGTCATTGAGTTGTCAATGTGTCAATGGATGACCACCCGTTTAG
CTTCAGCTTCAACAGATGGCGTGGTCAAGATTTGGGAAGACTGTAAGGCAGTACCACTTGCAGTATTCAGACCACATGATGGTAATCCTGTAAATTCTGT
GGCGTTCCTGACTGCCCCTGACCATCCTGATCACATTGTTCTTATCACTGGGGGCCCGCTGAATCAGGAACTGAAGATATGGGCCTCTGCAAGTGAGGAG
GGCTGGTTGTTGCCTAGTAATGCTGAATCATGGCAGTGCAATCAGACTTTGACCTTGAAGAGTTCGGTTGAATCGAACGCTGAGGATGCATTCTTTGATC
AAGTGGTAGCGCTGCCCTGTGCAGGCTTATTTCTGCTGGCAAATGCCAAGAAGAATGCCATATATGCTGTACACTTGGAATATGGGCCTTATCCTGCTGC
AACCCGCATGGATTACATAGCTGAGTTTACAGTTACAATGCCTATTTTGAGTCTTACTGGAACTAGTGATAGTTTACCAAATGGAGAACATATTGTTCAG
GTTTACTGTGTGCAAACACAAGCTATTCAACAATATGCGTTGAACTTGTCTCAATGCCTTCCACCACCTCTGGAGAACATGGAGTTGGAGAGAACAGAAT
CTAATGTTTCTCATGCTTTCGATGCTTCTAATTCTGATGGTTCCACTATTATGGAATCTTCTCATGGAAGTAAACCCACTTATATGTCTGCAGGTAACAT
CGCTTCCATACCACCAATGACTTCAAACAGTTCTGAAAATGCTCCTGCAGCAAATCATCCAGAAAGCTTATGTTCTTCTGATGTTAATAGCTCACTAGAT
ATAGCTTCTTCTGGTGGGCAAACCAAGGCTACTGCATCACATAATAATGCTGACAATACCAACACTGTGCCACCCCTTCTACCGATGAGTCCAAGGTTGC
CTCGAAAGTTGTCAGGTCTCCAGAGTCTGTCAAATAGCACAGATACTAGTTTGCAGCTCAGTGATCATGCTGGTGACCAGTCTGTTCCAGATTATTTGGT
GGACCGCAGAATTGAAACTGTGAAGGAAAATGCAAGTGATACATCCTCTGGTGATAATTTGAGTAAGGGTGAGAAGAATGTCAAACAAACTGATATTGCA
ATGGTTTCTGAAACTCCAATAATGTTTAAACACCCAACCCATCTTATAACTCCATCGGAGATATTATCCAGGGCCGTTTCTTCTGAAAATTCCCAAACCA
CTCAGGGCCTGAATGTCACAGAGGCAAAGATTCAGGATGTGCTTGTGAACAATGATATTGAAAGTGCTGAAGTGGAGCTTAAAGTTGTTGGAGAGACAGG
AACTGATCAAAATAATGACTTTGACTTGCCCAGAGAATCTCACACTGCTGTTGCAGAGAAGAAAGAGAAATCCTTCTACTCTCAGGCCTCAGATCTCGGC
ATTCAGATGGCAAGAGATTGTTGTGTGGAAGCATATAGTGTTGGGCCAGTCCAACAAGTTGATGAAGGTAGTATTACTGAGGTACTTGATAGACCTCCTA
GTGATGAAGATGAGAAGCAAGATATGACTAAAGATGTGCCTGCAAAACGTGGTGAACCAGAAACTTCTGTGGAAGTTCCCCAGCCCCCAGCCCCTACAAC
TAAAGCAAAAAAACCAAAAGGGAAAAGTTCGCAAGTATCCGTTCAATCTTCCCCTTCCCCGAGTCCATTCAACTCTACTGATTCATCCAAGGAACCAGGT
TGCAGTCCCTGTGCCCAATCTAGTGATGCTGCTTTACCTCAGATATTGGACATGCAGGATACACTTGATCAGTTAATGAACATGCAGAAAGAAATGCAGA
AGCAGATGAATACAATGATTTCTGTCCTGGTTTCCAAAGAAGGCAAAAGACTAGAGGCATCATTGGGCCGGAGTATTGAGAAAGTTGTTAGGGCTAACAC
AGATGCTCTATGGGTTCGTTTTCAAGAAGAAAACACTAAGCTTGAGAAATTAGAAAGGGACCGAATACAGCAGTTAGCAAATTTGATTACAAATTTTATA
AACAAGGATTTGCCAACAGCTTTGGAGAAAACTTTAAAGAAGGAAATAGCTGCAATTGGACCTGCTGTTGCTCGTGCTATCACTCCCATTTTGGAGAAAA
GCATATCATCATCTATTATGGAGTCATTCCAGAAAGGGGTGGGAGAGAAGGCAGTGAATCAGCTAGAGAAGACAGTTAGTTCAAAGCTTGAAGTTACAGT
GGCCAGGCAAATCCAATCACAGTTTCAAACTTCTGGCAAGCAAGCTCTTCAGGATGCATTAAGGTCTACTCTAGAAGCCTCAATTATTCCTGCGTTTGAG
ATGTCCTGCAAAGCTATGTTTGATCAAGTTGATGCCACATTTCAGAAAGAGCTTAGTAAGCATATAAATGACACTCAGCAACAGTTCAATTCCATGCATT
CTCCTTTGGCCATCGCTCTGAGGGATGCTATCAATTCAGCATCATCACTCACTCAAACCTTAAGTGGAGAGCTGGCTGATGGCCAACGACAGCTCTTAGC
TATGGCAGCTGCAGGTGCAAACTCAGAAGTAGGAAATCCATCAGCAAAGTTGGGCAATGGACCATTGCCTGGTTTGCATGAGATGCCCGAGGCACCTTTG
GATCCAACAAAAGAATTATCAAGGTTAATAGCTGAGCGGAAGTATGAGGAAGCATTCACTGTAGCTCTGCACAGAAATGATGTCACAATAGTTTCCTGGT
TATGTTCTCAGGTTGATCTACAAGGGATATTGTCCATGTCACCGCTACCACCTCTAAGCCAAGGAGTTCTGCTGGCCCTTCTACAGCAATTGGCATGTGA
TATTAGTAACGAAACCTCTAGAAAGTTGGGATGGATGACAGATGTGGCTGCTGCTATAAACCCTGTGGATCCAATGATTGCAGTGCATGTAAGACCAATC
TTTGAACAGGTCTACCAGATAGTTATCAATCAAAGGAGTCTACCTAGTACATCTGCTTCAGAAGCCCCAGGCATACGCCTTCTTCTGGTTGTCATAAATT
CTGTGCTAAGGAGCTGTAAATGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G001400.1 pacid=42769429 polypeptide=Potri.006G001400.1.p locus=Potri.006G001400 ID=Potri.006G001400.1.v4.1 annot-version=v4.1
MASPGIPHQQGGGGVIGGGGGGFDMNKLLKPSSANPMSIMQPPNQQQQQGPSPNSNTITTENLTTSPSFPLPSSSPPYLTPSSSYPPPTGPYHPFHHPHY
LSPYPPPPPPFQQLHNQFLTNTNIHHQNRPQPISSFAPPPPLSPSNSGGAVLMDILTNQNQQQPPLSSNLSGPFPSYASSAVSTATSAPPVPSAPPVSLA
SPTQQCCPPPVRMLSTKLPKGRHLNGNHVVYDIDVRLQGEVQPQLEVTPITKYVSDPGLVLGRQIAVNRNYICYGLKPGAIRILNINTALRSLLRGHNQK
VTDMAFFAEDVHLLASACVDGCVFIRKINEGPDEEEKPQIFERILLALHIIADGELVHPRVCWHPHKQEILVVAIGNLILKIDTNKVGKGAGFSAEQPLA
CPVDKLIEGVQLVGKHDGEVIELSMCQWMTTRLASASTDGVVKIWEDCKAVPLAVFRPHDGNPVNSVAFLTAPDHPDHIVLITGGPLNQELKIWASASEE
GWLLPSNAESWQCNQTLTLKSSVESNAEDAFFDQVVALPCAGLFLLANAKKNAIYAVHLEYGPYPAATRMDYIAEFTVTMPILSLTGTSDSLPNGEHIVQ
VYCVQTQAIQQYALNLSQCLPPPLENMELERTESNVSHAFDASNSDGSTIMESSHGSKPTYMSAGNIASIPPMTSNSSENAPAANHPESLCSSDVNSSLD
IASSGGQTKATASHNNADNTNTVPPLLPMSPRLPRKLSGLQSLSNSTDTSLQLSDHAGDQSVPDYLVDRRIETVKENASDTSSGDNLSKGEKNVKQTDIA
MVSETPIMFKHPTHLITPSEILSRAVSSENSQTTQGLNVTEAKIQDVLVNNDIESAEVELKVVGETGTDQNNDFDLPRESHTAVAEKKEKSFYSQASDLG
IQMARDCCVEAYSVGPVQQVDEGSITEVLDRPPSDEDEKQDMTKDVPAKRGEPETSVEVPQPPAPTTKAKKPKGKSSQVSVQSSPSPSPFNSTDSSKEPG
CSPCAQSSDAALPQILDMQDTLDQLMNMQKEMQKQMNTMISVLVSKEGKRLEASLGRSIEKVVRANTDALWVRFQEENTKLEKLERDRIQQLANLITNFI
NKDLPTALEKTLKKEIAAIGPAVARAITPILEKSISSSIMESFQKGVGEKAVNQLEKTVSSKLEVTVARQIQSQFQTSGKQALQDALRSTLEASIIPAFE
MSCKAMFDQVDATFQKELSKHINDTQQQFNSMHSPLAIALRDAINSASSLTQTLSGELADGQRQLLAMAAAGANSEVGNPSAKLGNGPLPGLHEMPEAPL
DPTKELSRLIAERKYEEAFTVALHRNDVTIVSWLCSQVDLQGILSMSPLPPLSQGVLLALLQQLACDISNETSRKLGWMTDVAAAINPVDPMIAVHVRPI
FEQVYQIVINQRSLPSTSASEAPGIRLLLVVINSVLRSCK
|
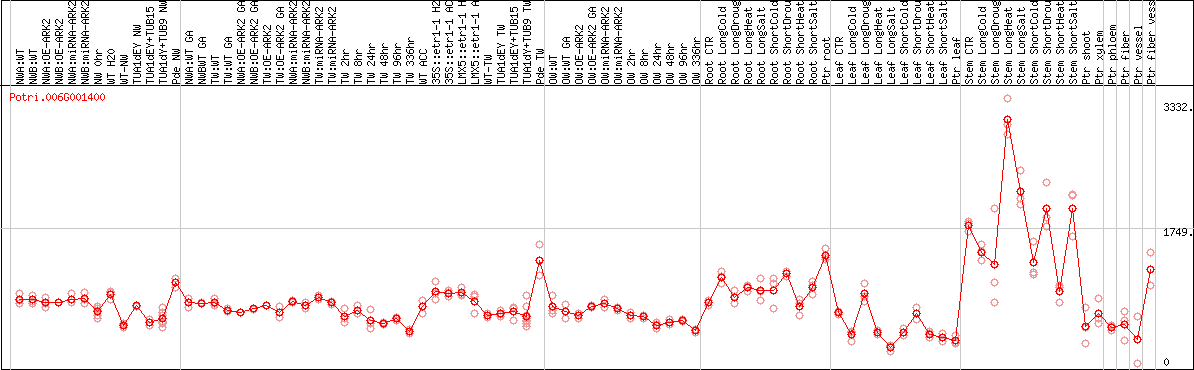
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G001400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
