External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12800 427 / 3e-152
SDRB, DECR
short-chain dehydrogenase-reductase B (.1)
AT2G07640 189 / 4e-60
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G05530 100 / 6e-25
SDRA, IBR1
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
AT3G05260 97 / 3e-23
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G54870 96 / 9e-23
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29260 91 / 5e-21
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G52340 87 / 1e-19
SIS4, SDR1, ISI4, GIN1, ATABA2, SRE1, ABA2, ATSDR1
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G29150 87 / 1e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G07440 87 / 1e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT4G03140 87 / 2e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G125900
430 / 4e-153
AT3G12800 421 / 9e-150
short-chain dehydrogenase-reductase B (.1)
Potri.006G089700
103 / 5e-26
AT5G06060 320 / 6e-111
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G200100
99 / 3e-24
AT3G26770 257 / 2e-85
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.011G022500
96 / 3e-23
AT4G05530 373 / 5e-132
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Potri.011G022400
96 / 4e-23
AT4G05530 330 / 4e-115
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Potri.005G039300
94 / 2e-22
AT2G29150 356 / 7e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G073700
94 / 3e-22
AT1G52340 242 / 1e-79
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G199900
94 / 4e-22
AT2G47140 256 / 1e-85
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.005G039500
92 / 9e-22
AT2G29150 351 / 5e-123
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031746
447 / 1e-159
AT3G12800 463 / 4e-166
short-chain dehydrogenase-reductase B (.1)
Lus10031170
439 / 9e-157
AT3G12800 454 / 8e-163
short-chain dehydrogenase-reductase B (.1)
Lus10018323
386 / 8e-136
AT3G12800 395 / 4e-139
short-chain dehydrogenase-reductase B (.1)
Lus10017129
382 / 6e-134
AT3G12800 391 / 7e-138
short-chain dehydrogenase-reductase B (.1)
Lus10020019
102 / 2e-25
AT4G05530 395 / 5e-141
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Lus10019777
96 / 1e-22
AT1G52340 391 / 3e-138
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016997
95 / 1e-22
AT1G52340 249 / 2e-82
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10015542
94 / 3e-22
AT4G05530 385 / 2e-136
SHORT-CHAIN DEHYDROGENASE/REDUCTASE A, indole-3-butyric acid response 1 (.1)
Lus10021314
94 / 3e-22
AT3G51680 238 / 6e-78
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10029937
94 / 4e-22
AT1G54870 503 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF08659
KR
KR domain
Representative CDS sequence
>Potri.006G025400.6 pacid=42768831 polypeptide=Potri.006G025400.6.p locus=Potri.006G025400 ID=Potri.006G025400.6.v4.1 annot-version=v4.1
ATGGAATCACCATTTAAAGCAAATATCCTGAAAGGAAAGGTAGCTTTGATAACAGGAGGTGGATCAGGAATTGGGTTTGAAATATCAACACAATTTGGTA
AACATGGTGCCTCTGTTGCCATCATGGGTAGGCGCAAGCAAGTTGTTGATTCTGCTGTTGCTAATCTGCAATCACTTGGGATTTCTGCTGCTGGATTTGA
GGGGGATGTTCGCAAGCAGGAAGATGCAAAAAGAGTTCTTGAATCAGCATTTAAACATTTTGGCAAGATTGATATTCTGGTGAATGGTGCAGCTGGCAAC
TTTCTTGTATCTCCTGAGGATTTGTCTCCTAATGGATTTCGAACAGTTTTGGATATTGATGCTGTTGGCACCTTCACAATGTGCCATGAAGCACTTCCTT
ATCTCAAGAAAGGAGGGCTTGGACAAAGCTTATCTGGAGGTGGAATAATTTTAAATATAAGTGCTACCTTACACTATACAGCAGCTTGGTATCAAATAAA
CGTAGCTGCAGCGAAGGCAGCTGTTGACGCCATAGGTAGAAACTTGGCACTGGAGTGGGGTACTGACTATGATATTAGGGTGAATGGGATCGCTCCGGGC
CCCATTAGTGGCACTCCTGGCATGAGTAAGCTTGTTCCGGAAGAGATAAATAGCAAAGCTAAAGATTTCATGCCTCTGTATAAACTAGGAGAGAAATGGG
ACATTGCTATGGCCGCTCTCTACCTTGCTTCTGATGCTGGTAAATACATCAATGGAACCACTCTCATTGTTGACGGGGGACTTTGGCTGAGCCGTCCTCG
CCACCTTCCAAAAGATGAAGTCAAGCAGGTTTCCCGAGCTGTCGAAAAGAAATCCCGAAATGCACCAGCTGGGGTTCCATCTAGCAAGCTCTAG
AA sequence
>Potri.006G025400.6 pacid=42768831 polypeptide=Potri.006G025400.6.p locus=Potri.006G025400 ID=Potri.006G025400.6.v4.1 annot-version=v4.1
MESPFKANILKGKVALITGGGSGIGFEISTQFGKHGASVAIMGRRKQVVDSAVANLQSLGISAAGFEGDVRKQEDAKRVLESAFKHFGKIDILVNGAAGN
FLVSPEDLSPNGFRTVLDIDAVGTFTMCHEALPYLKKGGLGQSLSGGGIILNISATLHYTAAWYQINVAAAKAAVDAIGRNLALEWGTDYDIRVNGIAPG
PISGTPGMSKLVPEEINSKAKDFMPLYKLGEKWDIAMAALYLASDAGKYINGTTLIVDGGLWLSRPRHLPKDEVKQVSRAVEKKSRNAPAGVPSSKL
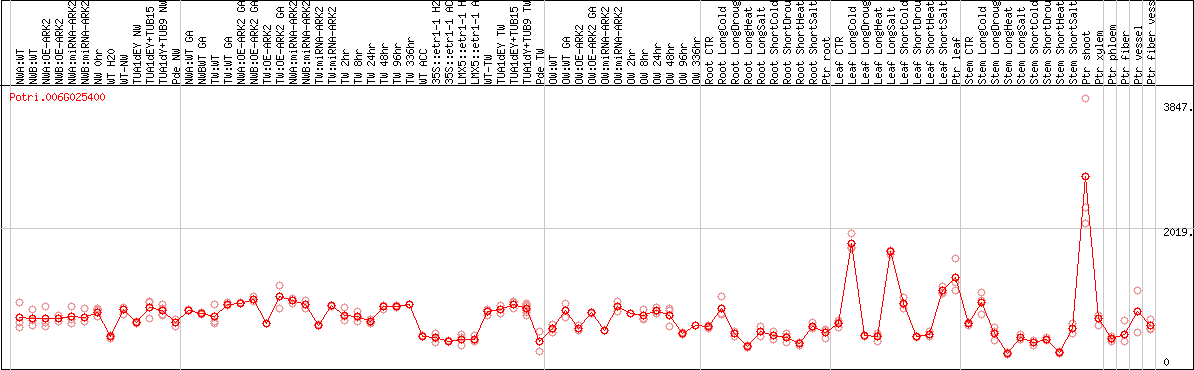
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G025400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.