Potri.006G031900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G031900.1 pacid=42769376 polypeptide=Potri.006G031900.1.p locus=Potri.006G031900 ID=Potri.006G031900.1.v4.1 annot-version=v4.1
ATGGGAAAAAACGTGGTTTCTGACGAAGAAGATGAGGTGGAGTTAGAAGACGAAGATGAGAGAGAGCCAGTTGATGGAGATAAAATTGGACGAGACGACG
ACGAGGAGGAGGATGAAGAAGGGCAGGATGAATATGAGAAGGATGATTTTATAGTGGATGATGTCGAGGAGGAAGAAGAAGTGGAAGAAGAAGAGGATAG
GGCAGATAGCGACGAGGAGAGGCACAAGAAGAAGAAGAAAAAGAAGAAAAGAGAAGCTGAGGATGTACTCGATGAAGATGACTACGAGCTCCTGCGAGAC
AACAATGTGTACCACCATCGGCCAAAGGATAGCAAAAAATTTAAGCGTTTGAAGAAAGCCCAGAGGGATTCAGATGAGGACCGATATGGGCTTTCTGATG
ACGAGTTTGATGGAAGTGGTAAAGGTGGTCGAACTGCTGAGGAGAAACTGAAGCGCAGCTTATTTGGTGATGATGAAGGGGTTCCACTTGAGGATATGCC
TGAAGAGGAGGAGCAAGAAGAAGTGGAAGAGGATGCGGACATTGGTGACGAGGATGAGATGGCTGATTTTATTGTGGATGAAGATGATGATGATGGAACT
CTAGTGAGACGCAAGAAGCTAAAGAAAAAGAAATCTCAGCAGGCATCTGGAGTTTCATCCTCTGCCTTGCAAGAAGCTCAAGAAATATTTGGTGATGTTG
ATGAACTCATACAGATACGCAGACAAGGGTTAGAATCTAGTGAATGGAGAGAAAGGAGACTGGAAGATGAATTTGAGCCAACTGTCCTTTCTGAAAAATA
CATGACTGAGAAAGATGATCAGATCCGGATGACTGATATTCCAGAAAGAATGCAGGTATCTGAGGGAAGCACTGGTCCTCCTCCATTGGATGATTTCAGC
ATAATGGAGGAGAGCAACTGGATTTATAGTCAGATTGCAAGTGGAACATTACCTTTGTTTGCGGAGAGTGGACTGCTTATAAACAAGGATGATGTCACCC
GATTTTTGGAATTACATCACATTCAGAAGTTAGATATTCCCTTCATTGCTATGTATCGGAAGGAAGAGTGTTTAAGCCTGTTGAAAGATCCCGAGCAACA
TGAAGATGATGAAAATCCTTATGACACTGGTAGAATTCCCACATTCAAGTGGCACAAGGTGCTTTGGGCCATCCAGGACTTGGACAGGAAGTGGTTACTT
CTTCAGAAACGAAAGAGTGCCCTCAATGCTTACTACAATAAGCGCTTTGAAGAAGAATCTCGACGGATATATGATGAAACAAGACTCAATTTAAACCAGC
AGCTGTTTGAGTCAATTCTTAAATCACTCAAGACTGCAGAATCGGAAAGAGAGGTTGATGATGTTGATGCAAAATTTAACCTGCATTTTCCTCCTGGTGA
GGTTGTCGTGGATGAAGGACAATACAAGAGGCCCATGCGGAGATCACAATACAGTGTCTGTAGTAAAGCTGGCCTATGGGAAGTTGCAAGCAAATTTGGG
TACAGTGCCGAGCAACTGGGAATGCAGTTATCTTTATTAAAGATGGAGGACGAGTTGCAGGATGCCAAGGAAACACCAGAGGAGATGGCTTCAAATTTTA
CATGTGCAATGTTTGAAAGTCCCCAAACAGTTCTTAAAGGGGCTAGGCACATGGCTGCAGTTGAGATAAGTTGTGAGCCCTGTGTCAGGAGATATGTCCG
TTTAATTTTTATGGACAAGGCTGTAGTGTCTACGAGTCCAACATCAGATGGGAAAGCTGCTATAGATTCTTTCCATCAGTTTGCAGGTATCAAGTGGTTG
CGTGAAAAGCCAGTCAAGAAGTTTGAGGATGCACAGTGGCTCCTTATCCAGAAAGCTGAAGAGGAGAAACTTCTTCAAGTCACTATTAAGCTTCCACAAA
AAGTTATGGATCAACTGATTGATGACTGTAACGGTCGCTACCTGAGTATTGGTGTTAGTAAATATGCTCAACTGTGGAATGAGCAAAGGAGTTTGATACT
GAAGGATGCACTCTTTGCTTTTTTGTTGCCTTCAATGGAGAAAGAGGCTAGGTCATTGTTGACAAGTAGAGCTAAAAACCGGTTGCTTTGGGAATATGGG
AAGGTTTTTTGGAATAAAGTCTCTGTGGGCCCATATCAACGGAAAGAAAGTGATATTTCCATGGATGATGAAGCTGCACCTAGGGTTATGGCTTGCTGTT
GGGGTCCTGGAAAGCCAGCAACTACTTTTGTGATGCTAGATTCATCTGGGGAGGTTTTAGATGTACTTTATGCTGGGTCCCTTACCTTACGATCTCAGCA
TGCTAGTGATCAGCAACGCAAGAAAAATGATCAGCAACGTGTTCTGAAATTTATGACAGACCACCAGCCACACGTGGTTGTTCTTGGAGCTGTGCACTTG
TCTTGTACTAAACTGAAGGATGACATTTACGAGATTATTTTTAAAATGGTGGAGGAAAATCCTAGAGATGTTGGACATGAGATGGATGAACTGAGTATTG
TGTATGGGGACGAGTCTCTCCCTCGTCTTTATGAAAATTCTCGGATTTCTTCTGATCAACTACCTGGGCAGTCAGGCATTGTTAAACGGGCTGTTGCTCT
TGGCCGTTATCTACAAAATCCCTTGGCTATGGTTGCAACGCTATGTGGACCAGCTAGGGAGATTCTATCTTGGAAACTAAATCCTTTGGAGAACTTTCTC
ACTCCTGATGATAAATACATGGTGATTGAACAAGTCATGGTAGATGCCACAAACCAGGTGGGCTTAGACATTAATTTGGCCACAAGCCATGAATGGTTAT
TTGCTCCTTTACAATTCATTTCTGGGCTTGGACCCAGAAAGGCAGCATCTTTGCAGAGGTCTCTGGTAAGAACTGGAGCGATTTTCACTCGCAAGGATTT
TGTGACAGCTCATGGTCTTGGTAAAAAAGTGTTTGTCAATGCTGTTGGATTTTTGCGTGTCCGGAGGAGTGGCCTTGCTGCTAGCAGCAGCCAGTTCATT
GACGTGTTGGATGATACAAGGATACATCCAGAGTCCTATGGTCTCGCACAGGAGTTGGCAAAAGTCATTTATGAAAAGGACAGTGGTGATGTAAATGATG
ATGATGATGCACTAGAGATGGCAATAGAGCATGTTAAAGAGCGGCCTAATTTATTGAAAACATTTGTTTTCGATAAGTATCTCGAAGACAAGAAGCGGGA
AAACAAGAAGGAGACTTTCATGGATATAAGAAGGGAATTAATACAAGGTTTCCAAGATTGGCGTAAACAGTATAAAGAACCAACCCAGGATGAGGAATTT
TATATGATTTCAGGTGAAACCGAGGATACCCTTGCTGAAGGGATAATTGTACAAGCCACAGTTCGCAGGGTGCAAGGTGGAAAGGCAATATGTGCATTGG
AATCTGGTCTGACTGGGATTCTTACGAAGGAGGACTACGCTGATGATTGGAGGGATATTCCTGAGTTGTCCGACAAGCTGCGTGAAGATGATATACTGAC
TTGCAAGATAAAATCAATTCAGAAGAATAGATATCAAGTGTTCCTTGTTTGCAAAGACAGTGAGATGAGAAATAACCGATATCAGCAGGCCCGAAACCTT
GATCGCTATTATCATGAAGACCAGAGCAGTCTGAGGAGTGAGCAGGAGAAGGTTCGCAAAGACAGGGAGCTTGCAAAGAAACATTTCAAGCCTAGGATGA
TTGTTCATCCACGCTTCCAGAATATAACGGCTGATGAAGCAATGGAGTTCTTATCTGACAAGGATCCTGGAGAAAGTATTATTCGTCCTAGTTCTCGTGG
GCCGTCGTATTTAACTCTAACTCTTAAAGTTTACAACGGAGTTTATGCTCACAAAGACATAGTTGAAGGTGGGAAGGAACACAAAGATATCACGAGTGTA
CTTCGTATTGGGAAGACATTGAAAATTGGGGAGGACACATTTGAGGATTTGGATGAGGTAATGGACCGCTATGTTGATCCCTTGGTGAGCTACCTAAAGG
CAATGCTAAGTTATCGCAAGTTCAGAAGTGGCACCAAAGTAGAAGTTGATGAGCTCCTTAGGATTGAGAAATCACAACAGCCTACTAGGATAGTTTATGC
ATTTGGCATTTGTCATGAACACCCAGGCACATTCATTCTCACTTATATAAGGAGTACAAATCCCCATCACGAGTATGTTGGTCTATATCCCAAGGGATTC
AAGTTCCGAAAGCGTATGTTTGAGGACATTGATCGGCTGGTGGCATATTTTCAAAAGCATATAGATGACTCCTTGCATGAATCAGCACCATCAATTAGGT
CAGTTGCTGCTATGGTCCCAATGCGAAGCCCTGCAACTGGGGGCTCGTCCTGGGGTGGTTCAACCTATGAAGGTGGTCGGAGAGGGCAGTCTTTTGACAG
AGATCGTTCTTCTGGTCCAGGTTCCAGGACAGGTCGAAGTGGTGGTAGTCGAGATGGTCACCAGAGTGGGGCACCAAGACCATACAGTGGACGAGGACGT
GGTCGTGGCTCATATAACAATGGCGGGGGAAGTAATAGTGGCAATGAGAGGCGGGATTCTGGTTATGATAAGCCCAGATGGGATTCAGGAACCAAGGATG
GTGATGAAGGCTGGGGCAGTTTCCCAGGAGCCAAAGTCCAAAACTCACCTGGCAGGGAAGCTTTTCCTGGTGGCTGGGGAGCTGGTGCAAGCAGTGGTGG
AAATGGTTGGGGTGGTGGAGCTAGTGGTGGTGATAATAGTGGTTGGGGCCATGGAACTGATGGTGGGACCGATAGTGGGAATTCTGGGCGGGGTACTACT
AGCTCGAAGAGAGATTCAGCACAAAGGGGGTGGGGTGCTGCTGGAGGTGGAAATGGCAATGGTAGTGACTCTGGTCATGGAAGTCGTGGTGGAGGCACTA
ACGATGAGAATTCTGAGTGGGGGCAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G031900.1 pacid=42769376 polypeptide=Potri.006G031900.1.p locus=Potri.006G031900 ID=Potri.006G031900.1.v4.1 annot-version=v4.1
MGKNVVSDEEDEVELEDEDEREPVDGDKIGRDDDEEEDEEGQDEYEKDDFIVDDVEEEEEVEEEEDRADSDEERHKKKKKKKKREAEDVLDEDDYELLRD
NNVYHHRPKDSKKFKRLKKAQRDSDEDRYGLSDDEFDGSGKGGRTAEEKLKRSLFGDDEGVPLEDMPEEEEQEEVEEDADIGDEDEMADFIVDEDDDDGT
LVRRKKLKKKKSQQASGVSSSALQEAQEIFGDVDELIQIRRQGLESSEWRERRLEDEFEPTVLSEKYMTEKDDQIRMTDIPERMQVSEGSTGPPPLDDFS
IMEESNWIYSQIASGTLPLFAESGLLINKDDVTRFLELHHIQKLDIPFIAMYRKEECLSLLKDPEQHEDDENPYDTGRIPTFKWHKVLWAIQDLDRKWLL
LQKRKSALNAYYNKRFEEESRRIYDETRLNLNQQLFESILKSLKTAESEREVDDVDAKFNLHFPPGEVVVDEGQYKRPMRRSQYSVCSKAGLWEVASKFG
YSAEQLGMQLSLLKMEDELQDAKETPEEMASNFTCAMFESPQTVLKGARHMAAVEISCEPCVRRYVRLIFMDKAVVSTSPTSDGKAAIDSFHQFAGIKWL
REKPVKKFEDAQWLLIQKAEEEKLLQVTIKLPQKVMDQLIDDCNGRYLSIGVSKYAQLWNEQRSLILKDALFAFLLPSMEKEARSLLTSRAKNRLLWEYG
KVFWNKVSVGPYQRKESDISMDDEAAPRVMACCWGPGKPATTFVMLDSSGEVLDVLYAGSLTLRSQHASDQQRKKNDQQRVLKFMTDHQPHVVVLGAVHL
SCTKLKDDIYEIIFKMVEENPRDVGHEMDELSIVYGDESLPRLYENSRISSDQLPGQSGIVKRAVALGRYLQNPLAMVATLCGPAREILSWKLNPLENFL
TPDDKYMVIEQVMVDATNQVGLDINLATSHEWLFAPLQFISGLGPRKAASLQRSLVRTGAIFTRKDFVTAHGLGKKVFVNAVGFLRVRRSGLAASSSQFI
DVLDDTRIHPESYGLAQELAKVIYEKDSGDVNDDDDALEMAIEHVKERPNLLKTFVFDKYLEDKKRENKKETFMDIRRELIQGFQDWRKQYKEPTQDEEF
YMISGETEDTLAEGIIVQATVRRVQGGKAICALESGLTGILTKEDYADDWRDIPELSDKLREDDILTCKIKSIQKNRYQVFLVCKDSEMRNNRYQQARNL
DRYYHEDQSSLRSEQEKVRKDRELAKKHFKPRMIVHPRFQNITADEAMEFLSDKDPGESIIRPSSRGPSYLTLTLKVYNGVYAHKDIVEGGKEHKDITSV
LRIGKTLKIGEDTFEDLDEVMDRYVDPLVSYLKAMLSYRKFRSGTKVEVDELLRIEKSQQPTRIVYAFGICHEHPGTFILTYIRSTNPHHEYVGLYPKGF
KFRKRMFEDIDRLVAYFQKHIDDSLHESAPSIRSVAAMVPMRSPATGGSSWGGSTYEGGRRGQSFDRDRSSGPGSRTGRSGGSRDGHQSGAPRPYSGRGR
GRGSYNNGGGSNSGNERRDSGYDKPRWDSGTKDGDEGWGSFPGAKVQNSPGREAFPGGWGAGASSGGNGWGGGASGGDNSGWGHGTDGGTDSGNSGRGTT
SSKRDSAQRGWGAAGGGNGNGSDSGHGSRGGGTNDENSEWGQ
|
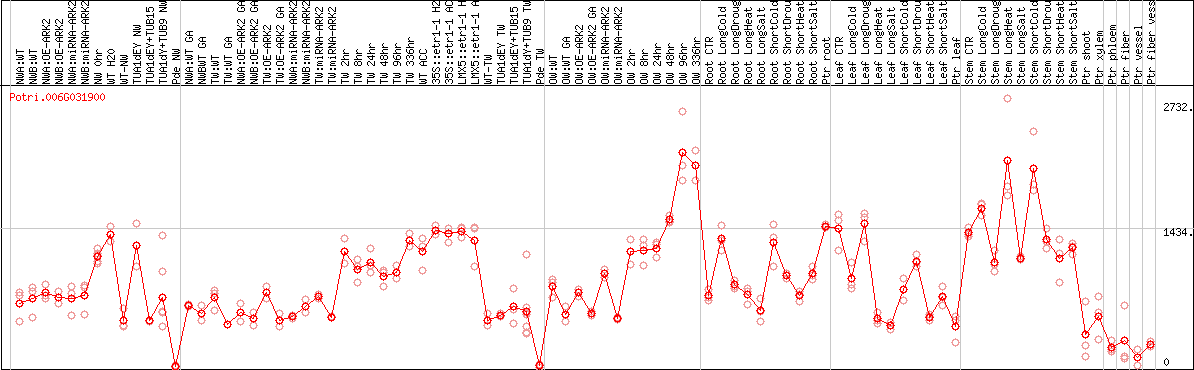
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G031900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
