Pt-GLU1.2 (Potri.006G038400) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-GLU1.2 | ||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G038400.4 pacid=42769065 polypeptide=Potri.006G038400.4.p locus=Potri.006G038400 ID=Potri.006G038400.4.v4.1 annot-version=v4.1
ATGGCTTCTCTTCAATCACCATTAATCTCTCCGGTTCCGCAACTCGTCAATGCAACAACACCGAACTCAGTTAATAAGAACTTATTATTTGTTGATTTTG
TTGGACTCTACTGTAAATCGAAGAGAACTAGAAGGAAAATCGGAGTTTCTTCTTCTTTTTCCTCGAGTTTCTCTCGATTTGCTAACAAAAAGAAGTCCTC
CTGTCCAGTCAACGCCACTCTCAGTGTTGACCGTCGTAATATCTCACCGCCGTCTTCTCCTCCTCATCCTCCTCCTGATTTGAAACCGCAGGTAGCAAAC
TTGGAAGATATATTATCAGAAAGAGGAGCATGTGGAGTTGGATTTATCGCGAATTTGGAAAACAAGCCGTCCCATGCGATTGTCAAGGATGCACTTACAG
CTCTTGGCTGCATGGAACATCGTGGAGGTTGTGGAGCAGATAATGACTCTGGTGATGGTTCAGGACTAATGACTTCGATTCCGTGGGAGCTATTCGATAA
GTGGGCTGAGAGTGAAGGGATTGGTTCTTTTGATAAATTGCATACTGGTGTCGGAATGATTTTTTTCCCTAAAGATGATAACCTTATGAAAGAAGCAAAA
GAAGTGATTGTAAATATTTTTAAACAAGAGGGCCTTGAGGTGCTCGGATGGAGGCCTGTTCCTGTAAATACATCTGTAGTTGGTTTTTACGCAAAAGAAA
CCATGCCCAACATAGAACAAGTATTTGTTAGAGTTATTAACGAAGAGGATGTTGATGACATTGAACGAGAACTGTATATCTGCCGGAAGCTGATTGAAAG
AGCGGCAAACTCAGAAAGTTGGGGAAATGAGCTTTATTTCTGTTCTTTGTCTAATCGAACAATTGTTTATAAGGGAATGCTTCGTTCTGAAGTTCTTAGG
CTGTTTTATTCTGACCTTCAGAATGATATCTACAAATCACCTTTTGCCATTTATCATAGAAGGTATAGCACAAATACTAGTCCCAGATGGCCTCTTGCTC
AACCCATGCGGTTTCTTGGTCATAATGGAGAGATCAACACCATACAGGGTAACTTGAACTGGATGCAATCACGAGAAACCTCATTGAAATCTTCTGTTTG
GCATGGTCGAGAAAATGAAATCCGTCCTTATGGGAACCCAAAGGCATCAGACTCAGCTAATCTTGACAGTGCAGCAGAATTGTTAATAAGAAGTGGGCGC
ACTCCTGAGCACGCGTTGATGGTTCTCGTACCAGAGGCATACAAGAATCATCCAACTTTGACCATCAAATATCCTGAGGTTGTAGATTTTTATGATTATT
ACAAGGGTCAGATGGAAGCTTGGGATGGACCTGCTCTACTCTTATTCAGTGATGGAAAGACAGTTGGGGCTTGTCTTGATCGGAATGGCCTTCGCCCTGC
TAGATATTGGCGGACAGTCGACAATTTTGTCTATGTTGCATCTGAGGTTGGGGTTGTACCAATGGATGAATCAAAAGTAACGATGAAAGGCCGTCTAGGT
CCCGGAATGATGATAACTGTAGATTTGCCTGGTGGCCAGGTTTATGAGAATACTGAAGTTAAAAAGCGAGTAGCACTGTCAAATCCGTATGGAAAATGGG
TTCATGAAAACCTGCGATCCTTGAAGTCTACAAACTTCCTCTCAGCAACAGTTATGGACAATGAATCAATTTTAAGATGCCAACAGGCATTCGGCTACTC
CAGTGAGGATGTCCAAATGGTTATTGAGAATATGGCTTCTCAAGGGAAGGAGCCTACATTTTGCATGGGGGATGATATTCCTTTGGCAATATTGTCTCAA
AAACCACACATGCTTTATGATTATTTCAAACAACGGTTTGCACAGGTTACAAATCCAGCTATAGACCCACTCAGAGAAGGATTAGTTATGTCACTTGAAA
TCAACATCGGAAAGCGAGGAAACATCTTGGAAGATGGACCAGAAAATGCTTCACAGGTTATCTTGTCTAGTCCTGTACTAAATGAAGGGGAGCTTGAGTT
ATTACTGAAGGATCCCTACTTAAAACCTCAAGTTTTACCAACCTTTTTTGATATACGTAAAGGGGTTGAGGGTTCTTTGGAGAAAACACTGATTAAGCTT
TGTGCGGCAGCTGATGAAGCTGTTAGAAATGGTTCCCAATTGCTTGTTCTCTCTGACCGATCTGATGATCTGGAACCCACTCGACCTGCAATCCCAATTC
TTCTTGCTGTTGGTGCTGTTCATCAGCATCTTATTCAGAATGGCCTGCGAATGTCAACTTCAATTGTAGCAGACACTGCTCAGTGCTTCAGCACTCATCA
TTTTGCCTGTTTGATTGGATATGGTGCCAGTGCTATATGCCCGTACTTGGCACTGGAAACTTGCCGGCAATGGCGCTTGAGTAAAAGAACTGTGAATCTG
ATGATGAATGGAAAGATGCCTACTGTAACAATCGAACAAGCTCAAAAGAACTTTTGCAAGGCAGTTAAATCTGGTCTTCTCAAAATTCTTTCTAAGATGG
GCATCTCATTGCTTTCAAGTTATTGTGGTGCACAGATATTTGAGATTTATGGATTGGGGAAGGAGGTTGTTGATCTAGCATTTTGTGGCAGTGTTTCAAA
TATTGGTGGAGTGACTTTTGATGAGTTGGCGAGGGAGACTTTGTCCTTCTGGGTGAAAGCTTTCTCTGAGGCTACGGCCAAAAGACTGGAAAATTATGGA
TTTATACAATTCAGACCTGGAGGGGAATATCATGGAAACAATCCAGAGATGTCGAAATTGCTTCACAAAGCTGTTCGCCAAAAGAGTGAAAATGCATTTT
CTATTTATCAGCAACATTTGGCTAACCGGCCTGTGAATGTTCTTCGTGATCTTCTTGAATTTAAAAGTGATCGTGCTCCAATTCCTGTGGGAAAGGTTGA
ACCTGCCATATCCATAGTTCAAAGGTTTTGCACTGGGGGGATGTCACTTGGAGCCATTTCAAGAGAAACTCATGAAGCAATTGCCATTGCTATGAACAGA
TTAGGTGGAAAATCTAATTCAGGGGAAGGTGGTGAGGATCCAATTCGCTGGACCCCACTTTCTGATGTTGTTGATGGGTACTCCCCTACATTGCCGCATC
TTAAAGGTCTTCAAAATGGTGATACTGCTACCAGCGCTATCAAGCAGGTTGCTTCAGGACGTTTTGGTGTCACTCCAACATTCTTGGTCAACGCTGTTCA
GTTAGAAATCAAAATCGCACAGGGTGCCAAGCCTGGTGAAGGTGGCCAATTGCCTGGGAAGAAAGTTAGTGCATATATTGCGAGGCTGAGAAATTCTAAA
CCTGGGGTTCCACTTATATCCCCACCCCCTCACCATGACATTTATTCCATTGAAGATCTTGCCCAATTGATCTATGACCTTCATCAGGTCAATCCTAAGG
CTAAGGTATCTGTAAAGCTTGTGGCAGAAGCAGGTATTGGCACAGTTGCATCTGGGGTAGCAAAGGGTAATGCTGATGTTATACAGATATCAGGGCATGA
TGGTGGAACTGGAGCTAGCCCCATAAGTTCCATTAAGCATGCCGGTGGTCCTTGGGAACTTGGACTTACAGAAACCCATCAGACTCTCGTTGCAAATGGG
CTTAGAGAAAGAGTGATCCTTAGGGTTGATGGGGGCTTCAAAAGCGGAGTTGATGTCCTGATGGCTGCGGCAATGGGTGCTGACGAGTATGGATTTGGTT
CTGTCGCAATGATTGCTACGGGATGTGTAATGGCCCGCATTTGTCACACTAATAATTGCCCTGTTGGTGTTGCTAGTCAGAGAGAAGAACTACGCGCCCG
GTTTCCTGGTGTGCCTGGTGATCTTGTTAATTTCTTTTTATATGTTGCTGAGGAGGTAAGAGGCATGCTGGCACAACTGGGTTATCAGAAGTTGGATGAT
ATCATTGGGCATACAGATTTGTTAAGACAACGAGATATTTCCCTTGTCAAAACTCAGCATCTTGATCTCAGTTATATTATGTCGAGTGTAGGATTACCAA
AACTGAGCAGTACTGATATCAGGAACCAAGATGTACACTCTAATGGTCCTGTTTTAGATGATGTTGTTCTAGCAGATCCAGAGATTCTAGATGCTATTGA
GAATGAAAAAGTTGTCAATAAAACCATAAAAATATACAATGTGGACCGTGCTGTTTGTGGGCGTATAGCAGGTGTGGTTGCAAAGAAATATGGTGACACT
GGTTTTGCTGGGCAGCTGAACATCACGTTTACTGGGAGTGCTGGGCAGTCGTTTGCATGTTTTTTGACACCTGGAATGAATATTCGCCTAATAGGAGAGG
CCAATGATTACGTGGGAAAGGGTATGGCTGGAGGAGAGTTGGTCGTAACTCCCGTGGAGAACACTGGGTTTGTTCCGGAGGACGCCACCATTGTCGGGAA
TACTTGCTTGTATGGGGCTACAGGCGGTCAAGTATTTGTCAGAGGGAAAGCTGGGGAGCGGTTTGCTGTGAGAAACTCACTTGCTGAAGCTGTAGTTGAA
GGCACTGGAGACCATTGTTGTGAGTACATGACTGGTGGTTGTGTGGTGGTACTTGGAAAAGTGGGTAGAAATGTAGCTGCTGGGATGACCGGAGGTTTGG
CGTACATGCTTGATGAGGATGACACTTTGATGCCCAAGGTAAATAAAGAAATAGTGAAGGTCCAAAGAGTGACTGCTCCTGTTGGCCAGATGCAGCTAAA
GAGCCTTATTGAAGCTCATGTTGAAAAAACCGGAAGCGGCAAAGGCGCTGCTATCCTGAAGGAGTGGGACACATATCTACCGCTCTTTTGGCAGCTAGTA
CCACCCAGTGAAGAAGACACCCCAGAGGCTTGTGCATCATTCGAGGCAACCAGTGCTGGGCAAGTGACATCATTCCAATCTGCATAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G038400.4 pacid=42769065 polypeptide=Potri.006G038400.4.p locus=Potri.006G038400 ID=Potri.006G038400.4.v4.1 annot-version=v4.1
MASLQSPLISPVPQLVNATTPNSVNKNLLFVDFVGLYCKSKRTRRKIGVSSSFSSSFSRFANKKKSSCPVNATLSVDRRNISPPSSPPHPPPDLKPQVAN
LEDILSERGACGVGFIANLENKPSHAIVKDALTALGCMEHRGGCGADNDSGDGSGLMTSIPWELFDKWAESEGIGSFDKLHTGVGMIFFPKDDNLMKEAK
EVIVNIFKQEGLEVLGWRPVPVNTSVVGFYAKETMPNIEQVFVRVINEEDVDDIERELYICRKLIERAANSESWGNELYFCSLSNRTIVYKGMLRSEVLR
LFYSDLQNDIYKSPFAIYHRRYSTNTSPRWPLAQPMRFLGHNGEINTIQGNLNWMQSRETSLKSSVWHGRENEIRPYGNPKASDSANLDSAAELLIRSGR
TPEHALMVLVPEAYKNHPTLTIKYPEVVDFYDYYKGQMEAWDGPALLLFSDGKTVGACLDRNGLRPARYWRTVDNFVYVASEVGVVPMDESKVTMKGRLG
PGMMITVDLPGGQVYENTEVKKRVALSNPYGKWVHENLRSLKSTNFLSATVMDNESILRCQQAFGYSSEDVQMVIENMASQGKEPTFCMGDDIPLAILSQ
KPHMLYDYFKQRFAQVTNPAIDPLREGLVMSLEINIGKRGNILEDGPENASQVILSSPVLNEGELELLLKDPYLKPQVLPTFFDIRKGVEGSLEKTLIKL
CAAADEAVRNGSQLLVLSDRSDDLEPTRPAIPILLAVGAVHQHLIQNGLRMSTSIVADTAQCFSTHHFACLIGYGASAICPYLALETCRQWRLSKRTVNL
MMNGKMPTVTIEQAQKNFCKAVKSGLLKILSKMGISLLSSYCGAQIFEIYGLGKEVVDLAFCGSVSNIGGVTFDELARETLSFWVKAFSEATAKRLENYG
FIQFRPGGEYHGNNPEMSKLLHKAVRQKSENAFSIYQQHLANRPVNVLRDLLEFKSDRAPIPVGKVEPAISIVQRFCTGGMSLGAISRETHEAIAIAMNR
LGGKSNSGEGGEDPIRWTPLSDVVDGYSPTLPHLKGLQNGDTATSAIKQVASGRFGVTPTFLVNAVQLEIKIAQGAKPGEGGQLPGKKVSAYIARLRNSK
PGVPLISPPPHHDIYSIEDLAQLIYDLHQVNPKAKVSVKLVAEAGIGTVASGVAKGNADVIQISGHDGGTGASPISSIKHAGGPWELGLTETHQTLVANG
LRERVILRVDGGFKSGVDVLMAAAMGADEYGFGSVAMIATGCVMARICHTNNCPVGVASQREELRARFPGVPGDLVNFFLYVAEEVRGMLAQLGYQKLDD
IIGHTDLLRQRDISLVKTQHLDLSYIMSSVGLPKLSSTDIRNQDVHSNGPVLDDVVLADPEILDAIENEKVVNKTIKIYNVDRAVCGRIAGVVAKKYGDT
GFAGQLNITFTGSAGQSFACFLTPGMNIRLIGEANDYVGKGMAGGELVVTPVENTGFVPEDATIVGNTCLYGATGGQVFVRGKAGERFAVRNSLAEAVVE
GTGDHCCEYMTGGCVVVLGKVGRNVAAGMTGGLAYMLDEDDTLMPKVNKEIVKVQRVTAPVGQMQLKSLIEAHVEKTGSGKGAAILKEWDTYLPLFWQLV
PPSEEDTPEACASFEATSAGQVTSFQSA
|
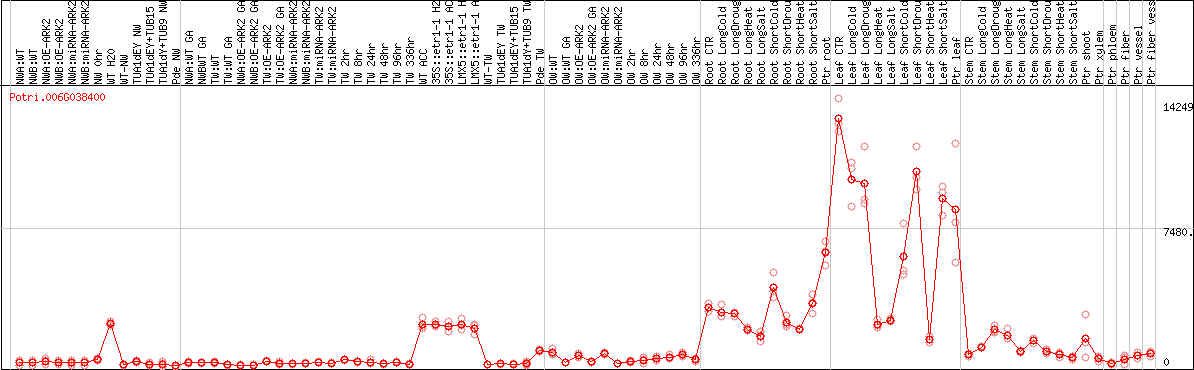
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G038400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
