Potri.006G039400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G039400.1 pacid=42767916 polypeptide=Potri.006G039400.1.p locus=Potri.006G039400 ID=Potri.006G039400.1.v4.1 annot-version=v4.1
ATGGCGCGTCTTCTTCTAAAACCTCACAACAGCAGTAGACACTGCATCTCTTTACTCAGAGCTACCATGACCTCCTCCTCCTCTTCTTCTGTTACTTGGT
CTCACTCATTCTCTTCTCTCTCCCGCCCCTTAGTCTCCTTCAAGCCAAGGGTGAGTTTCCCATTGAGGACGGTTCCAAGTAGATTTTTGGTGCAAAATGG
GGTCTCACCAAGAGCTTTAATGTCTACCAGTGCAGCAACTGAATCACAGAAAGAAAGCGCGAGCTCGAAAGCTTATGGGTCTGATCAAATTCAGGTGTTG
AAAGGATTAGAGCCTGTAAGGAAAAGGCCAGGGATGTATATTGGAAGTACGGGACCGCGTGGTCTGCACCATTTGGTGTATGAGATATTAGATAATGCTG
TTGATGAGGCCCAAGCAGGATATGCTTCAAAGATAAATGTTGTTCTGCATTCGGATAATTCCATTAGCATTACTGACAATGGGCGTGGGATTCCCACTGA
CTTGCATCAGGTGACAAAGAAATCTGCCCTGGAGACTGTGTTAACGGTCTTACATGCTGGCGGCAAATTTGGTGGTTCTAACAGTGGATATAGTGTCTCT
GGTGGGCTACATGGTGTGGGTTTATCTGTTGTCAATGCTTTGTCTGAGGAACTAGAAGTTACAGTTTGGCGTGATGGAAAGGAGTACCAGCAGAAATATT
CTCGTGGAAACCCTGTGACAACTCTAATGTGTTATGAACTTTCAGCAGAATCGAAAGACCAAAAAGGGACTCGCATCAGATTCTGGCCTGACAAAGAAGT
ATTCACCACTGAGATTCAGTTTGACTACAACACAGTAGGTGGACGGGTCAGGGAGCTTGCATTTTTGAACCCAAAGCTTGCTATTTCATTGAAAAAAGAG
GACAATGATCCGGAGAAGAATCAGTATGATGAACATTTCTATGCAGGTGGATTGATTGAATATGTGAATTGGCTAAATACTGATAAGAAATCGCTTCATG
ATGTTGTGAGTTTCAGAAAAGAAGTAGATGGAATTGCAATTGATATGGCTCTCCAATGGTGTTCTGATGCTTACTCAGATACAATCCTGGGATATGCTAA
TAGCATACGTACTGTTGATGGTGGAACCCATATAGATGGGGTGAAGGCTTCATTGACAAGAACTCTCAATAACTTTGGGAAAAAGTCAAAAGTCATTAAG
GATAAGGATATTAGTTTAAGCGGTGAACATGTAAGGGAGGGATTGACATGCATTATCTCAGTCAAAGTTCCAAGCCCAGAATTTGAAGGTCAGACAAAGA
CAAGGTTGGGAAATCCGGAGGTGCGGAAAGTGGTAGAGCAATCTGTCCAGGAGTATCTTACTGAATATTTGGAGTTGCATCCTGATGTTCTTGATTCAAT
CTTATCAAAGTCTCTAAATGCTCTCAAGGCAGCTTTGGCAGCAAAGAGGGCAAGAGAATTGGTGAGACAAAAAAGTGTTTTGAAATCATCATCTCTTCCA
GGAAAACTGGCTGATTGCTCATCTACAGATCCTGAAGAATCTGAAATTTTTATAGTTGAAGGAGATTCAGCTGGAGGGAGTGCCAAACAAGGCCGTGATC
GACGCTTTCAGGCTATTCTCCCTTTGAGGGGTAAAATTCTGAATATAGAAAGAAGAGATGAGGAGGCAATGTACAAAAATGAAGAGATCCAAAATCTTAT
ACTTGCCCTTGGACTTGGAGTGAAGGGAGAGGATTTTAAAAAAGATGCTCTTCGTTATCATAGGATCATCATCTTAACAGATGCTGATGTGGACGGTGCT
CACATCCGAACTCTTCTGCTGACTTTCTTCTTTAGATATCAGAGAGCTTTATTTGATGAAGGCTGCATTTATGTTGGTGTGCCACCCCTATATAAGGTTG
AGAGGGGAAAGCAGGTTTACTATTGTTTTGATGATGCAGAACTCAGAAAGGTTCAAAGTTCATTCCCTCAAAATGCATCATACAACATGCAAAGGTTCAA
AGGCTTGGGGGAAATGATGCCTGCACAACTGTGGGAAACAACAATGGACCCAGAGCAAAGGCTGCTGAAACAATTAGTGGTTGAAGATGCAGCAGAGGCC
AGTGTTGTCTTTTCATCTCTTATGGGTACCCGGGTGGATGGCCGCAAGGAACTTATTCAGAAAGCGGCAAGAATGATTAACGTTGATCATTTGGACATCT
GA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G039400.1 pacid=42767916 polypeptide=Potri.006G039400.1.p locus=Potri.006G039400 ID=Potri.006G039400.1.v4.1 annot-version=v4.1
MARLLLKPHNSSRHCISLLRATMTSSSSSSVTWSHSFSSLSRPLVSFKPRVSFPLRTVPSRFLVQNGVSPRALMSTSAATESQKESASSKAYGSDQIQVL
KGLEPVRKRPGMYIGSTGPRGLHHLVYEILDNAVDEAQAGYASKINVVLHSDNSISITDNGRGIPTDLHQVTKKSALETVLTVLHAGGKFGGSNSGYSVS
GGLHGVGLSVVNALSEELEVTVWRDGKEYQQKYSRGNPVTTLMCYELSAESKDQKGTRIRFWPDKEVFTTEIQFDYNTVGGRVRELAFLNPKLAISLKKE
DNDPEKNQYDEHFYAGGLIEYVNWLNTDKKSLHDVVSFRKEVDGIAIDMALQWCSDAYSDTILGYANSIRTVDGGTHIDGVKASLTRTLNNFGKKSKVIK
DKDISLSGEHVREGLTCIISVKVPSPEFEGQTKTRLGNPEVRKVVEQSVQEYLTEYLELHPDVLDSILSKSLNALKAALAAKRARELVRQKSVLKSSSLP
GKLADCSSTDPEESEIFIVEGDSAGGSAKQGRDRRFQAILPLRGKILNIERRDEEAMYKNEEIQNLILALGLGVKGEDFKKDALRYHRIIILTDADVDGA
HIRTLLLTFFFRYQRALFDEGCIYVGVPPLYKVERGKQVYYCFDDAELRKVQSSFPQNASYNMQRFKGLGEMMPAQLWETTMDPEQRLLKQLVVEDAAEA
SVVFSSLMGTRVDGRKELIQKAARMINVDHLDI
|
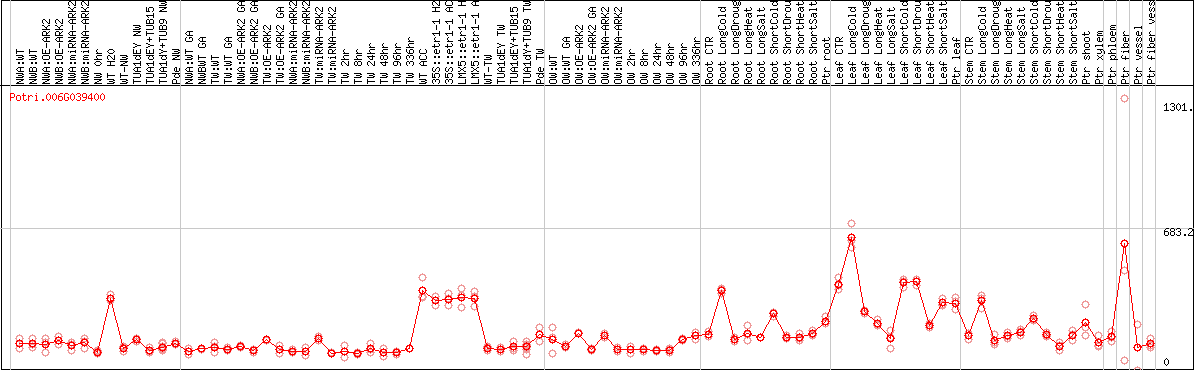
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G039400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
