Potri.006G041500 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G041500.1 pacid=42770089 polypeptide=Potri.006G041500.1.p locus=Potri.006G041500 ID=Potri.006G041500.1.v4.1 annot-version=v4.1
ATGGCTCCTTATTTTGTGTTTCCACAAAATCTAAAATCCTTAGAAGAAGAAAACGAAGACAATCGTCTCCACGTTCTGAACCCAACCGACGTCGCTTCTC
TCCGTCTTCCTGAACTCGAAGAATTTGTCAAAGGAGTATCCTTTGATTTGTCGGATAAAGAGATATTCTGCATTGAGGAGCAAGAAGTGTTTGATCACGT
ATATTCGTTAGTTAAAGGGTTTTCTAGTCTAACTCCATCCGGTAAAGTCAATCTTGTGGAGAGTCTTCGGTCGAATCTAAGTGTTTTGCTCCCAAATGTG
GATTCTCTCTTGCGCGTTTTTCAAGGTCAGGATGATGATAATGACAATGGGAATGAGACTCCTCCGGTGCTTGATCGAGTTTCTTCTTATCGAAATGCTC
TTAAAATTTACACATTTTTCCTCGTCAGTATTGTTCTGTCTGAGGAGTCAAGTGCTAGTTCAAATAACAAAACTAAGATGACAGGGCCTAATAGGAAGAA
ACAGTCGGTACATTCATGGAATTGGGAACCACAGAGGGGCCGGATACTTAATTTGATTGCTAATTCACTAGAGATCAACCTTGCGTTGCTCTTTGGATCG
ACAGACCCTGATGAAAATTATCTTTCTTTTATTACAAAAAATGCATTTGGTTTGTTTGAGAATGCAACACTTATAAAGGACTCTGAGACAAAAGATGCTC
TTTGCCGAATAATTGGGGCTTGTGCTACGAAATACCACTATACAGCACAGTCATGTGCTTCAATCATGCACCTAGTTCACAAATATGACTATGTTGTTAC
CCACATGGCTGATGCGGTTGCTGGGGCTGAGAAGAAGTATGCTGATGGCACCCTTGCCAGTTCTTTGATCAGAGAGGTTGGGAGGACTAATCCAAAAGCT
TATGTGAAAGACACTGTTGGGGCTGAAAATGTTGGGCGTTTTCTTGTAGAGCTTGCTGATCGGCTACCAAAGTTGATTTCAACTAACATTGGGGTCTTGG
TCCCACATTTTGGAGGAGAATCTTATAAGATAAGGAATGCTCTTGTTGCTGTTCTGGGGAAGTTGGTGGCAAAGGCATTTAAGGATGTTGAGGGTGATGT
CAGTTCTAAATCTGTTCGTCTCCGCACAAAGCAAGCCATGTTGGAAATCTTGCTTGAGCGCTGCCGAGATGTTTCTGCTTATACCAGGAGTCGGGTTCTT
CAAGTTTGGGCTGAACTATGTGAAGAGCATTCTGTTTCAATTGGTCTGTGGAATGAGGTTGCTGCAGTTGCTGCTGGGAGATTGGAGGATAAATCTGCAA
TAGTTAGAAAAGCTGCATTAAATTTGCTTATCATGATGCTGCAGCATAACCCTTTTGGCCCCCAGCTTCGAATAGCTTCATTTCAAGCAACATTAGAACA
ATACAATAAGAAGTTGAATGAGCTTGAACCAGATAAAAGTGCAGAAAGTGTTTTGGATGGATTGCAATCTGATAATGAGACCTATGATGGAGGTGAGGTG
GATGATGTGAATATGGAAGAACCAGTTAAGGAACAGCAGGAGAGTTTAACTGACAGTGTGCCTAATCTGGAAGAAGGGATTCCTCATAAGGATAGCTCAG
TGCCAGATATTGGGAATTTGGAGCAAACCAGGGCTTTGGTTGCATCTCTGGAGGCTGGCTTGATATTTTCTAAGTGCGTATCTGCCACAATGCCTACGCT
TGTCCAGTTGATGGCTTCATCTTCAGCCACAGATGTTGAGAACACTATTTTGTTGCTGATGAGGTGCAAACAGTTCCAAATTGATGGTGCCGAGGCTTGC
CTCCGTAAGATGTTGCCATTGGTATTCTCTCAGGATAAATCCATCTATGAAGCTGTGGAGAATGCCTTCATTACAATTTATGTCCGGAAAAATCCATTAG
ATACTGCTAAGAATCTTTTGGATCTTGCCATAGATTCAAATATAGGAGATCTTGCGGCTCTGGAATTTATTGTCAATGCACTGGTTTCCAAAGGTGATAT
CTCCACAAGCACAATATCAGCATTATGGGATTTCTTTTGCTTTAACATTAGTGGTACAACTCCAGAGCAAAGTCGTGGAGCTTTATCAGTGCTTTGCATG
GCTGCAAAAGCGTCCCCTGGAGTTCTTGGCTCACACTTGCAAGATATTATTGATATTGGCTTTGGCCGTTGGGCTAAAGTGGACCCTTTGCTTGCCAGGA
CAGCATGCATTGCCATTCAGAGATTGTCAGAAGAGGACAAGAAAAAGCTGTTGGCTAGTAATGGTAGCCGTGTGTTTGGTTTCTTAGAGAACTTGATTAG
TGGCTCTTGGCTTCCAGAAAACACATGGTATGCTGCTGCTGATAAAGCAATAGGTGTCATATATGCAATTCATCCCACTCCGGAAACCCTAGCTGCTGAC
CTTGTAAAGAAGTCTCTTAGTTCTGTTTTCATTTGTAGTGGAGGAGATGACTTGCAGAATGATATTGAAAGTGGCAGCGCTGATATTCTTACAACAGTTC
AAGTAGCAAAAATTAGTAGATATTTATTTGTCACAAGTCATGTTGCCATGAATCAGTTACTATACATAGAAACCTGTGTTCGGAAGATCCAAAAACAGAA
GCTGAAGAGGGATAAATTGGGAGCTGATGGACAGAATGGTCATAATAATGGTATAAAACAAGATGATACACCAAAGGATAACATCAATGCGGAGCTAGGT
GTTTCTGCCTCTGAAGATGCAATTCTGGATACACTCTCTGAGAGAGCAGAGAAAGAGATTGTTGCTGGTGGTTCCAAAGAGAAATATTTGATAGGGCTTT
GTGCACCTTTCCTGTCGAAGCTTTGTAGAAATTTTAGTTTGATGCAAAAGTATCCTGAACTACAGGCATCTGGAATGCTTGCTCTTTGTCGATTTATGAT
TATTGATCCAGATTTTTGTGATGCAAATCTCCAGCTTCTCTTCACAGTTGTGGAAAGTGCACCATCAGAAACTGTTCGTTCAAACTGCACTATTGCTCTT
GGAGATTTGGCAGTTCGGTTTCCTAATCTGTTAGAACCATGGACTGAAAACATGTATGCTCGGTTACGAGATCCTTCGGTTTCTGTTAGGAAAAATGCTG
TGCTGGTGCTTTCACATCTCATATTAAATGATATGATGAAGGTTAAAGGTTATATAAATGAGATGGCTATAAGATTAGAAGATGAACAGGAGAGGATTTC
AAATCTCGCTAAACTTTTCTTTCATGAGCTGTCAAAGAAAGGAAGCAATCCTATATATAACTTACTTCCAGATATACTTGGCAAATTATCCAACCAGGAG
CTGAAAAGGGAGACTTTCTGCAACATCATGCAGTTTTTAATTGGGTCCATTAAAAAGGACAAACAAATGGAGTCTCTTGTTGAAAAGCTTTGCAACAGGT
TCAGTGGAGTGATAGATACTAGGCAGTGGGAATACATCTCGTATTGCCTCTCCCAGCTGGCATTTACTGAAAAGGGAATGAAAAAACTCATTGATTCATT
TAAGACGTTTGAACATGTCCTGTCTGAGGATTCTGTCATGGACAATTTTAAAAGCATAATCATTAAGGCTAAGAAGTTTGCAAAACCAGAACTTAAATTA
TGCATTGAGGAATTTGAGGAGAAGCTTACTAAGTTCCACATGGAAAAGAAAGAACAGGAAGTGACAGCAAGAAATGCCCAAATTCATCAGCAGAAAATCG
GTGGCATGGAAGGTTGTGCGGTGGCTAGAAATGAAGGAGAAGTGAGTGAAGAATCAGATGTTTTTGAAGATGGTGAAATCGATGATCCATCCATGGAAGA
AATGGTTCAATCATCGGATTCTGAGGTCAACAGGTCAGGTGAATATTCAGGTACTTCAAGTGAGGTGACAGGCATGGAGTCAGATGGCACTGAAGTGCAG
TCAAAAGTGGCCTCCAAATCTAGAGCTACGGAAAGCAAAGTGAAGGGTGAGAGTGGTGATATTTCTGCATCATCTAGAAGATCCACCAGGTCAAAGCAAA
GATAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.006G041500.1 pacid=42770089 polypeptide=Potri.006G041500.1.p locus=Potri.006G041500 ID=Potri.006G041500.1.v4.1 annot-version=v4.1
MAPYFVFPQNLKSLEEENEDNRLHVLNPTDVASLRLPELEEFVKGVSFDLSDKEIFCIEEQEVFDHVYSLVKGFSSLTPSGKVNLVESLRSNLSVLLPNV
DSLLRVFQGQDDDNDNGNETPPVLDRVSSYRNALKIYTFFLVSIVLSEESSASSNNKTKMTGPNRKKQSVHSWNWEPQRGRILNLIANSLEINLALLFGS
TDPDENYLSFITKNAFGLFENATLIKDSETKDALCRIIGACATKYHYTAQSCASIMHLVHKYDYVVTHMADAVAGAEKKYADGTLASSLIREVGRTNPKA
YVKDTVGAENVGRFLVELADRLPKLISTNIGVLVPHFGGESYKIRNALVAVLGKLVAKAFKDVEGDVSSKSVRLRTKQAMLEILLERCRDVSAYTRSRVL
QVWAELCEEHSVSIGLWNEVAAVAAGRLEDKSAIVRKAALNLLIMMLQHNPFGPQLRIASFQATLEQYNKKLNELEPDKSAESVLDGLQSDNETYDGGEV
DDVNMEEPVKEQQESLTDSVPNLEEGIPHKDSSVPDIGNLEQTRALVASLEAGLIFSKCVSATMPTLVQLMASSSATDVENTILLLMRCKQFQIDGAEAC
LRKMLPLVFSQDKSIYEAVENAFITIYVRKNPLDTAKNLLDLAIDSNIGDLAALEFIVNALVSKGDISTSTISALWDFFCFNISGTTPEQSRGALSVLCM
AAKASPGVLGSHLQDIIDIGFGRWAKVDPLLARTACIAIQRLSEEDKKKLLASNGSRVFGFLENLISGSWLPENTWYAAADKAIGVIYAIHPTPETLAAD
LVKKSLSSVFICSGGDDLQNDIESGSADILTTVQVAKISRYLFVTSHVAMNQLLYIETCVRKIQKQKLKRDKLGADGQNGHNNGIKQDDTPKDNINAELG
VSASEDAILDTLSERAEKEIVAGGSKEKYLIGLCAPFLSKLCRNFSLMQKYPELQASGMLALCRFMIIDPDFCDANLQLLFTVVESAPSETVRSNCTIAL
GDLAVRFPNLLEPWTENMYARLRDPSVSVRKNAVLVLSHLILNDMMKVKGYINEMAIRLEDEQERISNLAKLFFHELSKKGSNPIYNLLPDILGKLSNQE
LKRETFCNIMQFLIGSIKKDKQMESLVEKLCNRFSGVIDTRQWEYISYCLSQLAFTEKGMKKLIDSFKTFEHVLSEDSVMDNFKSIIIKAKKFAKPELKL
CIEEFEEKLTKFHMEKKEQEVTARNAQIHQQKIGGMEGCAVARNEGEVSEESDVFEDGEIDDPSMEEMVQSSDSEVNRSGEYSGTSSEVTGMESDGTEVQ
SKVASKSRATESKVKGESGDISASSRRSTRSKQR
|
DESeq2's median of ratios [POPLAR]
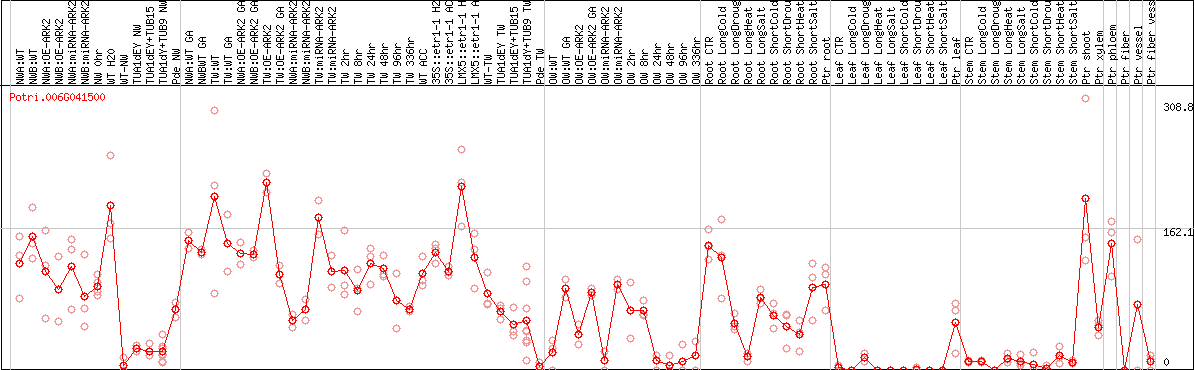
Coexpressed genes
Potri.006G041500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
