Potri.006G043900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G043900.1 pacid=42768336 polypeptide=Potri.006G043900.1.p locus=Potri.006G043900 ID=Potri.006G043900.1.v4.1 annot-version=v4.1
ATGATGCCTACCAAGCTTGGAGCTGAATTTGACTCTGATTATTCCTTCTCATCTGAAAGTTCATGTATGAGCAGCAAATCGTCATCATCCTCATCATGTT
ACCAGAAAGAAGATAATAGCTACGAGGCAGGAAGAGAGCTGAAGAGGAAAGTAAAGAAGTTAAGATCATCCAAGCTTGCAAGATTGCCAAGCTTGAAGCT
CATCACAAGACAAGCCAAAATTCAATCTGATGATGTTTCTGTCCTGTCTAGTGACGCAGCAAGATCACAACATTCAGGCATGTCACCAAATTATATGAGG
ACCACCACTTCTTCAAATGCAAAGAAAGAGAATTTGCAGAAATCAGCAAGAACTTTGGCAAGGCGATCTAGTTTTAAGCCAGCAAAGAGTTTGCCAAGGT
TGTCTAGTATAAAATTCAGGAGACCTTTGATAAGGAAATCTTCTGGAGGAACTGACCTGAAGAAAAAAATGAAGAGGTCAAGATCAATCAAGCTTGCCGA
CCGATCTTCCATTGTTTTTGCAAGTGATGCGGATGCATCGGTGGATTATTTGGAGGCAAAAAACTGTTCTGATGGGAGGAATGTTCATTTTCAGGCAAGT
CCCCATCTTGAATCTACTTTGATTAGCAACAATGAGGATAGGAAAAAATCAAGTAATTCAAAGCAGAATTTAGCTTCTTCTGGTAACAACTCTATGAGAG
TCATGACAAGAACTTCCACATTGAGACCTGTAAGGATCTTCACAAAAGTGGCGAGCTTCAGAACAAAGCGTTCTCAAATTCCAGACTCAAGCATCCAAAA
AACCACCTGTTCTTCGGCCATCAAAGACTCCAAGTTTCCAAATCATCTTGAGCTGCAACCAGGAGGAAGTGAATCTGAAGGAAATTCGATCATGAAGGTT
TGTCCATATAGTTACTGCTCCCTTCATGACCATCGCCATTCCGATGTACCTCCGCTTAAACGCTTTGTATCAATGAGGCGACGTCTGTTAAAGACCCAAA
AAAGCATGAAATCAGAAAGTCAATCCTCTCGAAGAGCGAAGCATTCTGGTATTTCAAAGAAGGGAACACAGACAAGTCAATCGGCATCTTGTGGAGACTT
GGCAGTTCTAGAAACAGCCCATGACAAGATGGCTGTCTCATCGTCCATTGGCAGAAAAGCTGGACAAAGAGCTGAGTCAAAGAGTGCTCACGGAGGAGAT
GAAAAAGATTACAGAGATGTAATAAGTGTCACTGAGAACCAAACTCTTCCAGAAGAAGCTGATGAAGGTAGAATTGCTAGTTTGAACTTGAATGTATTTA
AAGGTGATTCACAATTGAACACTGCAAAGGAAAATGCTTCAACAAGTGTAGCTGATGAACGAGTCAATAAACCCAGAAGCCTAAGCCTGAATAGATTCGT
GGAGTCCACAGAAATCGACAATAAGGTCTCTTCTGTTTCCATTGGTAAACCATCGCAAGAGGAAACAGCATCCTGTGAAGAAAAGAACCAAGATGCTGTG
CAAGATTATCGGTTCCTTGGTGCAGATTCTGAGCATGATTATACTGTGGATACAGGCCACAGAAATCCATGGGAGAAACAGAAGCCCATGGGATTGTGGA
ACCTAATATATCAGCATATGGCATCAGGTGTTGCTGCAGAAGATGGAACCCGACCACATCTTAATAAAGAAGCCAAGGAAGAAGAAGAAGAAGAGAATAC
ATTGCCTGGCATGAGCAAATCTGGTTCTTTTCAGGATTTTTCTAGTACAGATCACAGCATTGGTGAGGAATATCATGATGAACGCAGCCAGAAGATTCAG
CAATACCAATGTGATGCCATTAAGTTGGTGCAAGAAGCATTTGATAGAATACTCTCAGAAATTCCAGACCAGCCAACTGATGATCTATCAGTTACTAGTG
ACACAACTTCAGACAAGAAGATTGCAGAAAACGATCATGGTGAAGACAGACAATTGAACATCTCAACTTCCTATGATTCATGTGGAGACAGCATGGTGCA
GGAACCAGAAGAAACGAGGCTCCAAGCTGATAATGCCTTCCAAAAAGAGAAGGCAGAATCAAGTGTGGAAAGCAAGTCTAACCAGCAAACGCCCAAGAGC
TGGAGTAATCTGAGAAAAATACTCATCCTCAAGAGATTCATCAAGGCATTGGAAAAGGTGAGGAACTTCAGTCCACAGAAGCCACGGAATCTAAATGTAG
AGGCTGACCCAGAAGCAGAAAAGGTTCATCTGCGTCATCAAACCATGGGAGAGAGAAAAAATTCTGAGGAATGGATGCTTGATCATGCACTTCAACAGGT
TATTTCTACACTAGCCCCAGCTCAAAAGAGAAAGGTAGCTCTGCTTGTACAAGCATTTGAGAAAGTCACCCTGCCTACAGAAGTTGGTACTTCGCCAAGG
TCCAACATAGAAGCTTCTTCTCAAACAACTCCAGTTAAAACCAGCACCGGTGCCTCTGACTGCAAAGGGTCTAGAGAAGGAAAAGAAACTATATTTGGAA
TCACTCTTTGTAAGACATCAAGCCTTGAAACAAGCTTCAAACAGAATCAAGACCAGGCCAGTGACTTCTACAAAGTGGATGAGCACATTCAAGGTTCTTG
TTCTGAGGTGAAAGAAACAAGTTTAAAAAATGGTTGCATTCATCTAGCATCGAGTCCTTCGTCCACAAAGAACACTGCTGCAGAGTTGAAGAATGAATTT
GTTGCTTTCAATCTTGGTAATGGTGAAACTAATTCCACTGTTAAAGATGATGAGCCTGATTTCGTCAGCCACTGTTTGGTGGAGGATACAGATTCCAACT
TATGTGACAATCCATTGCCAAGGTCAGCTGATGTTCTACGAACTTCTAGTGAGGAACTTGTGATATATGGGGAAACCCTCCAAGAGGATGCCAAAGAAGC
AAGTGCAGTTTCTGCCTCAGAGGTACATGATCGTAATTTTGGATTAAACGGCCAAAAGTCAGACATCAATAACAAAAACAACGGAACTTGTGATGAATCA
GATGAACCCAAAAGCCAAACCCTAAAAGATTACGAGGGTTCCATTGCAAATACCGATGTGGTTTCTTCATCTTCAGTTTCTGTGCCACTAAAGGAATCAT
CTGAAGTTGCTGGAGAAGAAAATAAGCTCCTCCAAGGATCCACCCTGCTTGACGATTCAGAACCAGGTTGTACAACTGATGCTGCACATGAAAAGCAGAA
ACATATGAAGTTCTGGTTCTTGATATATAAGCACATGGTATCAGGCAATGCGACATTACTTGAGGGAGCAGAGAATGAAGAACAAGGGGATGGTGGCAAC
CAATTGGTTGAAATGAACACATTGGATAATGATGACGCGGGCAACCAAAAGATTAAACTCCAGCAGATTGAAACCATTAGACTGGTAGAAGAAGCAATTG
ATCAAATCCCTCTTCCAGAATTTCAAGAAGACTCACCAGATGATCAATCAGTTGCCTGCGACATAATTCAAGACCAGGACCAGGAGCACACAGAAAAGAA
AGCTGGGGAAGGGGAGGAACCTTTCATCTCATCTTCCTTTGAGGACACCAATGAAAGCTTTGAAAAATCTGACAGCACAAAAGTAGAGGAGAGCACCACA
CTGTATCAACAAGAACAGCAGCTCAATTCTGATAACATCAGTGCACAAGAAAAGGCAAAACCAATACCACCAGCAGGAAACAAGCCAAAGCCAGCTATGC
AAAACTGGAGCAATCTGAAAAAGGTGATCCTCCTCAAGAGATTCGTTAAGGCATTGGAAAAGGTGAAAAAATTCAATCCAAGGGAGCCACGATTTCTGCC
CTTGGACCCTGCAAGTGAAGCTGAAAAGGTTCATTTAAGGCACCAAGACACGGGTGATAGGAAAAATGCTGATGAATGGATGCTTGATTATACACTTCAA
CAGGTTGTTGCTAAACTGACTCCGGCTCGGAAAAGAAAAGTGTCATTGCTTGTAGAAGCCTTTGAGGCAGTCACCCCTATTGGAAGTTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G043900.1 pacid=42768336 polypeptide=Potri.006G043900.1.p locus=Potri.006G043900 ID=Potri.006G043900.1.v4.1 annot-version=v4.1
MMPTKLGAEFDSDYSFSSESSCMSSKSSSSSSCYQKEDNSYEAGRELKRKVKKLRSSKLARLPSLKLITRQAKIQSDDVSVLSSDAARSQHSGMSPNYMR
TTTSSNAKKENLQKSARTLARRSSFKPAKSLPRLSSIKFRRPLIRKSSGGTDLKKKMKRSRSIKLADRSSIVFASDADASVDYLEAKNCSDGRNVHFQAS
PHLESTLISNNEDRKKSSNSKQNLASSGNNSMRVMTRTSTLRPVRIFTKVASFRTKRSQIPDSSIQKTTCSSAIKDSKFPNHLELQPGGSESEGNSIMKV
CPYSYCSLHDHRHSDVPPLKRFVSMRRRLLKTQKSMKSESQSSRRAKHSGISKKGTQTSQSASCGDLAVLETAHDKMAVSSSIGRKAGQRAESKSAHGGD
EKDYRDVISVTENQTLPEEADEGRIASLNLNVFKGDSQLNTAKENASTSVADERVNKPRSLSLNRFVESTEIDNKVSSVSIGKPSQEETASCEEKNQDAV
QDYRFLGADSEHDYTVDTGHRNPWEKQKPMGLWNLIYQHMASGVAAEDGTRPHLNKEAKEEEEEENTLPGMSKSGSFQDFSSTDHSIGEEYHDERSQKIQ
QYQCDAIKLVQEAFDRILSEIPDQPTDDLSVTSDTTSDKKIAENDHGEDRQLNISTSYDSCGDSMVQEPEETRLQADNAFQKEKAESSVESKSNQQTPKS
WSNLRKILILKRFIKALEKVRNFSPQKPRNLNVEADPEAEKVHLRHQTMGERKNSEEWMLDHALQQVISTLAPAQKRKVALLVQAFEKVTLPTEVGTSPR
SNIEASSQTTPVKTSTGASDCKGSREGKETIFGITLCKTSSLETSFKQNQDQASDFYKVDEHIQGSCSEVKETSLKNGCIHLASSPSSTKNTAAELKNEF
VAFNLGNGETNSTVKDDEPDFVSHCLVEDTDSNLCDNPLPRSADVLRTSSEELVIYGETLQEDAKEASAVSASEVHDRNFGLNGQKSDINNKNNGTCDES
DEPKSQTLKDYEGSIANTDVVSSSSVSVPLKESSEVAGEENKLLQGSTLLDDSEPGCTTDAAHEKQKHMKFWFLIYKHMVSGNATLLEGAENEEQGDGGN
QLVEMNTLDNDDAGNQKIKLQQIETIRLVEEAIDQIPLPEFQEDSPDDQSVACDIIQDQDQEHTEKKAGEGEEPFISSSFEDTNESFEKSDSTKVEESTT
LYQQEQQLNSDNISAQEKAKPIPPAGNKPKPAMQNWSNLKKVILLKRFVKALEKVKKFNPREPRFLPLDPASEAEKVHLRHQDTGDRKNADEWMLDYTLQ
QVVAKLTPARKRKVSLLVEAFEAVTPIGS
|
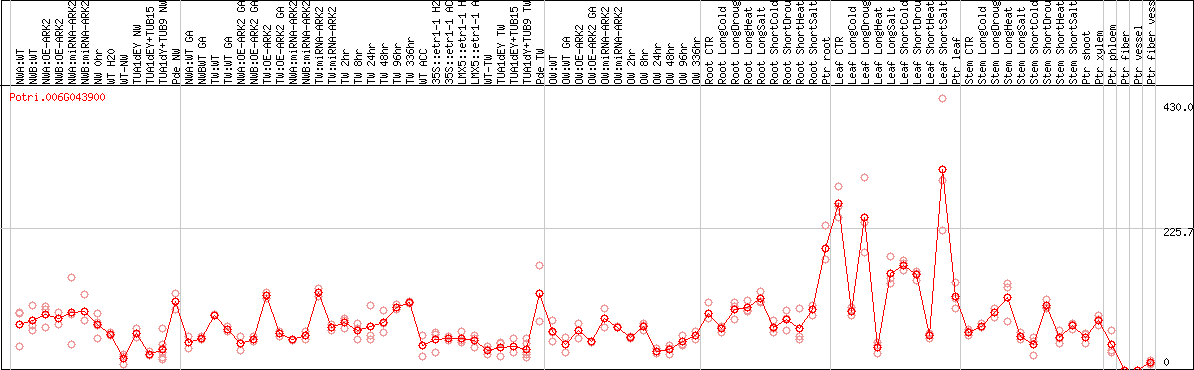
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G043900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
