External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G51510 134 / 1e-39
Y14
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G11400 53 / 1e-08
ATEIF3G1, EIF3G1
eukaryotic translation initiation factor 3G1 (.1.2)
AT5G06000 49 / 5e-07
ATEIF3G2, EIF3G2
ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITIATION FACTOR 3G2, eukaryotic translation initiation factor 3G2 (.1)
AT5G64200 48 / 1e-06
ATSC35, At-SC35
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
AT2G21440 48 / 1e-06
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G27000 47 / 4e-06
ATRBP45C
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G51300 47 / 4e-06
splicing factor-related (.1.2.3)
AT3G50670 46 / 6e-06
U1SNRNP, U1-70K
U1 small nuclear ribonucleoprotein-70K (.1.2)
AT1G73530 45 / 8e-06
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G47120 45 / 1e-05
C3HZnF
RNA recognition motif (RRM)-containing protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009963
133 / 7e-39
AT1G51510 265 / 3e-90
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10003632
130 / 2e-37
AT1G51510 260 / 3e-88
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030482
55 / 4e-09
AT3G11400 459 / 9e-165
eukaryotic translation initiation factor 3G1 (.1.2)
Lus10028823
55 / 6e-09
AT3G11400 459 / 9e-165
eukaryotic translation initiation factor 3G1 (.1.2)
Lus10017459
53 / 2e-08
AT3G11400 447 / 3e-160
eukaryotic translation initiation factor 3G1 (.1.2)
Lus10038932
51 / 1e-07
AT5G51300 897 / 0.0
splicing factor-related (.1.2.3)
Lus10003957
51 / 1e-07
AT5G64200 217 / 9e-70
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
Lus10023802
50 / 2e-07
AT5G64200 217 / 1e-68
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
Lus10040854
44 / 4e-06
AT3G46020 112 / 1e-33
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10042589
47 / 5e-06
AT2G21440 825 / 0.0
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.006G048300.3 pacid=42770307 polypeptide=Potri.006G048300.3.p locus=Potri.006G048300 ID=Potri.006G048300.3.v4.1 annot-version=v4.1
ATGAGAGGCAGAGTCATGGTAGACGAAGACAACGGCGACGTCGTGATGACGGAACAGCCAAGAGGAAGAGGAAGAGAAAGGCGCAAAGGCAGAGGTCTCA
GACTCGGAGGACGAGGAGAATCAAGGCAACAGACACTACTCCATGACGAAGAAGAAGAGGAAGAGGGTGGGAATGACCGAAATGTCCTTCAACTCTTCAC
TCCTCTTCGCTCTGTAGAGGGTTGGAGTGTTTTGTTGAGTGGAGTGCATGAAGAAGCACAAGAAGATGATATTCTCACTCTATTTGGTGCTTTTGGCCAC
ATCAACAACTTGCATCTCAATCTTGATCGCCGCACCGGATTTCTTAAGGGGTATGCTCTGATAGAATATGAGAACTTTATGGAAGCTCAAGCTGCTATAT
CTGGAATGAATGGGACTAAACTCTTCTTTCGGATCATCTCTGTTGACTGGGCTTTCAGCACTGGTCCTTTCGAAGGGAGAAGGTCTGCTTGTTTTCATGA
ATTATCATATGCTATAGCTATATTTGGTGTTAGCCTAACCTGTAACCCTTTCTCAGCTTGA
AA sequence
>Potri.006G048300.3 pacid=42770307 polypeptide=Potri.006G048300.3.p locus=Potri.006G048300 ID=Potri.006G048300.3.v4.1 annot-version=v4.1
MRGRVMVDEDNGDVVMTEQPRGRGRERRKGRGLRLGGRGESRQQTLLHDEEEEEEGGNDRNVLQLFTPLRSVEGWSVLLSGVHEEAQEDDILTLFGAFGH
INNLHLNLDRRTGFLKGYALIEYENFMEAQAAISGMNGTKLFFRIISVDWAFSTGPFEGRRSACFHELSYAIAIFGVSLTCNPFSA
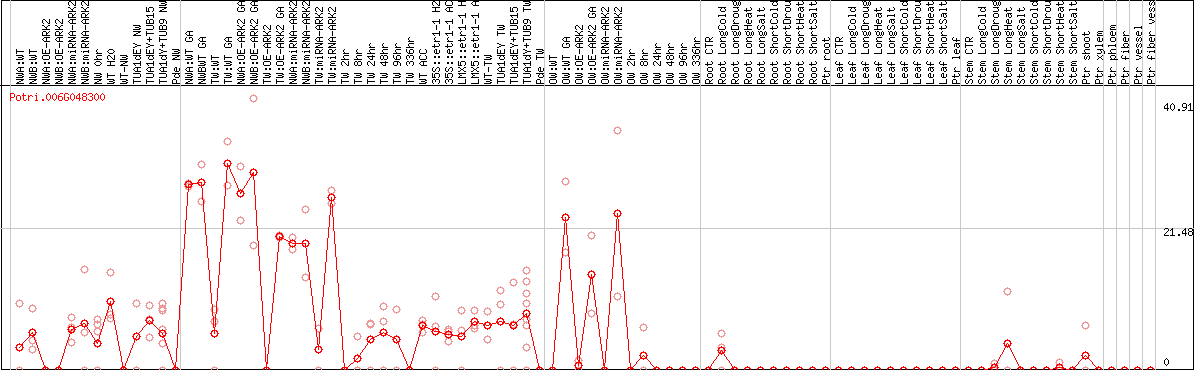
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G048300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.