Potri.006G049300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G049300.2 pacid=42769929 polypeptide=Potri.006G049300.2.p locus=Potri.006G049300 ID=Potri.006G049300.2.v4.1 annot-version=v4.1
ATGGGGAATTCAACGAGGCAGTTGAGGGCTATGCTTCGTAAGAATTGGCTTCTCAAAATTCGCCATCCTTTTATCACTTCTGCTGAGATCTTGCTTCCTA
CAATTGTAATGTTACTGCTAATAGCGGTGCGGACAAGAGTTGATTTGCAAATTCACCCTGCGCAGGCGTGTATAAAGGAAAATATGCTTGTAGAAGTAGG
AAAGGGTATGTCCCCGAATTTTCAAGAAGTTTTGGAGGCATTGCTGGTGAGGGGAGAGTTTTTGGCTTTTGCACCGGATACAGAAGAAACAAGGATGATG
ATTAACTTAATGTCAATCAAGTTCCCTCTACTGCAGCAAGTTTCTCTGATCTACAAAGATGAACTCGAGTTGGAAACATATTTAACCTCTGATCTTTATG
GCACCTGTAGTCAAGTCAAGAATTGCTCAAATCCAAAAATCAAAGGAGCAGTAGTATTTCATAATCAAGGTCCTCAGCTATTTGATTACAGTATACGACT
TAATCACACATGGGCTTTCTCTGGGTTTCCTGATGTTAGAACCATAATGGATGTGAATGGACCTTATCTCAATGACTTGGAACTGGGTGTTAATATAATA
CCGACAATGCAATACAGCTCCAGCGCTTTTTTCACTCTTCAGCAGGTAGTGGATTCATTCATAATATTTGCCTCTCAGCAAACTGAAACTGAGTCTTCAA
CTGAACATATAGAACTTCCATCGTCCAATTCATTCAACAAATCTTCCTCGCTCAAGTTGCCATGGACAAAATTTAGCCCTTCAAAAATTAGAATCGCTCC
CTTTCCAACACGTGAATACACCGATGATCAATTTCAGTCAATTATCAAGAGGGTCATGGGAGTATTGTATCTTTTGGGATTTCTCTACCCAATCTCTGGC
CTTATCAGTTATTCTGTGTTCGAGAAGGAACAAAAGATAAGAGAAGGTCTCTACATGATGGGCTTGAAAGACGGGATATTCCATCTTTCTTGGTTTATTA
CATATGCTTTGCAGTTTGCAATTTCATCTGGGATTATTACAGCTTGCACCTTGAATAACCTTTTCAAGTACAGCGACAAGTCAGTGGTGTTTGTGTATTT
TTTTTCATTTGGACTCAGTGCAATTATGCTGTCCTTTTTGATATCTACATTCTTCACACGAGCGAAAACAGCAGTGGCAGTTGGTACCCTTTCGTTTTTT
GGGGCCTTCTTTCCTTATTACACTGTCAATGACCCGGCTGTCCCTATGATATTGAAGGTCCTTGCCTCTTTGCTATCACCTACAGCCTTTGCTCTAGGAT
CAATTAACTTTGCCGATTATGAGCGTGCTCATGTTGGACTTCGCTGGAGCAACATATGGCGGGAGTCATCTGGGGTGAATTTTTTGGTCTGTCTTCTGAT
GATGTTATTTGACACATTGATATACTGTGCTATTGGCCTCTACCTTGATAAGGTCCTTCCCAGGGAAAATGGGATGCGCTACCCATGGAACTTCTTATTC
CAGAAGTGCTTCTGGAGGAAGAATAATTTTGTCAAACATCATGGTTCCAGTTTAGAATCCAACTTTAACGATGAGCTTTCCAATGAAAGGGCAAGCTTTT
TAGGAAATAATACCCAAGAACCTGCTGTTGAAGCGATAAGCTTAGATATGAAGCAACAAGAACTTGATAAAAGATGCATCCAGATTAGGAATCTGCGTAA
GGTGTATGCTTCTAAAAGGGGAAACTGTTGTGCTGTCAATTCTTTGCAGCTTACCTTATATGAAAATCAGATTCTTGCTCTTCTAGGACATAATGGAGCT
GGAAAAAGCACAACGATTTCAATGCTTGTTGGTCTTCTTCCTCCTACTTCTGGGGATGCTTTAGTATTTGGAAAGAATATCACAACAGACATGGATGAGA
TACGGAATGGACTGGGTGTGTGCCCTCAAAATGATATCCTTTTTCCAGAATTGACTGTGAGAGAACATTTAGAAATTTTTGCTGCATTGAAGGGTGTGAA
GGAAGATATTTTGGAGAGGGACGTAACTGATATGGTCAATGAAGTGGGTCTGGCTGATAAGGTCAATACTGCTGTGAGGGCTCTTTCTGGTGGCATGAAG
AGGAAATTATCTCTGGGAATTGCACTGATAGGAAATAGCAAGGTCGTAATTCTTGACGAACCTACGAGTGGGATGGATCCATACTCAATGCGCTTGACAT
GGCAGTTAATAAAAAGAATAAAGAAGGGCAGGATAATCTTACTTACAACACACTCGATGGATGAAGCTGATGAACTAGGAGATCGGATAGCTATCATGGC
TAATGGATCCTTGAAATGCTGTGGAAGCTCCCTATTTTTGAAGCATCAGTATGGGGTTGGTTATACCCTTACATTAGTGAAGTCGTCCCCTACTGCTTCT
GTGGCTTCTGATATTGTTTACCGCCATGTTCCATCTGCAACGTGTGTGAGCGAGGTTGGAACTGAGATTTCTTTTAAGCTGCCTTTGGCATCATCGGTTT
CTTTTGAAAGCATGTTTAGGGAAATTGAAAGTTGCATGAGAAGATCAATTTCTAAATCAGAAATGAGTAGTAGTGAGGACAAAAGTTATCCTGGCATTGA
AAGTTATGGTATTTCAGTCACAACTCTGGAAGAGGTATTCCTGAGAGTTGCAGGATGCGGGTACGATGAAACTGATGATTTCGTGGACAGAAACAACATT
CTTTCATCTAACTCTACAGTTCCTGCAGCTTATGACAATCGCCCTTCCGAAACAATTTTTGATGCAAAAATTTTGGGAAATTATAAAAAGATTATTGGAT
TCATATCTGCCATGGTGGGGAGAGTCTCTGGTTTAATGGCCGCCACAATCTTAAGTTTCATCAATTTCTTAGGTATGCAGTGCTGTTGCTGCTGTATCAT
TTCAAGATCAACATTTTGGCAGCATACTAAGGCATTATTCATAAAGAGGGCTATATCTGCTCGGAGAGATCGGAAAACAATTGTTTTCCAACTTCTAATT
CCTGCCATTTTCTTGCTTTTTGGTCTTCTTTTTCTCAAGCTTAAGTCACATCCTGATCAGCAGTCTGTTACTCTAACAACTTCACATTTTAATCCTCTCT
TAAGTGGTGGTGGTGGTGGTGGTCCAATTCCTTTTGATCTATCCTTGCCTATTGCAAAAGAGGTTGCTGGATATATTAAAGGGGGGTGGATTCAAAACTT
TAGACAGAGTGCATACAGATTCCCTGATGCAGAGAGGGAATTGGCTGATGCAATCAAGGCAGCAGGACCAACTTTGGGACCAGTTTTACTTTCTATGAGT
GAATTTTTAATGTCTAGCTTTAATGAATCCTATCAGTCAAGGTATGGGGCCGTTGTGATGGATAAAAAACATGATGATGGAAGCTTGGGATATACCATTC
TACACAACAGTTCCTGTCAGCATGCTGCTCCAACCTTTATCAATCTAATGAATGCTGCGATTTTGAGGCTTGCTACTGGTGACCAAAATATGACAATTCA
AACACGCAACCACCCTCTGCCGATGACCAAAAGCCAGCACTTACAACATCATGATTTGGATGCCTTTTCTGCTGCAATTATTGTCAATATCGCCTTCTCA
TTCATTCCTGCTTCATTCGCAGTTGCCATTGTGAAGGAACGTGAAGTTAAGGCAAAGCACCAACAATTGATTAGTGGGGTTTCTGTACTTTCATATTGGG
TTTCTACATACATCTGGGACTTCATTAGCTTCTTAATTCCTTCATCTTTTGCTCTCTTACTGTTCTATATTTTTGGTCTGGATCAGTTTATTGGAAAGGA
TTGCTTCTTGCCAACGTTCCTCATGTTCTTAGAGTATGGACTAGCAATTGCGTCTTCAACATATTGCCTTACCTTCTGCTTTTCTGAACACTCTATGGCT
CAGAATGTGGTTCTGTTGGTCCACTTTTTCACTGGGTTGATTCTCATGGTTATCTCATTTATCATGGGCCTTATACAAACAACAGCAAGTGCAAATAATT
TGCTTAAGAACTTCTTCAGATTATCACCAGGATTTTGCTTTGCTGATGGACTAGCTTCACTGGCTCTTCTAAGACAAGGGATGAAAGATAAATCTAGTAA
TGCAGTTTTTGACTGGAATGTAACTGGTGCTTCCCTTTGTTATCTCGGCTTTGAGAGCATCGGTTACTTTCTTCTCACACTTGGGTGGGAACTGTTGCCT
TTTCACAAATTGACTCCAGTAGGAATTAAGCAGTATTGGAGGAGCATCATGAATCTCCAGCATGACACTCATGATTTAGAACCACTTCTAAAATCTCCAT
CAGAAACTGTTGATCTCAACTTTGATGAGGACATAGATGTGCAAACAGAAAGAAATAGGGTACTTGCAGGTTCCATTGACAATGCTATCATCTACTTGCG
TAACCTTAGGAAGGTATATCCTGGAGAAAAACATCGTACAAAAGTTGCTGTGCGTTCTTTGACTTTCTCAGTTCAAGCAGGAGAATGTTTTGGCTTTTTA
GGAACGAATGGAGCTGGGAAGACAACTACATTGTCAATGTTAACTGGAGAGGAATCTCCAACTGATGGAAGTGCTTTCATTTTTGGTAAAGACACGCGTT
CAGATCCCAAGGCTGCCCGTCGGCATATTGGGTATTGTCCCCAATTTGATGCTCTATTGGAATTTTTAACTGTCCAAGAGCATCTCGAGCTTTATGCAAG
AATAAAAGGAGTTGCAGATTACAGAATTGACGATGTTGTAATGGAAAAGTTGTTGGAGTTTGACTTGCTGAAGCATGCCAACAAACCATCTTTCACTCTT
AGTGGGGGAAACAAACGCAAATTGTCTGTTGCAATTGCAATGATTGGAGATCCACCTATTGTAATTCTTGATGAGCCATCGACAGGCATGGATCCTATTG
CTAAACGATTCATGTGGGAAGTCATATCTCGTTTGTCAACCAGACAAGGGAAGACAGCAGTAATTCTTACTACCCACAGCATGAATGAAGCTCAAGCACT
GTGCACCCGAATAGGAATAATGGTTGGAGGCCGATTAAGATGCATTGGAAGTCCTCAGCATTTGAAAACACGATTTGGAAATCACCTTGAACTAGAGGTT
AAGCCTACCGAAGTCAGCTCTGTAGATTTGGAAAACCTTTGCCAAACCATTCAAAGCAGACTTTTCGACATTCCTTCACATCCAAGAAGTTTACTTGATG
ACATTGAAGTTTGCATTGGGAGAATTGACTCCATTACATCTGAAAATGCATCAGTGATGGAGATCAGCTTGTCTCAAGAAATGATTATTTTGATTGGACG
CTGGCTTGGAAATGAAGAAAGAGTGAAGACACTTGTATCTTCAACACCCATTTCTGATGGGGTCTTTGGTGAACAGTTATCGGAGCAGTTGGTTCGAGAT
GGTGGGATACCTTTGCCAATATTTTCAGAGTGGTGGTTAGCTATAGAGAAGTTTTCTGCTATTGATTCATTTATCCTGTCATCCTTTCCTGGGGCAGCAT
TTCAAGGGTGCAATGGTTTGAGTGTTAAATATCAGTTGCCATATAGTAAGGACCTATCGCTTGCGGATGTTTTTGGACACATCGAACAAAATAGAAATCA
ACTGGGCATAGCAGAATACAGTATCAGCCAGTCAACACTTGAGACCATTTTTAATCATTTTGCAGCAAGTTCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G049300.2 pacid=42769929 polypeptide=Potri.006G049300.2.p locus=Potri.006G049300 ID=Potri.006G049300.2.v4.1 annot-version=v4.1
MGNSTRQLRAMLRKNWLLKIRHPFITSAEILLPTIVMLLLIAVRTRVDLQIHPAQACIKENMLVEVGKGMSPNFQEVLEALLVRGEFLAFAPDTEETRMM
INLMSIKFPLLQQVSLIYKDELELETYLTSDLYGTCSQVKNCSNPKIKGAVVFHNQGPQLFDYSIRLNHTWAFSGFPDVRTIMDVNGPYLNDLELGVNII
PTMQYSSSAFFTLQQVVDSFIIFASQQTETESSTEHIELPSSNSFNKSSSLKLPWTKFSPSKIRIAPFPTREYTDDQFQSIIKRVMGVLYLLGFLYPISG
LISYSVFEKEQKIREGLYMMGLKDGIFHLSWFITYALQFAISSGIITACTLNNLFKYSDKSVVFVYFFSFGLSAIMLSFLISTFFTRAKTAVAVGTLSFF
GAFFPYYTVNDPAVPMILKVLASLLSPTAFALGSINFADYERAHVGLRWSNIWRESSGVNFLVCLLMMLFDTLIYCAIGLYLDKVLPRENGMRYPWNFLF
QKCFWRKNNFVKHHGSSLESNFNDELSNERASFLGNNTQEPAVEAISLDMKQQELDKRCIQIRNLRKVYASKRGNCCAVNSLQLTLYENQILALLGHNGA
GKSTTISMLVGLLPPTSGDALVFGKNITTDMDEIRNGLGVCPQNDILFPELTVREHLEIFAALKGVKEDILERDVTDMVNEVGLADKVNTAVRALSGGMK
RKLSLGIALIGNSKVVILDEPTSGMDPYSMRLTWQLIKRIKKGRIILLTTHSMDEADELGDRIAIMANGSLKCCGSSLFLKHQYGVGYTLTLVKSSPTAS
VASDIVYRHVPSATCVSEVGTEISFKLPLASSVSFESMFREIESCMRRSISKSEMSSSEDKSYPGIESYGISVTTLEEVFLRVAGCGYDETDDFVDRNNI
LSSNSTVPAAYDNRPSETIFDAKILGNYKKIIGFISAMVGRVSGLMAATILSFINFLGMQCCCCCIISRSTFWQHTKALFIKRAISARRDRKTIVFQLLI
PAIFLLFGLLFLKLKSHPDQQSVTLTTSHFNPLLSGGGGGGPIPFDLSLPIAKEVAGYIKGGWIQNFRQSAYRFPDAERELADAIKAAGPTLGPVLLSMS
EFLMSSFNESYQSRYGAVVMDKKHDDGSLGYTILHNSSCQHAAPTFINLMNAAILRLATGDQNMTIQTRNHPLPMTKSQHLQHHDLDAFSAAIIVNIAFS
FIPASFAVAIVKEREVKAKHQQLISGVSVLSYWVSTYIWDFISFLIPSSFALLLFYIFGLDQFIGKDCFLPTFLMFLEYGLAIASSTYCLTFCFSEHSMA
QNVVLLVHFFTGLILMVISFIMGLIQTTASANNLLKNFFRLSPGFCFADGLASLALLRQGMKDKSSNAVFDWNVTGASLCYLGFESIGYFLLTLGWELLP
FHKLTPVGIKQYWRSIMNLQHDTHDLEPLLKSPSETVDLNFDEDIDVQTERNRVLAGSIDNAIIYLRNLRKVYPGEKHRTKVAVRSLTFSVQAGECFGFL
GTNGAGKTTTLSMLTGEESPTDGSAFIFGKDTRSDPKAARRHIGYCPQFDALLEFLTVQEHLELYARIKGVADYRIDDVVMEKLLEFDLLKHANKPSFTL
SGGNKRKLSVAIAMIGDPPIVILDEPSTGMDPIAKRFMWEVISRLSTRQGKTAVILTTHSMNEAQALCTRIGIMVGGRLRCIGSPQHLKTRFGNHLELEV
KPTEVSSVDLENLCQTIQSRLFDIPSHPRSLLDDIEVCIGRIDSITSENASVMEISLSQEMIILIGRWLGNEERVKTLVSSTPISDGVFGEQLSEQLVRD
GGIPLPIFSEWWLAIEKFSAIDSFILSSFPGAAFQGCNGLSVKYQLPYSKDLSLADVFGHIEQNRNQLGIAEYSISQSTLETIFNHFAASS
|
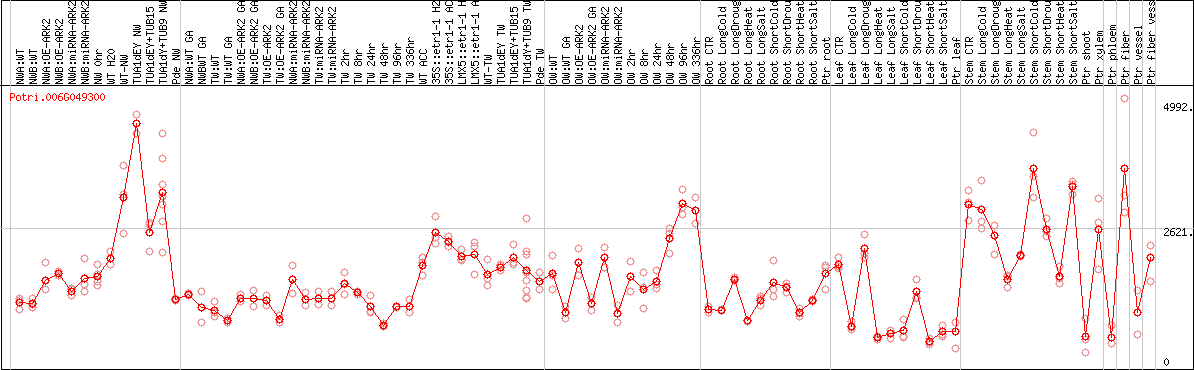
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G049300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
