Potri.006G055300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G055300.1 pacid=42768806 polypeptide=Potri.006G055300.1.p locus=Potri.006G055300 ID=Potri.006G055300.1.v4.1 annot-version=v4.1
ATGGCTGCCCTAGAGGGTGCAACTGTGTCAGTTGATTCGGTAGCGTTTAGCTTCTTAACGGATGAAGAGGTTCATAAGCATAGTTTTGTAAAGATTACCA
GCGCGAGATTGCTTGATACTTTGGACAAACCTGTGCCTGGTGGATTATATGACCCAGCAATGGGGCCTCTAGGGGATGAGCCTTGCAAAACATGCGGTCA
ACGTTCAACTAATTGCACAGGTCATTGTGGCCATATAGATCTGATTAGTCCAGTTTACAATCCTCTGCTCTTTAATTTTCTTCACAAACTTCTGCAAAGA
ACCTGTTTCTTCTGCTTTCATTTTAGAGCAGATAGCAACCAGGTTGAGAAATTTGTTTCTCAATTGGAACTCATCATAAAAGGTGATGTTGTTGGGGCTA
AAAGGTTGGATTCTTTCTCGCCAATTGAAGCCTCATTGCCAGAAGACAGTGATGGCAGCAGTGAATCTTGTTCTACAATTCATTCAGGAGCACGGCATCC
AAACAATGAGCAGTCAAAGCAGAGCGAATGGACCTCCCTTCAGTTATCTGAAGCAATGTCTATCTTAAATAATTTTCTGAAGTTGGAATCAAAGAAGTGC
AAGAACTGTTCAGCATCAAATCCTAATATTCGGAAGCCTACTTTTGGTTGGTTTCATTGGGCAGGTTTATCAAATGCAGCTATTAGATCAAATCTGATTA
AGCAGCAGACAATTGAAGGGCCATTTGGGGGTGCATTTGAAGAGTTGATAGATGCAGAAGATGCAACCAAATCTCCTAGTAACAAAGAGTCTGCCACAAA
CAGGAATTTGAAAGAACACCAGAAATTGCAACATCAGTTCACTAGCCAGAAAGATGCTTTGTCCTCGCAGTTATTACCATCAGAAGCGATGGATATTTTA
AAGCTTTTGTGGAAAAATGAAGCTCGACTCTGCTCATTGATGAGTGACATTCAGCAGCAAGGAGTAGGAAAGAAAAAGGCAGGACATTCTATGTTCTTTC
TGAACACTGTTTTAGTGCCTCCAATCAAATTCAGGCCTCCGACCAAGGGAGGCGACTCTGTGATGGAACATCCTCTATCTGTTTTGCTGAGCAAGGTGCT
GGAATTGAATGGATCTTTGGCAGATGCGCATAGAAGCAATGATTTCCCATTAATAGCTAGGCGGTGGTTGGAACTTCAGCAATCTCTAAATGTCTTGTTT
GATTCCAATACTGCCAAGGGTCAGAAGGATGTGATTTCTGGGATATGCCAGATACTTGAGAAAAAGGAGGGTATGTTTCGGCAGAAAATGATGGGAAAGA
GGGTTAATTATGCATGCCGGTCTGTTATTTCCCCAGATCCTTATTTAGATGTTAACGAAATTGGAATTCCTCCGTGCTTTGCAGTGAAGTTAACATATCC
TGAGAGAGTGACACCCTGGAATGTTGCCAAATTGAGAAATGCTGTAATCAATGGTGCTGAAAGTCATCCTGGAGCTACACACTATGTGGATAAGCTGTCT
ACTACTAAGCTTCCACTGAATCGAAAAATGCGTGTTTCTGTAGCAAGGAAATTATCCGGGAGGAGTGTTGATTATGAATATGAAGGGAAAATTGTATATC
GCCATTTGCAAGATGGAGATATTGTGCTTGTCAATCGACAGCCTACACTTCACAAACCAAGTATAATGGCACATGTAGTTCGTGTATTGAAGGGGGAGAA
GACACTACGCATGCACTATGCAAATTGCAGTACATACAATGCTGATTTTGATGGTGATGAAATGAATGTTCATTTCCCACAAGATGAAGTTTCTCGAGCT
GAAGGTTACAACATTGTTAATGCAAATAATCAATATGTCAGGCCTTCCAATGGCGAACCAATTAGATCCTTGATTCAGGATCATATTATTAGTGCTGTGC
TTTTAACAAAGAAGGACACTTTTTTGACTGAGGATGAAGTTTATCAACTGCTTTATAGTTCTGGCGTGTCAAATGCTCGTCCAACATCTTTCAGTGGCAA
AGCTGGCCGGAAAGTTATATTCTTAAGTTATGAAGATGAGATTGAAACCGTTGATCCTGCAATAAGGAAACCCATTTATCTGTGGTCAGGAAAACAGTTG
ATTACTGCTGTGTTGAATCACATTACAAGGGGCCATCCACCATTCACTGTTGAGAAAGGTGGGAAGCTCTCGTATGATTTTTTTAAGAGTAAAATTAAGA
ATGGCAAATCTAGCAATGGAGAAAAAGTTGGTGTAAGCAAACCAATGAAGGAAAAAGAGTCTGGCAAAGTCAATCCAAAAGAAAATCAGCTTGAAGATGA
TAAGATGATTATCTTTAGGAATGTATTGGTCCAAGGGGTGATTGACAAGGCCCAGTTTGGTGAGTATGGTTTGGTGCACACAGTTCAAGAATTGTTTGGT
GCAAAAGCAGCTGGCACCCTGCTCTCTGTTTTCAGTCGTTTATTCACTGCTTATTTGCAGATGCATGGATTCACATGTGGAGTTGATGATCTTTTGATAA
CGAAAATTAAAGACGACGAAAGGAAGAAGCAACTTGAAAACTGTGAAAAGTGTGGGGAACAAATTCATCGTAAATTTATCGGAATCAAGGATGAGAACAT
TAAGATAGATCCATTGGAATTACAATCAAATATCGAGAAAACTATACGCAGTGATGGTGAGTCTGCCTTGACATATCTGGATAGGCAGATGACAAATGAA
CTAAATTCCAAAACAAGCTCAGGAGTCATCAATGAGCTGTTGTCTGAAGGGTTGCTAAAACCCTCTGGAAAGAATTGCATATCACTTATGACTACATCTG
GGGCTAAAGGTAGCAAGGTCAATTTCCAACAGATATCTTCTTTTCTAGGCCAACAGGAGTTGGAAGGTAAAAGAGTTCCTCGAATGGTTTCTGGAAAGAC
ATTGCCCTGCTTTCATCCATGGGATTGGGCAGCCCGTGCTGGAGGCTATATTATTGATCGTTTTCTCACCGGTCTGCGTCCTCAAGAATATTATTTCCAT
TGTATGGCTGGTCGTGAAGGGTTGGTTGATACAGCTGTCAAAACATCTCGCAGTGGATACCTCCAAAGATGCCTGATTAAAAATCTTGAGTGCCTCAGAA
TCGGCTATGATCATACAGTTCGTGATGCTGATGGCTCAATTGTTCAATTTTACTATGGAGAAGATGGTGTGGATGTTCATCAAACGGGCTTTATTGCTAA
ATTTGAAGCCTTGGCAGCGAATCGAGAAATAATATACGAAAAAAGTGATGAGCTTGGCACATGTAATGCTTATATTTCGGAATTGCCTAAAGCTCTTAAA
GAAAAAGCAGAAACATTCCTACGTAAAATCGCAAAGGAGCAGAGTTCTTTACATGATCCCACAAAGGATCGGAGTTCTAATTTAGTGGAACATGATTTCT
ATAAGTTACTGAAGCAAAAGTTTTTCACGAGTCTTGCACAACCAGGAGAACCAGTTGGTGTCCTTGCTGCTCAGTCTGTAGGAGAACCATCAACACAGAT
GACGCTGAACACTTTCCATCTAGCCGGGCGTGGTGAAATGAATGTAACACTTGGAATTCCACGTCTCCAAGAGATTTTGATGACTGCTTCAGCTGATATC
AAGACTCCGATTATGACCTGCCCCTTGCATGAAGGGAGAACTAAAGAGGATGCTGAGCGCCTTTCTGATAAGTTGAAGAAGGTCACCGTAGCCGATATAG
TAGAGAGCATGGAAGTATCTGTCATGCCATTTGCTGTTCAGAATGACGACATTTGCAGAATTTATAAGCTTAAAATGAAGCTTTATACACCAGCGCACTA
CCCACAACATGCTGATATATCAGTAGAAAATTGGGAAGAAACCCTTGAGGTGTTGTTTGTGAGAGAGTTGGAGGATGCTATACAAAATTATCTGGTTTTG
CTTTCTAAAATTAGCGGCATTAAGAATTTTTTGAAGGAATCACATTCAGGGACGCCAATTGAAGCAGAAGAAGATGTTTCTGGCAACATATCACGCGAGG
GTGAAAACGATGATGATAGTGATGATGAAGAAGGGGAAGAGGCTGATGATCTAGGTTTAGATGTGCAGAAAAGAAAACAGCAAGTTACAGATGAGATGGA
TTATGATGATGGCTCTGAGGGGGTACTGAATGAAGATGAAGGGGATTTATCAGGAAGCCAGGCTCCATCTGGATCTGAAAGTGATACTGAACCGGCTGAT
AAAGAAAGTGAGATCAGTAATACCGATATGGTTGACTATGACAGTGAATATTTTGAAAAGCCAACACATCTTGGAAATTACTCTAAGCCAAAGTCAAGGA
AGAAAACCTCAGAAAGTTCATCACAAGTCGAAATGCATTCTAAGCTAAAGTCAACAGAAAAGAAGAAACAAAAGGCCAAGGGAAAGAAAGTTAGATCGAA
ATTAGTTAAAAAGGACTTTGATAGATCAATCTTTGTGGAAGCCAAGGGATTGCACTTTGAAATCCATCTTAAATTCACCAATGAGCCTCATATTTTACTG
GCCGAGATTGCTCAGAAGACAGCCAAGAAGGTCTGTATCCAGAATCCCGGGAAGGTTCAACGGTGTCAAGTGACTGATTGCAAAGAAAATCAAGTGATTT
ACTATGGGAAAGACCCCAAGAGAAGAATTGATATTGAACCTGGGGAAAAACAAAAAATCCCAGCGCTTCATACAATTGGAGTGGACTTCAATACATTTTG
GAAAATGCAAGATCACCTTGATGTTAGGTATATGTATTCAAATAGCATTCATGGTATGTTGAAAGCTTATGGAGTAGAGGCCGCAAGAGAGACAATCATT
AGGGAGATAAAGCACGTGTTCAATTCTTATGGAATATCCGTTAATACACGGCATTTGTCTCTTATTGCTGATTATATGACCCATACCGGTGAGTACCGTC
CGATGAGCCGTATTGGGGGCATATCAGAGTCGATATCACCTTTAAGTAAAATGTCTTTTGAGACAGCGTCCAAATTCATAGTTGAAGCAGCATTGCACGG
GGAGGTCGATAATTTGGAAGCTCCATCTGCAAGGGTTTGTCTTGGATTGCCAGTGAAAATGGGTACTGGATCTTTTGATTTGATGCAGAAACTGGAAATT
TGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G055300.1 pacid=42768806 polypeptide=Potri.006G055300.1.p locus=Potri.006G055300 ID=Potri.006G055300.1.v4.1 annot-version=v4.1
MAALEGATVSVDSVAFSFLTDEEVHKHSFVKITSARLLDTLDKPVPGGLYDPAMGPLGDEPCKTCGQRSTNCTGHCGHIDLISPVYNPLLFNFLHKLLQR
TCFFCFHFRADSNQVEKFVSQLELIIKGDVVGAKRLDSFSPIEASLPEDSDGSSESCSTIHSGARHPNNEQSKQSEWTSLQLSEAMSILNNFLKLESKKC
KNCSASNPNIRKPTFGWFHWAGLSNAAIRSNLIKQQTIEGPFGGAFEELIDAEDATKSPSNKESATNRNLKEHQKLQHQFTSQKDALSSQLLPSEAMDIL
KLLWKNEARLCSLMSDIQQQGVGKKKAGHSMFFLNTVLVPPIKFRPPTKGGDSVMEHPLSVLLSKVLELNGSLADAHRSNDFPLIARRWLELQQSLNVLF
DSNTAKGQKDVISGICQILEKKEGMFRQKMMGKRVNYACRSVISPDPYLDVNEIGIPPCFAVKLTYPERVTPWNVAKLRNAVINGAESHPGATHYVDKLS
TTKLPLNRKMRVSVARKLSGRSVDYEYEGKIVYRHLQDGDIVLVNRQPTLHKPSIMAHVVRVLKGEKTLRMHYANCSTYNADFDGDEMNVHFPQDEVSRA
EGYNIVNANNQYVRPSNGEPIRSLIQDHIISAVLLTKKDTFLTEDEVYQLLYSSGVSNARPTSFSGKAGRKVIFLSYEDEIETVDPAIRKPIYLWSGKQL
ITAVLNHITRGHPPFTVEKGGKLSYDFFKSKIKNGKSSNGEKVGVSKPMKEKESGKVNPKENQLEDDKMIIFRNVLVQGVIDKAQFGEYGLVHTVQELFG
AKAAGTLLSVFSRLFTAYLQMHGFTCGVDDLLITKIKDDERKKQLENCEKCGEQIHRKFIGIKDENIKIDPLELQSNIEKTIRSDGESALTYLDRQMTNE
LNSKTSSGVINELLSEGLLKPSGKNCISLMTTSGAKGSKVNFQQISSFLGQQELEGKRVPRMVSGKTLPCFHPWDWAARAGGYIIDRFLTGLRPQEYYFH
CMAGREGLVDTAVKTSRSGYLQRCLIKNLECLRIGYDHTVRDADGSIVQFYYGEDGVDVHQTGFIAKFEALAANREIIYEKSDELGTCNAYISELPKALK
EKAETFLRKIAKEQSSLHDPTKDRSSNLVEHDFYKLLKQKFFTSLAQPGEPVGVLAAQSVGEPSTQMTLNTFHLAGRGEMNVTLGIPRLQEILMTASADI
KTPIMTCPLHEGRTKEDAERLSDKLKKVTVADIVESMEVSVMPFAVQNDDICRIYKLKMKLYTPAHYPQHADISVENWEETLEVLFVRELEDAIQNYLVL
LSKISGIKNFLKESHSGTPIEAEEDVSGNISREGENDDDSDDEEGEEADDLGLDVQKRKQQVTDEMDYDDGSEGVLNEDEGDLSGSQAPSGSESDTEPAD
KESEISNTDMVDYDSEYFEKPTHLGNYSKPKSRKKTSESSSQVEMHSKLKSTEKKKQKAKGKKVRSKLVKKDFDRSIFVEAKGLHFEIHLKFTNEPHILL
AEIAQKTAKKVCIQNPGKVQRCQVTDCKENQVIYYGKDPKRRIDIEPGEKQKIPALHTIGVDFNTFWKMQDHLDVRYMYSNSIHGMLKAYGVEAARETII
REIKHVFNSYGISVNTRHLSLIADYMTHTGEYRPMSRIGGISESISPLSKMSFETASKFIVEAALHGEVDNLEAPSARVCLGLPVKMGTGSFDLMQKLEI
|
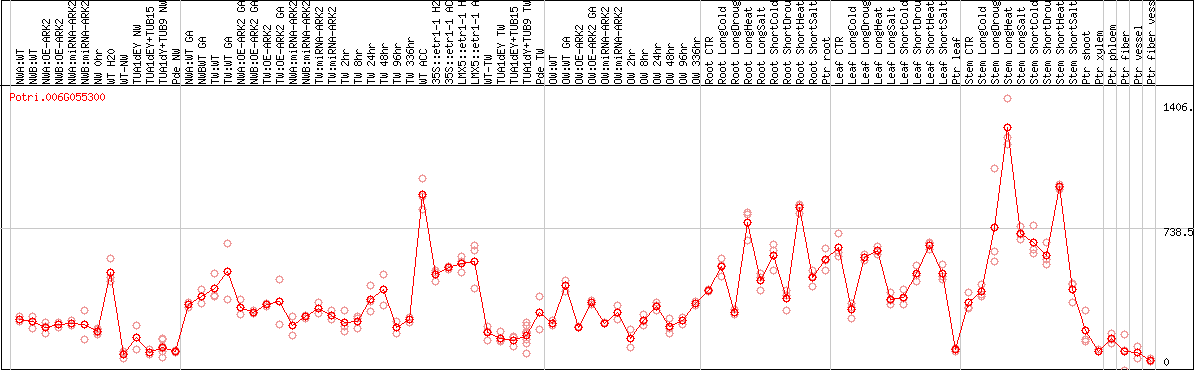
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G055300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
