External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06710 163 / 4e-49
HD
HAT14
homeobox from Arabidopsis thaliana (.1.2)
AT2G22800 160 / 7e-49
HD
HAT9
Homeobox-leucine zipper protein family (.1)
AT4G37790 154 / 2e-46
HD
HAT22
Homeobox-leucine zipper protein family (.1)
AT4G16780 145 / 7e-43
HD
ATHB2, HAT4, ATHB-2
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 2, homeobox protein 2 (.1)
AT3G60390 146 / 9e-43
HD
HAT3
homeobox-leucine zipper protein 3 (.1)
AT2G44910 143 / 2e-41
HD
ATHB4, ATHB-4
ARABIDOPSIS THALIANA HOMEOBOX-LEUCINE ZIPPER PROTEIN 4, homeobox-leucine zipper protein 4 (.1)
AT5G47370 142 / 2e-41
HD
HAT2
Homeobox-leucine zipper protein 4 (HB-4) / HD-ZIP protein (.1)
AT4G17460 138 / 4e-40
HD
HAT1
Homeobox-leucine zipper protein 4 (HB-4) / HD-ZIP protein (.1)
AT2G01430 129 / 2e-36
HD
ATHB17, ATHB-17
ARABIDOPSIS THALIANA HOMEOBOX-LEUCINE ZIPPER PROTEIN 17, homeobox-leucine zipper protein 17 (.1)
AT1G70920 119 / 2e-33
HD
ATHB18
homeobox-leucine zipper protein 18 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004161
166 / 4e-50
AT5G06710 263 / 5e-86
homeobox from Arabidopsis thaliana (.1.2)
Lus10021064
163 / 6e-50
AT5G06710 265 / 3e-88
homeobox from Arabidopsis thaliana (.1.2)
Lus10016254
162 / 3e-48
AT5G06710 274 / 4e-90
homeobox from Arabidopsis thaliana (.1.2)
Lus10029293
160 / 1e-47
AT5G06710 274 / 5e-90
homeobox from Arabidopsis thaliana (.1.2)
Lus10040175
152 / 6e-45
AT4G16780 280 / 1e-93
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 2, homeobox protein 2 (.1)
Lus10004759
152 / 8e-45
AT4G16780 305 / 1e-103
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 2, homeobox protein 2 (.1)
Lus10004377
152 / 1e-44
AT4G16780 265 / 1e-87
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 2, homeobox protein 2 (.1)
Lus10007849
151 / 1e-44
AT4G16780 303 / 6e-103
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 2, homeobox protein 2 (.1)
Lus10027633
138 / 8e-41
AT4G37790 138 / 3e-40
Homeobox-leucine zipper protein family (.1)
Lus10011935
134 / 3e-39
AT4G37790 142 / 1e-41
Homeobox-leucine zipper protein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00046
Homeodomain
Homeodomain
CL0123
PF02183
HALZ
Homeobox associated leucine zipper
Representative CDS sequence
>Potri.006G071600.1 pacid=42768105 polypeptide=Potri.006G071600.1.p locus=Potri.006G071600 ID=Potri.006G071600.1.v4.1 annot-version=v4.1
ATGGAAGAAGAAGGGTACTGTAACACAAGACTTGGGCTTGGACTAGGAGGTGAAAACTATGCGCCATGGCCGCAGAAACAGAAGGAGAAGCCTGTGGTAC
GTCTAGACCTCTCATTTACACTCTGCCCTGATCAAGACGATTCCATGAATATAGATCATCATGGCAAGGCAGAGGGGACATGCTTCAAGTCTGAAGAAGA
TGAGGATTATGGAAATAAGAGAAGTGATCATTCCATTGATAATAGCTGCATGTATGGTACTGGTAGAAAGAAACTCAGGCTTACAAAAGACCAATCAAGC
TACCTTGAGGAGAGCTTTCGAAGGCATCCCACTCTTAATCCGGCTAAGAAGCATGCACTCGCTGAGCAATTGAATTTAAAGCCGAGACAAGTTGAAGTCT
GGTTTCAAAACAGGAGGGCAAGGACCAAGCTGAAGCAAACAGAGGCAGACTGTGAACTCTTGAAGAAATGTTGTGAAAGTTTGAGCAATGAGAATAGAAG
ATTGAAGAGAGAATTGCAAGAATTAAGGTCACAAAAAACTGGACGGTCGTCGTCGTCGCACTCTCAGCTTGCAAAGGATTTGGGTACAATTACAAAGTGT
CCTTCATGTGAGGAAAGCACCACAACTGATCAGAACAAAATGTAG
AA sequence
>Potri.006G071600.1 pacid=42768105 polypeptide=Potri.006G071600.1.p locus=Potri.006G071600 ID=Potri.006G071600.1.v4.1 annot-version=v4.1
MEEEGYCNTRLGLGLGGENYAPWPQKQKEKPVVRLDLSFTLCPDQDDSMNIDHHGKAEGTCFKSEEDEDYGNKRSDHSIDNSCMYGTGRKKLRLTKDQSS
YLEESFRRHPTLNPAKKHALAEQLNLKPRQVEVWFQNRRARTKLKQTEADCELLKKCCESLSNENRRLKRELQELRSQKTGRSSSSHSQLAKDLGTITKC
PSCEESTTTDQNKM
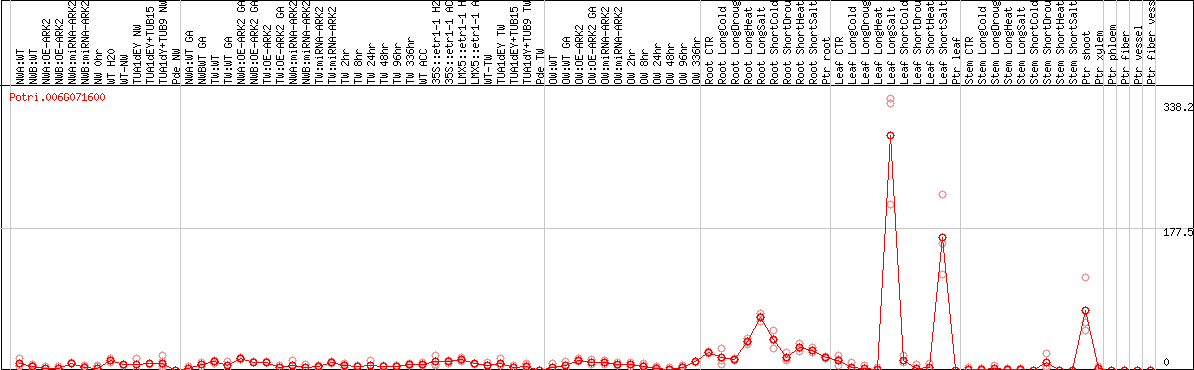
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G071600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.