External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G57150 175 / 1e-54
bHLH
bHLH035
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT4G29930 150 / 9e-45
bHLH
bHLH027
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT2G28160 83 / 2e-18
bHLH
ATFIT1, ATBHLH29, bHLH029, FIT1, ATBHLH029, FRU
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
AT2G16910 82 / 8e-18
bHLH
AMS, bHLH021
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G10610 72 / 2e-14
bHLH
bHLH090
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G12860 69 / 4e-13
bHLH
SCRM2, ICE2, bHLH033
SCREAM 2, INDUCER OF CBF EXPRESSION 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G26744 68 / 8e-13
bHLH
SCRM, ATICE1, ICE1, bHLH116
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
AT1G01260 67 / 2e-12
bHLH
bHLH013, INU4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT4G16430 64 / 1e-11
bHLH
bHLH003, INU3
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G46510 60 / 3e-10
bHLH
ATAIB, bHLH107, bHLH017, INU1, JAM1, MYL1
ABA-inducible BHLH-type transcription factor (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G141700
322 / 4e-112
AT5G57150 205 / 4e-66
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.006G074800
309 / 4e-107
AT5G57150 181 / 1e-56
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.018G141600
261 / 6e-88
AT5G57150 191 / 2e-60
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.018G141500
256 / 3e-86
AT5G57150 214 / 1e-69
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.006G074600
257 / 6e-86
AT5G57150 168 / 3e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.006G074900
226 / 3e-74
AT5G57150 200 / 3e-64
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.018G141800
217 / 1e-70
AT5G57150 256 / 6e-86
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.009G136300
85 / 1e-18
AT2G16910 461 / 1e-156
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G189600
78 / 3e-16
AT1G10610 223 / 3e-67
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006951
199 / 4e-64
AT5G57150 243 / 2e-81
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10016151
85 / 4e-19
AT2G28160 235 / 1e-75
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
Lus10030609
74 / 4e-15
AT2G16910 239 / 1e-74
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10030880
72 / 3e-14
AT2G16910 224 / 7e-69
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10021416
68 / 6e-14
AT2G28160 191 / 7e-61
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
Lus10018395
69 / 9e-14
AT4G21330 131 / 1e-37
DYSFUNCTIONAL TAPETUM 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10007618
67 / 3e-13
AT4G21330 141 / 2e-42
DYSFUNCTIONAL TAPETUM 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10036068
63 / 6e-12
AT5G57150 79 / 3e-18
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10032542
65 / 9e-12
AT3G26744 386 / 6e-129
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
Lus10043199
65 / 9e-12
AT3G26744 387 / 1e-129
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.006G074700.1 pacid=42767187 polypeptide=Potri.006G074700.1.p locus=Potri.006G074700 ID=Potri.006G074700.1.v4.1 annot-version=v4.1
ATGGAAATTATTGAACATATTGGAGAGTGCGAAAATTCCTGGGAAACTAACTGGTTTTGGAATGAAGAGCTTAATTTTAGACACAGTTGGGCAACGAATC
AGCAATTTAATCTCGGGAGCAATGATTCAAGCGCGACGGATGGAACCACTTCAACAATATTCTCTAAAAATATTGTTTCAGAGAGGAGCAGGAGGAAGAA
TCTCAGTGACAAGCTCTTAGCACTTAGAGAAGCTGTCCCCAAAATCAGCAAGATGGACAAGGCTTCTATAATCAAAGATGCTATTGATTACATCCAAGAT
TTGCAAGAACAAGAGAAGGGACTCCAAGCTGAGATAATGGAACTTGAATCGAATAGATTGAAGGAAGATCTTGGTTATGACTTTGATCAAGAGCTACCAG
TCTTGTTAAGATCCAAGAGAACAAGATATGACCAAATTTATGATCACAGGATGGCCAGAAATACTTGTCCCATTCAAGTTCATGAATTTAGTGTTACTTC
GATGGGAGGGAAAAATCTGTTCGTGAGCTTGACATGTAATAGAACAACAGATGCTATGAGTAGAATTTGTGAGGTTTTTGAATCTTTGAAGCTGAAAATT
ATCACAGCAAATATCACAACTCTCTCAGAATTGGTCAAGAAGACAGTCTTGATAGAGGTAGATGAAGAAGAGAAAGAGCATGTAAAGATAAAAATTGAAA
GAGCCTTTTCAGTTCTAAGATCCACACATGGTACCAGGATGATGTAA
AA sequence
>Potri.006G074700.1 pacid=42767187 polypeptide=Potri.006G074700.1.p locus=Potri.006G074700 ID=Potri.006G074700.1.v4.1 annot-version=v4.1
MEIIEHIGECENSWETNWFWNEELNFRHSWATNQQFNLGSNDSSATDGTTSTIFSKNIVSERSRRKNLSDKLLALREAVPKISKMDKASIIKDAIDYIQD
LQEQEKGLQAEIMELESNRLKEDLGYDFDQELPVLLRSKRTRYDQIYDHRMARNTCPIQVHEFSVTSMGGKNLFVSLTCNRTTDAMSRICEVFESLKLKI
ITANITTLSELVKKTVLIEVDEEEKEHVKIKIERAFSVLRSTHGTRMM
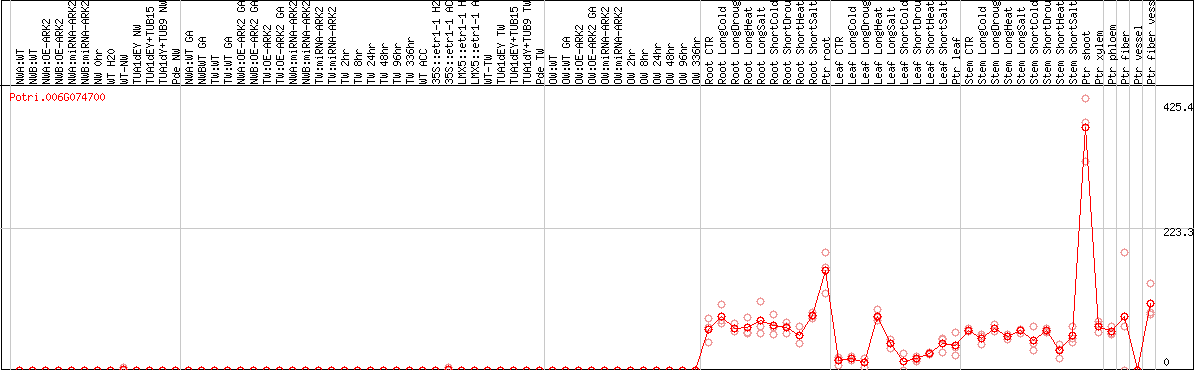
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G074700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.