Potri.006G080700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G080700.1 pacid=42767628 polypeptide=Potri.006G080700.1.p locus=Potri.006G080700 ID=Potri.006G080700.1.v4.1 annot-version=v4.1
ATGGCCGGCGATCTTTCAACCGATTCTCTTAGATTTATTATCGTCATCTCCGCCTTTTTGACACTGCTGCTAGCTACGGGTTCATATGGGTCTCCTTCTG
GAAGTCGCAAAACGGGCAAGTCTTCATTATCATCGGTGTTTTCTTTATTTAATCTCAAAGAGAAGAGTAGGTTTTGGAGTGAATCTGTTATTCACAGTGG
CGATTTCGATGATTTGGAATCTTCTAGCCCTGCCAAAATGGGACCTATCAATTTCACCGAAGCCGGTAACATTGCAAGTTATCTTAAGCTCCAAGAAGTG
GATTCCATGTACCTTCCAGTTCCTGTAAATTTCATTTTCATAGGCTTCGAAGGAAAAGGGAACCAAGCATTCAAGCTTCATTCAGAAGAAATTGAGCGTT
GGTTCACAAAAATTGATCACATCTTTGAACATACACGCGTACCAAAAATTGGAGAAGTTCTCACCCCATTTTACAAAATTTATGTAGACAAAGAGCAACA
TCACCATCTTCCCTTAGTCAGTCACATAAATTACAATTTTTCTGTTCATGCCATTCAAATGGGTGAAAAGGTTACTTATATCTTTGAGCATGCCATCAAT
CTTTTGGCTCGCAAGGATGATGTGTCTGATAACAGTGATAACAAGGATGTCCTGTGGCAAGTAGATATGGACGTGATGGATGCTCTTTTCAGTAGTCTTG
TTGATTATCTTCAACTTGACAATGCATATAACGTTTTCATTTTGAATCCCAAGCATGACCTAAAGAGGGCTAAATATGGCTACAGGAGAGGTTTGTCGGA
TTCAGAAATAACATTTCTAAAAGAGAATAAGAGCTTGCAAACAAAAATTCTTCAGTCTGGAGGCGTTTCAGAAAGTGTTCTCGCACTTGATAAGATTAAG
AGACCTCTGTATGAAAAGCATCCTATGACCGCATTTACCTGGACGATAACTGAAGAAACTGATACAGTGGAATGGTATAATATCTGCCTGGATGCATTGA
ACAATGCTGAGAAGTTATATCAAGGGAAGGACACTTCTGACATCATTCAGAATAAAGTCTTACAATTATTGAAAGGGAAGAATGAGGATATGAAGCTTCT
TCTTGAAAAAGAGTTAAAATCTGGGGGCTTCAGTGATTTTCCTGCAGAGTGTCTTACAGATACATGGATCGGAAGGGACAGGTGGGCCTTTATTGATTTA
ACTGCTGGCCCCTTCTCGTGGGGTCCTGCTGTTGGTGGAGAAGGTGTGCGCACAGAGCGTAGTTTGCCAAATGTGCAGAAAACTATTGGTGCAGTTGCTG
AGATTTCAGAAGATGAAGCTGAAGAGCGCTTGCAAGAAGCAATTCAGGAAAAGTTTTCAGTACTTGGTGATAAGGATCATCAAGCCATCGATATTCTTTT
GGCTGAGATTGATATTTATGAGCTTTTTGCTTTCAAGCACTGCAAAGGAAGGAGAGTTAAGCTAGCTCTTTGTGAAGAGCTTGATGAGAGAATGCGGGAT
TTGAAAAATGAGCTCCAGTCTCTTGACCATGAAAAGCACGATGAAAGTCATAAGAAGAAGGCCGTAGAGGCATTAAAACGAATGGAGAGTTGGAATCTTT
TCAGTGATACCCATGAGCAGGAGTTTCGAAACTACACAGTTGCAAGGGATACCTTTCTTGCACATTTAGGTGCAACCCTTTGGGGGTCTATGAGACACGT
AATCTCACCTTCACTATCTGATGGAGCATTCCACTATTACGAGAAGATATCCTTTCAGTTTTTTTTCGTCACACATGAAAAAGTTAGGAATGTTAAACAT
CTACCTGTTGATCTTGAGGCTCTAAAGAATGGGCTATCCTCCTTACTTGTATCATCACAAAAGGCTATGTTCAGTGAGAACCTAGTAGTGCTGTCAGAGG
ATCCTGCATTAGCAATGGCCTTTTCAGTGGCTCGGCGAGCAGCAGCTGTCCCATTATTGCTTGTTAATGGAACATATAGGAAAACCACTCGGTCCTATCT
TGATTCCTCCATTCTCCAGCATCAGTTGCAGAGGCATCTCCATGACCATGGTTCTCTCAAAGGAGCACATGCCCATAGTAGGTCCACACTTGAAGTTCCT
ATCTTTTGGTTCATTTATGGCGAGCCACTATTAGTTGACAAGCATTATCAGGCAAAGGCTCTTTCTGACATGGTTATTGTGGTCCAGTCAGAACCATCAT
CTTGGGAAAGCCATCTGCAGTGTAATGGGCAATCAGTTTTGTGGGATTTGAGGAGCCCTGTAAAAGCTGCGTTGGCTTCTGTCTCCGAACATCTGGCTGG
TTTACTTCCACTTCATCTTGTCTACAGCCATGCCCATGAAACTGCAATTGAGGATTGGGTATGGTCGGTGGGATGCAATCCTTTTTCAATCACTTCCAGA
GGCTGGCATATGTCACAATTTCAGTCTGATACAATTGCTCGGAGCTACATCATCACAGCGCTTGAAGAGTCAATACAACTGGTCAATGCAGCCATTCGTC
GTCTACTCATGGAACATACATCTGAAAAGACCTTCAAAATGTTTCAGTCTGAGGAGCGAGAACTTGTGAACAAGTATAATTATGTTGTTAGCCTTTGGAG
AAGGATTTCAACTATTCATGGAGAGCTGCGTTACATGGATGCAATGAGACTTCTTTATACTTTAGAGGATGCATCTGAAAGGTTTGCTAATCAAGTGAAT
GCGACTATGGCCGTTCTTCACCCAATTCACTGCATGAGAGAGAGAAAAGTACATGTGGTGATTGATATGACAACAGTCCCTGCTTTCTTGGTTGTTCTTG
GTGTTCTGTATATGGTTCTGAAGCCTAGACGACCAAAACCCAAGATTAATTAA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G080700.1 pacid=42767628 polypeptide=Potri.006G080700.1.p locus=Potri.006G080700 ID=Potri.006G080700.1.v4.1 annot-version=v4.1
MAGDLSTDSLRFIIVISAFLTLLLATGSYGSPSGSRKTGKSSLSSVFSLFNLKEKSRFWSESVIHSGDFDDLESSSPAKMGPINFTEAGNIASYLKLQEV
DSMYLPVPVNFIFIGFEGKGNQAFKLHSEEIERWFTKIDHIFEHTRVPKIGEVLTPFYKIYVDKEQHHHLPLVSHINYNFSVHAIQMGEKVTYIFEHAIN
LLARKDDVSDNSDNKDVLWQVDMDVMDALFSSLVDYLQLDNAYNVFILNPKHDLKRAKYGYRRGLSDSEITFLKENKSLQTKILQSGGVSESVLALDKIK
RPLYEKHPMTAFTWTITEETDTVEWYNICLDALNNAEKLYQGKDTSDIIQNKVLQLLKGKNEDMKLLLEKELKSGGFSDFPAECLTDTWIGRDRWAFIDL
TAGPFSWGPAVGGEGVRTERSLPNVQKTIGAVAEISEDEAEERLQEAIQEKFSVLGDKDHQAIDILLAEIDIYELFAFKHCKGRRVKLALCEELDERMRD
LKNELQSLDHEKHDESHKKKAVEALKRMESWNLFSDTHEQEFRNYTVARDTFLAHLGATLWGSMRHVISPSLSDGAFHYYEKISFQFFFVTHEKVRNVKH
LPVDLEALKNGLSSLLVSSQKAMFSENLVVLSEDPALAMAFSVARRAAAVPLLLVNGTYRKTTRSYLDSSILQHQLQRHLHDHGSLKGAHAHSRSTLEVP
IFWFIYGEPLLVDKHYQAKALSDMVIVVQSEPSSWESHLQCNGQSVLWDLRSPVKAALASVSEHLAGLLPLHLVYSHAHETAIEDWVWSVGCNPFSITSR
GWHMSQFQSDTIARSYIITALEESIQLVNAAIRRLLMEHTSEKTFKMFQSEERELVNKYNYVVSLWRRISTIHGELRYMDAMRLLYTLEDASERFANQVN
ATMAVLHPIHCMRERKVHVVIDMTTVPAFLVVLGVLYMVLKPRRPKPKIN
|
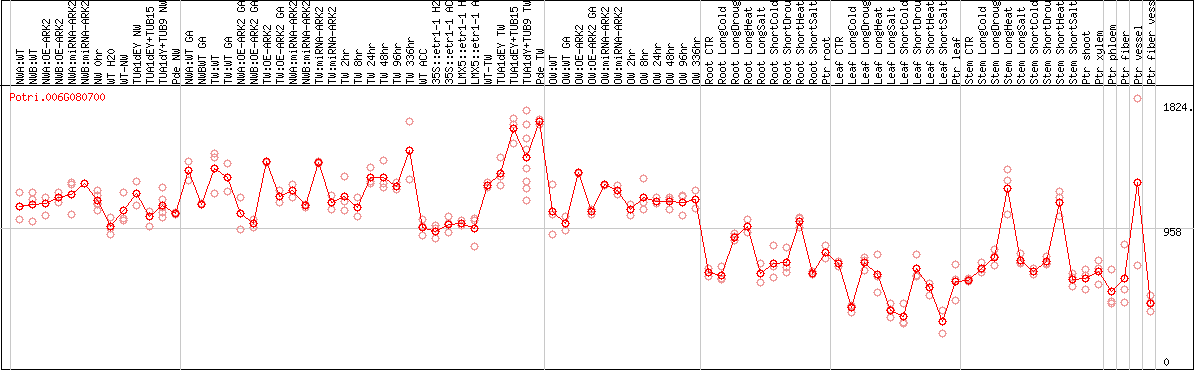
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G080700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
