Potri.006G083600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G083600.8 pacid=42767405 polypeptide=Potri.006G083600.8.p locus=Potri.006G083600 ID=Potri.006G083600.8.v4.1 annot-version=v4.1
ATGGTTTATGTGGATATATCAGCAGCAGCAGCAGCAGCAGCAGATGTGAGTATGGTAGCTGCAAAAGGTGATTTGAAGAGAGAGCGTAATCCATTTCCGG
TGAATGAGAATGAGACCGGGGAATTATTTCCTAACAAGAAACAAGCAAAACAAGAAGAAGCTTCAAACGACGACACAAAATCTGAAGTTTCGAATCCTGT
AATAACACTAGTTTCTCCTAAAGGCAACGGATCATCATCCCATGATATCTCTGAAGAATCGCCTACGAATGCGTGCCCCTCTTCAGAGGAGACTCTGACT
GTGAGCCAGGAAGGAGGTGGTAGCAGCAGTGAAGATAACACCTCCAATCAAAGTCCTCGAAATGATACTTGTGATTCTGTCTCTATGTCTCCTGTTGTGT
TGGAGATTCCTGAGCATGCTAGTACCACGGGAGTTAGGAAGATTACTTTTAAGTTCAGCAAAAGAAAAGAGGATTATGATACCAAAATTTCGCCTCACCC
TCTACATGGTGGAATTGATCAAGGACTGCTGTATCATCGTAATGGGGATTATTATCCAAGGAACCACTCTGTTTGGGTTAATTCCTGCACAGAAATGCCT
CAGACCAGAGAAAGATATATGGAGTTGAACATGTCTAAGAAGGTCGTTCCCAACAACTACCCTACGAATGTCAAAAAGTTGTTAGCAACTGGTATTCTCG
ATAGGGCTCGAGTCAAGTATATTTGTTTTTCTTCAGAGCGAGAGCTTGATGGAATTATAGACGGCGGTGGATATTTGTGTGGCTGTTCGTCATGTAATTT
CTCCAAAGTCCTTAGTGCATATGAATTTGAGCAGCATGCTGGTGCTAAAACTAGACATCCAAACAATCACATATACTTGGAGAATGGGAAACCAATTTAT
AGCATTATTCAAGAACTAAAAACGGCTCCACTCAGCATGATAGATGGAGTCATAAAAGACGTGGCTGGTTCATCTATCAATGAGGAGTTCTTCCGGGTTT
GGAAAGCTAGTCTTAATCAAAGCAATGCACTTGTTGGAGCAGATAAAAAATGTCACAGCGAACTGCCTTGTTTGCCTCATTCTCATGTAAGCTACGCTAG
TCAAGCTTTGAAAGAAAGCTTTTGCCCAATTTCAAGCTCCTTTTTGTATAATAACAACTTTGTCAGCCAGCAAATGTATATGGAGACTTCAGGCGTGAAT
AAGCAAACATCGAAGAGGCCTAGCCTTTACTTCCCTGGCTCTGCTACGAAGCAAAAGAAAACAGCTGAGAGTGGTGTCAGGAAGAGGGATAATGATCTTC
ACAGGTTACTCTTTATGCCGAATGGTCTTCCAGATGGGACCGAATTGGCTTACTATGTCAAAGGGCAGAAAATACTAGGGGGTTACAAGCAGGGTAATGG
TATTGTCTGTAGTTGTTGTGAAGTAGAGATTAGCCCTTCACAGTTTGAGTCTCATGCTGGAATGTCTGCCAGGCGTCAACCCTACCGTCACATTTATACT
TCTAATGGACTGACCCTCCATGATATAGCTATTTCGTTGGCAAATGGACAAAACATTACCACAGGTATTGGTGATGATATGTGTGCAGAATGTGGAGATG
GAGGGGATCTAATGTTTTGTCAGAGTTGCCCTCGTGCTTTTCATGCAGCATGTTTGGATTTACATGATACTCCTGAAGGTGCTTGGCATTGTCCAAATTG
CAATAAACTTGGCCACGGAGGCAATTTTGCAAGACCTATTGTCATCCGGCTGACTCGAGTTGTTAAAACTCCAGAATATGATGTTGGTGGTTGTGCTGTT
TGCAGGGCCCATGATTTTAGTGGCGATACATTTGATGACCGAACAGTTATCCTTTGTGACCAATGCGAAAAGGAATTTCATGTTGGTTGCTTGCGGGAAA
GTGGACTATGTGATCTAAAAGAAATCCCCAAAGATAATTGGTTCTGCTGTCAAGACTGCAATAATATCTATGTAGCTCTCCGGAATTCTGTTTCTACTGG
AGTGCAGACAATTCCCGCTTCGCTGTTAAATATCATAAATAGAAAGCATGTCGAGAAAGGGCTACTTGTTGATGAAGCTGCATATGATGTGCAGTGGCAA
ATTCTGATGGGAAAGAGTCGCAACCGAGAAGATCTTTCATTGCTTTCAGGGGCTGCTGCAATTTTTCGAGAGTGTTTTGATCCTATTGTTGCAAAAACTG
GCCGTGATCTGATTCCTGTTATGGTCTATGGGAGAAATATATCTGGCCAGGAATTTGGGGGCATGTATTGTGTGCTTTTGACTGTAAGGCATGTTGTTGT
GTCTGCTGGTCTTCTTAGGATTTTCGGTAGGGAGGTTGCCGAGCTTCCTTTAGTGGCCACAAACAGAGAGCATCAAGGGAAAGGTTATTTCCAAGCATTA
TTCTCGTGCATTGAGAGGTTATTGTGCTCCCTGAATGTGGAACAACTGGTGCTCCCAGCTGCTGAGGAAGCAGAATCAATCTGGACAAGAAGATTTGGCT
TCAGAAAGATGAGTGAGGGACAATTGTTGAAGTATACAAGGGAGTTTCAGCTGACGATTTTCAAGGGAACATCCATGCTCGAGAAGGAAGTCCCACGGAT
AATTGATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G083600.8 pacid=42767405 polypeptide=Potri.006G083600.8.p locus=Potri.006G083600 ID=Potri.006G083600.8.v4.1 annot-version=v4.1
MVYVDISAAAAAAADVSMVAAKGDLKRERNPFPVNENETGELFPNKKQAKQEEASNDDTKSEVSNPVITLVSPKGNGSSSHDISEESPTNACPSSEETLT
VSQEGGGSSSEDNTSNQSPRNDTCDSVSMSPVVLEIPEHASTTGVRKITFKFSKRKEDYDTKISPHPLHGGIDQGLLYHRNGDYYPRNHSVWVNSCTEMP
QTRERYMELNMSKKVVPNNYPTNVKKLLATGILDRARVKYICFSSERELDGIIDGGGYLCGCSSCNFSKVLSAYEFEQHAGAKTRHPNNHIYLENGKPIY
SIIQELKTAPLSMIDGVIKDVAGSSINEEFFRVWKASLNQSNALVGADKKCHSELPCLPHSHVSYASQALKESFCPISSSFLYNNNFVSQQMYMETSGVN
KQTSKRPSLYFPGSATKQKKTAESGVRKRDNDLHRLLFMPNGLPDGTELAYYVKGQKILGGYKQGNGIVCSCCEVEISPSQFESHAGMSARRQPYRHIYT
SNGLTLHDIAISLANGQNITTGIGDDMCAECGDGGDLMFCQSCPRAFHAACLDLHDTPEGAWHCPNCNKLGHGGNFARPIVIRLTRVVKTPEYDVGGCAV
CRAHDFSGDTFDDRTVILCDQCEKEFHVGCLRESGLCDLKEIPKDNWFCCQDCNNIYVALRNSVSTGVQTIPASLLNIINRKHVEKGLLVDEAAYDVQWQ
ILMGKSRNREDLSLLSGAAAIFRECFDPIVAKTGRDLIPVMVYGRNISGQEFGGMYCVLLTVRHVVVSAGLLRIFGREVAELPLVATNREHQGKGYFQAL
FSCIERLLCSLNVEQLVLPAAEEAESIWTRRFGFRKMSEGQLLKYTREFQLTIFKGTSMLEKEVPRIID
|
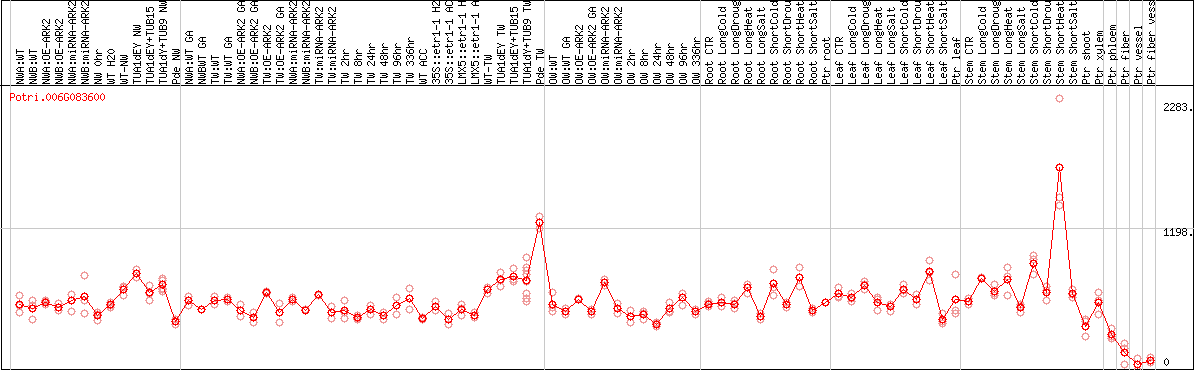
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G083600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
