External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G37660 451 / 2e-160
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G02240 406 / 6e-144
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G31530 92 / 3e-21
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G34460 81 / 4e-17
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G18890 70 / 4e-13
AtTic62
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G16720 54 / 7e-08
HCF173
high chlorophyll fluorescence phenotype 173 (.1)
AT4G18810 49 / 3e-06
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G80820 44 / 6e-05
CCR2, ATCCR2
cinnamoyl coa reductase (.1)
AT1G15950 42 / 0.0003
IRX4, ATCCR1, CCR1
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
AT2G33590 42 / 0.0003
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G253900
80 / 8e-17
AT4G31530 399 / 1e-139
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G150300
73 / 4e-14
AT3G18890 481 / 1e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.T124404
71 / 2e-13
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G111323
71 / 2e-13
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.011G079600
64 / 2e-11
AT2G34460 370 / 6e-130
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.014G000400
55 / 3e-08
AT1G16720 863 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Potri.007G003500
52 / 3e-07
AT1G16720 852 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Potri.003G063800
50 / 1e-06
AT1G72640 348 / 1e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G183100
49 / 2e-06
AT4G35250 650 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010846
466 / 1e-166
AT2G37660 451 / 2e-160
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10024383
464 / 7e-166
AT2G37660 458 / 1e-163
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10009715
86 / 4e-19
AT4G31530 408 / 1e-144
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10022420
86 / 7e-19
AT4G31530 415 / 4e-146
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10040884
74 / 1e-14
AT2G34460 401 / 7e-142
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004936
73 / 2e-14
AT2G34460 399 / 4e-141
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10042346
67 / 6e-12
AT3G18890 483 / 2e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026321
64 / 5e-11
AT3G18890 479 / 6e-163
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10034781
55 / 4e-08
AT1G16720 870 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Lus10033322
55 / 4e-08
AT1G16720 874 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF13460
NAD_binding_10
NAD(P)H-binding
Representative CDS sequence
>Potri.006G086800.1 pacid=42769567 polypeptide=Potri.006G086800.1.p locus=Potri.006G086800 ID=Potri.006G086800.1.v4.1 annot-version=v4.1
ATGACAACCACCGTCACCGTGACACGTGTTCCATTGATTTTCTCCGGTCATTTTCTTAATAATAAAAACGGGCGTGCTTCACAATCGGCCGTACACCTTT
TAGGTAGCATTAGCAATAGCAATACTAGCAGTGCTAGTCGTAGCTCATTTTCTTCGTCACTAACAGCAACAACGCGCATAAGGAGAGCAGGGTTCGCTGT
ATTAGCTTCAAATTCAATGGCTCCAAGCATTGTCCTTGTCACTGGAGCTGGTGGCAGAACTGGTTCCATAGTTTATAAGAAACTGAAGGAGAGGTCTGAA
CAATATGTTGCAAGGGGTTTGGTAAGAACTGAAGAGAGCAAGGAGAAAATTGGAGGAGCTGAGGATGTTTTTGTTGGGGATATAAGGGAATCCAAAAGCA
TTGTTCCTGCGATCCAAGGAATTGATTCTCTGATCATCCTGACAAGTGCAGTGCCAAAGATGAAACCTGGGTCTGATCCATCCAAAGGCAGGCCTGAGTT
CTATTTTGAAGATGGAGCATTTCCTGAGCAGGTTGACTGGATTGGTCAGAAAAATCAAATAGATGCTGCTAAGGCCGCAGGAGTTAAGCAAATTGTGCTG
GTTGGATCTATGGGTGGAACAAACCTAAATCATCCCTTGAACAGTTTGGGTAATGGGAATATACTGGTTTGGAAGAGAAAGGCTGAGCAGTATTTGGCCG
ACTCTGGTGTTCCATACACAATTCTAAGGGCTGGAGGCCTGCAAGACAAAGAAGGAGGTGTCCGAGAATTACTTGTTGGAAAAGATGACGAGCTTCTTCA
GACAGAGACAAGAACCATTGCCAGAGCTGATGTTGCAGAAGTCTGCATTCAGGCACTACAGTACGAAGAGGCTCAATTTAAGGCATTTGATCTGGCCTCA
AAACCAGAGGGAACTGGCACCCCAGCCAATGATTTTAAGGCTCTCTTTTCTCAAGTCACTGCTCGTTTTTGA
AA sequence
>Potri.006G086800.1 pacid=42769567 polypeptide=Potri.006G086800.1.p locus=Potri.006G086800 ID=Potri.006G086800.1.v4.1 annot-version=v4.1
MTTTVTVTRVPLIFSGHFLNNKNGRASQSAVHLLGSISNSNTSSASRSSFSSSLTATTRIRRAGFAVLASNSMAPSIVLVTGAGGRTGSIVYKKLKERSE
QYVARGLVRTEESKEKIGGAEDVFVGDIRESKSIVPAIQGIDSLIILTSAVPKMKPGSDPSKGRPEFYFEDGAFPEQVDWIGQKNQIDAAKAAGVKQIVL
VGSMGGTNLNHPLNSLGNGNILVWKRKAEQYLADSGVPYTILRAGGLQDKEGGVRELLVGKDDELLQTETRTIARADVAEVCIQALQYEEAQFKAFDLAS
KPEGTGTPANDFKALFSQVTARF
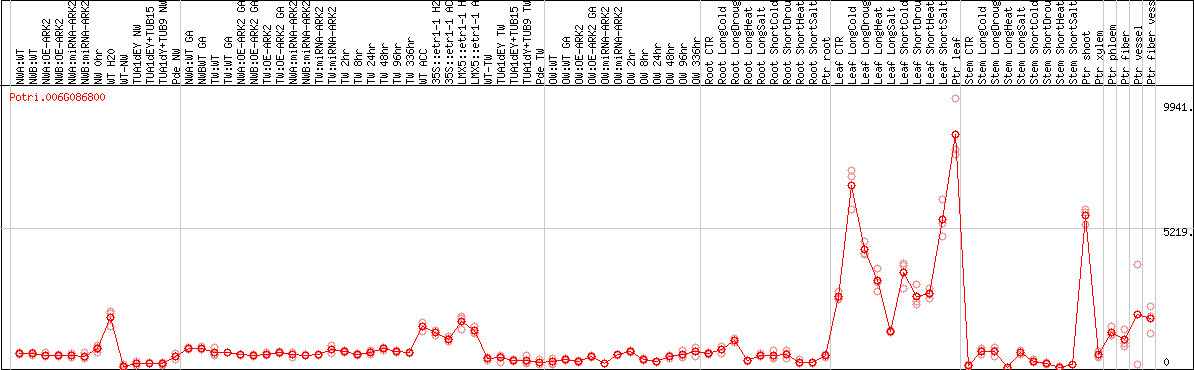
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G086800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.