Potri.006G105700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G105700.1 pacid=42769935 polypeptide=Potri.006G105700.1.p locus=Potri.006G105700 ID=Potri.006G105700.1.v4.1 annot-version=v4.1
ATGTCTTCCTTAATTCTCAACAAGTCAGCACACATCAAAATCCGCCTCTTTCAAGGCTTTAACTCTCTCAAACACCTAAAACATGTCCACGCAGCCCTTC
TCCGCCTCGGCCTCGACGAAGACAGTTACCTCCTTAACAAGGTCTTACGTTTCAGCTTCAACTTTGGCAACACAAACTATTCCCATCGCATATTTCATCA
AACCAAAGAACCTAATATTTTCCTTTTCAATACAATGATCCATGGCTTAGTTTTGAATGATTCTTTCCAAGAATCCATTGAAATTTACCACTCAATGAGA
AAAGAAGGGCTTTCTCCTGATAGTTTTACTTTCCCTTTTTTGTTGAAAGCGTGTGCCAGGCTTTTGGATTCTAAATTGGGTATTAAGTTGCATGGCCTTG
TGGTTAAAGCGGGTTGTGAATCGGATGCGTTTGTTAATACTAGTTTGGTCAGTTTGTATGGGAAATGTGGGTTTATAGACAATGCATTTAAGGTGTTTGA
TGATATTCCCGAGAAGAATGTTGCTGCGTGGACGGCGATAATAAGTGGGTATATTGGTGTTGGGAAATGTAGGGAGGCGATTGATATGTTTCGTAGGTTG
TTGGATATGGGTTTGAGGCCGGATAGTTTCAGTCTTGTTCGGGTTTTGGCTGCCTGCACACGAATTGGGGATTTGAGAAGTGGAGAGTGGATTGATGATT
ACATAACGAAGATTGGCATGGTAAGGAATGTGTTTGTGGCTACTTCTTTGGTGGATTTTTATGTCAAATGTGGAAACATGGAGAGAGCATGTAGTGTCTT
TGATGGGATGCTGGAGAAGGATATTGTTTCTTGGAGTTCTATGATTCAGGGTTATGCTTCCAATGGGCTGCCTAAAGAAGCTTTAGACCTCTTTTTTAAA
ATGTTGAATGAGGGTTTCAGACCTGATTGTTATGCTATGGTGGGGGTTCTTTGTGCTTGTGCGAGGTTAGGAGCACTAGAATTAGGGAACTGGGCTAGCA
ATTTGATGGATAGAAATGAGTTCTTGGGCAATCCTGTCTTGGGTACGGCATTGATTGACATGTATGCAAAATGTGGGAGAATGGATAGCGCCTGGGAAGT
TTTCAGGGGAATGAGGAAGAAAGATATAGTAGTTTGGAATGCAGCTATATCAGGTCTTGCCATGAGTGGACATGTCAAAGCAGCATTTGGACTTTTTGGC
CAAATGGAGAAATCTGGGATTGAGCCGGATGGGAACACTTTTGTTGGCCTACTTTGTGCTTGTACTCATGCGGGTCTAGTTGACGAGGGTCGTCAATATT
TTAATAGTATGGAACGTGTTTTTACCTTGACTCCAGAAATCGAACACTATGGATGCATGGTGGATCTCCTGGGCCGTGCAGGTTTCTTAGATGAAGCTCA
TCAGCTTGTCAAAAGTATGCCAATGGAGGCCAATGCTATTGTTTGGGGAGCATTGCTGGGTGGATGCAGGCTGCATAGAGATACCCAATTGGTGGAAGGT
GTGCTGAAACAGCTCATAGCATTAGAACCGTCCAACTCAGGTAATTACGTTCTTTTGTCAAATATATATTCAGCAAGTCATAAATGGGAAGATGCAGCAA
AGATTAGGTCAATCATGAGTGAGAGGGGGATTAAGAAAGTACCAGGGTATAGTTGGATTGAAGTGGATGGAGTTGTTCATGAATTCCTTGTGGGGGATAC
ATCCCATCCCTTATCGGAGAAAATATATGCAAAACTTGGCGAATTGGTCAAGGACTTGAAAGCATCAGGTTATGTTCCCACAACAGATTATGTGCTGTTT
GATATAGAAGAAGAGGAGAAGGAGCATTTCATTGGTTGTCACAGTGAGAAGCTGGCTATTGCCTTTGGTTTGATAAGCACAGCTCCAAATGACAAAATTC
GTGTTGTGAAAAATCTTCGTGTTTGTGGTGATTGTCATGAGGCAATAAAGCACATTTCTAGGTTTACAGGTAGAGAGATAATTGTGAGAGATAACAACCG
ATTCCATTGTTTTAATGATGGTTCTTGTTCATGTAAAGATTATTGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G105700.1 pacid=42769935 polypeptide=Potri.006G105700.1.p locus=Potri.006G105700 ID=Potri.006G105700.1.v4.1 annot-version=v4.1
MSSLILNKSAHIKIRLFQGFNSLKHLKHVHAALLRLGLDEDSYLLNKVLRFSFNFGNTNYSHRIFHQTKEPNIFLFNTMIHGLVLNDSFQESIEIYHSMR
KEGLSPDSFTFPFLLKACARLLDSKLGIKLHGLVVKAGCESDAFVNTSLVSLYGKCGFIDNAFKVFDDIPEKNVAAWTAIISGYIGVGKCREAIDMFRRL
LDMGLRPDSFSLVRVLAACTRIGDLRSGEWIDDYITKIGMVRNVFVATSLVDFYVKCGNMERACSVFDGMLEKDIVSWSSMIQGYASNGLPKEALDLFFK
MLNEGFRPDCYAMVGVLCACARLGALELGNWASNLMDRNEFLGNPVLGTALIDMYAKCGRMDSAWEVFRGMRKKDIVVWNAAISGLAMSGHVKAAFGLFG
QMEKSGIEPDGNTFVGLLCACTHAGLVDEGRQYFNSMERVFTLTPEIEHYGCMVDLLGRAGFLDEAHQLVKSMPMEANAIVWGALLGGCRLHRDTQLVEG
VLKQLIALEPSNSGNYVLLSNIYSASHKWEDAAKIRSIMSERGIKKVPGYSWIEVDGVVHEFLVGDTSHPLSEKIYAKLGELVKDLKASGYVPTTDYVLF
DIEEEEKEHFIGCHSEKLAIAFGLISTAPNDKIRVVKNLRVCGDCHEAIKHISRFTGREIIVRDNNRFHCFNDGSCSCKDYW
|
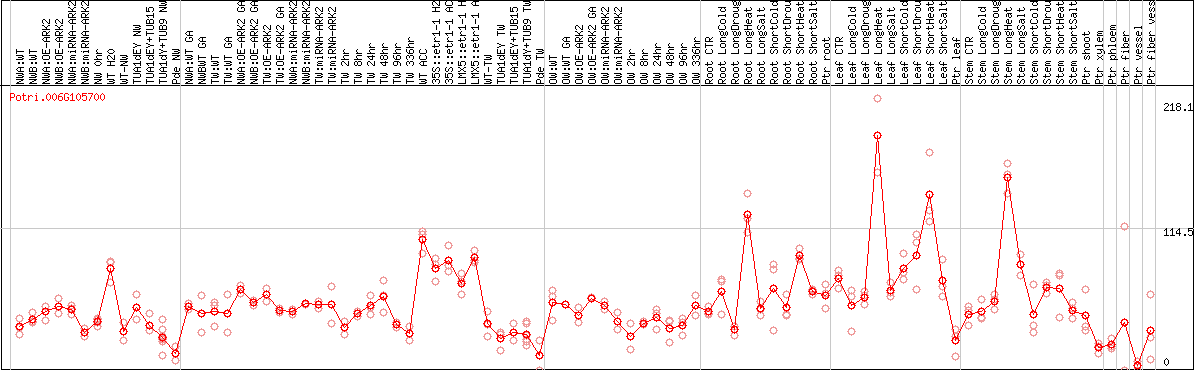
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G105700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
