WAVE4.1 (Potri.006G105800) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | WAVE4.1 | ||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G105800.2 pacid=42767800 polypeptide=Potri.006G105800.2.p locus=Potri.006G105800 ID=Potri.006G105800.2.v4.1 annot-version=v4.1
ATGCCGTTGACGAGATATCAGATAAGGAACGAATACCGTTTAGCCGATCCAGAGTTGTTTAAAGCAGCTGATAAAGATGATCCAGAAGCGCTTTTGGAAG
GTGTCGCCATGGCTGGCCTCGTCGGCGTCTTGCGCCAGCTCGGTGACCTCGCCGAGTTCGCTGCTGAGATTTTCCATGACCTGCATGAGGAAGTAATGAC
AACTGCAGCAAGAGGCCATGGGCTGATGGCTCGTGTACAACAGCTCGAGGCAGAATTTCCTTCAATTGAGAAGGCATTTCTCTCCCAAACTAATCATTCA
CCATTCTTTTCCAGCTCAGGTGCTGACTGGCATCCTAATTTACAAATGGAGCAGAATCTAATTACTCGTGGAGGCTTACCTCACTTTGTAATGGACTCTT
ATGAAGAATGCCGTGGCCCACCACAGTTATTCCTTTTGGATAAGTTTGATGTTGCGGGGGCTGGGGCATGTTTGAAACGTTATACCGATCCATCATTCTT
TAAAGTTGAAGCAGCATCCTCTGGAATAGCAACTGTAGAAGTTCAGAGGGGGAAGAAAATTCGAAAGAAGAAGAAAGGATCACGGTATAAAAATGGAGAA
ACTCCTGAAGTTGTACCAACATCACATGCCAAATTGCATGAGTTGTTTTTGGAGGAGCGTTCTGAAAATGGCCATTCTGACCCTGCACGGCTAGTGAAAT
TAAAGAGAAGGCTGTTCAACGGATCTCCATTTGATCTGAAACCTGGGAAGAGTTACATGCAGAAATTTGTGCTTACTCCTTCACCAGATCGTAAACAAGT
TTGTGAAGATTCTGTCACTCGTTCACCTTTGAAGTTGACATTAGATAATTCTAGTGAATCAAGGTATGAAATACATGAGGTCAGTGTGGCAAGCCCTGTG
AAGCAATCATCACATGGAGGGGAAAGCACATCTTCATCACCCAGTGAACGGGAGGCCACGCTTAAAACTTTTATGGACGAGTTGAATGGAGAGCCTGTTG
ATAGTAGAATAATTAAGGTACTTAACCCAATTGTTGATCGTGAGATGGATGAATATCCCCTTATTGTCCAAAAGATGGTGATTGAAGAGGAATCATCAGT
TGATGCAGATGGAAAAGCAGAGGGCACTGTAGATGGGGATCATTCTGATGACATGACAAGCGAGGTGGAAAACTACATGGATGCTCTTACTACCATGGAT
TCTGGAATGGAAACAGACAATGAGTATAAACCTATGAATGGTCAGGACTTCATGGATGTGAGGGCACACGGTGCAGATTCTGATGCGAACGAGGAACAGC
TGGATGCTCAAGCTAATTTTTCAGATTCTCAATCCATTGGAAACTCTTCTCTGTCAGAGGGTGGGAACAGTTCGTTTAAGAAAGGAACGTCTAGCTTTTC
TTACTCTGACACTCTCAGCAATGTGGCTGAGAATACAGCATCTGATGGTGAAGGAGCAGGTAAATGGTTCCCTTCTATCTCTTCTACTGAAAACTATCCA
AGAGACATTGCTGATTTGCCATCAGATAGTCCATCTGTATTTGTGGAGAGTGGCATAACTGAATCCCATCACCTTGTAACATTCAATGATACCGAGGAAG
ACAAAATCCCTGATTCTGGTGAAGCTTCTCGTAGTTCATGTCTTACAGATTGGAATCTTGTGTTCCTGCATGCAGCTCCTGTTGCAGGTTCCATGGTAAG
CCCCCTGGCAGGACCTGAACTAGATGAGGCATCCTCTGGCAGTATTGAACCTGGCTCTGAATCACCAAACTCCGATAGGAATGGTTTAAACCTGGCTGAT
TTCCCTTCGCAACTGGGGCATGATACTTCACTTACAGATTCTTCTAAAACTCATTCTGTTGGTGAGTTGGATCATGAAGATCAAAAAATGTTAACCGATG
CTGTTGTGCTTGTGTCAAATGTGTCAGATCTGGCTTTTGAAAAGAAAGGCAGTGATGATTCTGTGAATGGAGTGCTTCAGACAGATTATGCAGCTGAACA
TTCAACTATGACACCGGCAGAGGAGCGATTTCCTAAGTCAACTTTGCCTGTAGTCGAGTTGGATTCAGGGGTCCTATCTCTGCCTGATAATTTGGATTTT
GTAAAGCCTGACGTTTTAGTTTCTGAAGTTGACGATGCAATTGCAACAAGGGAGACTAGAGCAGAAAATTTAACACTAGTGGTGGATACTTCAGAGACCG
AATGCGTTAGCGAGCATCACTTCTCTGATATGACTATTGATGCTTCACAATTGGAACTTGACTCATCTAAATTAGGAGTTCCATGTTCTGAAGTGAATAT
AAATCTTGAAGAAATTCCCAACGGTTTTGATGCTGAGGAAAACATTGCATTCACAAAGGTGGATATTACAAGAGGAGATGCTGCTTCATTCGAACATCAG
TCATTGAGTTCAGATAAACCAATACTTGAAGATCATGTAAATTTGGATGATGCTGTAACTGAAACTGGTCAAGCAGAAGATATGGCTGTATCTAGTGCTG
CTTCAAGTGGTGCGAACAATGAAGATGTCAGCAATGTGATTTGTCCCTCTTCAGAGTTAGTTTGTTCCCCGCCTAGGAATGCTACAGAGCCACTAGAAGC
TCTTTCCATCCCTGAGGATCCACATCTGACAAGGTTGGACCTCGATGAGGTAATTTCCGCCAAACCTTTGTCAGAATCACAGGTGCAGATGGAAGTTACT
TCTATAGATTGGGATTCTAACCCATATAAACCAGTTTCTGAAGACCATCCAAATCAAGAGGTAAGTGAAGTTCATAACCTGTCTTTGGAACTGAGCAACC
AGGAGTCAGAAACAAAAGATAACCATCAGCATCATTATGCGGAGGCTAGCGACAACACTGTTTGTTTGCCTTTGTGTTATCTACCAGAGTCGGGAAATAC
TCTAGAACAATCAACTGAGGTGCAAGATGATCAGTTTAGTGCAGAGTCCAGTCATGCAGACAACACAAATACTTTGTTGAGTAGTCAAACTTCATCTACT
GGGTACCTTGTAGGAACTGGAATTCCTCTGGAGCACACATTGGAATTGCAATCTGATCAACTAGACAGGGGATGCTTGAAATTAGGTGAAGCAAGTTCTA
TATCAACAGATCTTCAGTCTGAAAGTTCCTGTTTGAAAGATCTCTCCAGTCAAGAGCATTTACTGCAATCATTCTGTCAGGAACGTAATGCAACTGTTCT
GGAAACAAATCCATTTGATTCTGCTTTTCCTAGCTTTGGTGTACTCCCTGTCCCTGAGGCAAGTCAGGTCTATCCAGAGGCAATGCCACCATTGCCACCT
CTACCTCCTATGCAATGGAGGTTAGGGAAAATTCAACCTGCTTCACTAGATGCAGACAGAGATATGATTGATAACAGTGAAGGCACTTTCCCACTGATCC
AGCCATTTATGGTTGATCAGCAGGTCCATTTTGATTTCCCATCATTAGATAGAGAGATCGCGCACCCTTCAAACCCATTTCTGTCTCTCCCTGTTGAAGA
AAGTCGGATGTTTCCACACTCGACTACAGAGTCAATGGGAAATTCCTTGCTTCCAACTCCACTCTTGTCAGAAACACCAATCATTGACAATGATGCACAC
TGTCAACAGGATCACCTCCGATCAGACACAACACAATCTGTGAGCTCATCTTTGGCTTTACCAGAAATGTCCGATGAGAGGCATGAGCATGGTTTCCTTC
CATTAGGAGGAGAATCCGCACAATCTAGTTCCAATCCATTCTCACTAGAGCCAAACATAGAACATACGACTGCTGTAAATGATCCCATGCCAACACAGGG
CCTGCCAATCCATCCATTTAATCAATCAGCACCCAAAACAGGATTAGACATGAAGTTTCCTGGACAGAGTTCACAAAGTTCAGAAGAGGAACTAGGGAAT
TCTTATGGTAAATCTGCAGCGCCACTAACTATGGAAGAAGAGCCGCATCATGATTTTGTAACTTCACAGGGATTAACAATGTGGCCGCCGACTGCATTAG
CTATGACACCACCAACCTCTGAAGTTGGAAAGCCAAATGGAAACAAGATCCCTCGTCCTAGAAATCCTCTCATTGATGCTGTTGCTGCACATGATAAAAG
CAAGTTGAGAAAGGTTGCTGAACTAGTTCGACCTCAGGTTGGACCAAAAGTAGAAGAAAGAGATTCACTTCTGGAACAGATACGAACCAAGTCCTTTAAC
TTGAAGCCTGCAACCGTGACAAGGCCTAGCATTCAGGGTATTCAGGGTCCAAAAACCAACTTGAAGGTCGCAGCCATCTTAGAGAAAGCAAATGCAATTC
GCCAGGCATTGACTGGAAGCGATGAAGATAACGATTCAGACAGCTGGAGTGATTCTTAG
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G105800.2 pacid=42767800 polypeptide=Potri.006G105800.2.p locus=Potri.006G105800 ID=Potri.006G105800.2.v4.1 annot-version=v4.1
MPLTRYQIRNEYRLADPELFKAADKDDPEALLEGVAMAGLVGVLRQLGDLAEFAAEIFHDLHEEVMTTAARGHGLMARVQQLEAEFPSIEKAFLSQTNHS
PFFSSSGADWHPNLQMEQNLITRGGLPHFVMDSYEECRGPPQLFLLDKFDVAGAGACLKRYTDPSFFKVEAASSGIATVEVQRGKKIRKKKKGSRYKNGE
TPEVVPTSHAKLHELFLEERSENGHSDPARLVKLKRRLFNGSPFDLKPGKSYMQKFVLTPSPDRKQVCEDSVTRSPLKLTLDNSSESRYEIHEVSVASPV
KQSSHGGESTSSSPSEREATLKTFMDELNGEPVDSRIIKVLNPIVDREMDEYPLIVQKMVIEEESSVDADGKAEGTVDGDHSDDMTSEVENYMDALTTMD
SGMETDNEYKPMNGQDFMDVRAHGADSDANEEQLDAQANFSDSQSIGNSSLSEGGNSSFKKGTSSFSYSDTLSNVAENTASDGEGAGKWFPSISSTENYP
RDIADLPSDSPSVFVESGITESHHLVTFNDTEEDKIPDSGEASRSSCLTDWNLVFLHAAPVAGSMVSPLAGPELDEASSGSIEPGSESPNSDRNGLNLAD
FPSQLGHDTSLTDSSKTHSVGELDHEDQKMLTDAVVLVSNVSDLAFEKKGSDDSVNGVLQTDYAAEHSTMTPAEERFPKSTLPVVELDSGVLSLPDNLDF
VKPDVLVSEVDDAIATRETRAENLTLVVDTSETECVSEHHFSDMTIDASQLELDSSKLGVPCSEVNINLEEIPNGFDAEENIAFTKVDITRGDAASFEHQ
SLSSDKPILEDHVNLDDAVTETGQAEDMAVSSAASSGANNEDVSNVICPSSELVCSPPRNATEPLEALSIPEDPHLTRLDLDEVISAKPLSESQVQMEVT
SIDWDSNPYKPVSEDHPNQEVSEVHNLSLELSNQESETKDNHQHHYAEASDNTVCLPLCYLPESGNTLEQSTEVQDDQFSAESSHADNTNTLLSSQTSST
GYLVGTGIPLEHTLELQSDQLDRGCLKLGEASSISTDLQSESSCLKDLSSQEHLLQSFCQERNATVLETNPFDSAFPSFGVLPVPEASQVYPEAMPPLPP
LPPMQWRLGKIQPASLDADRDMIDNSEGTFPLIQPFMVDQQVHFDFPSLDREIAHPSNPFLSLPVEESRMFPHSTTESMGNSLLPTPLLSETPIIDNDAH
CQQDHLRSDTTQSVSSSLALPEMSDERHEHGFLPLGGESAQSSSNPFSLEPNIEHTTAVNDPMPTQGLPIHPFNQSAPKTGLDMKFPGQSSQSSEEELGN
SYGKSAAPLTMEEEPHHDFVTSQGLTMWPPTALAMTPPTSEVGKPNGNKIPRPRNPLIDAVAAHDKSKLRKVAELVRPQVGPKVEERDSLLEQIRTKSFN
LKPATVTRPSIQGIQGPKTNLKVAAILEKANAIRQALTGSDEDNDSDSWSDS
|
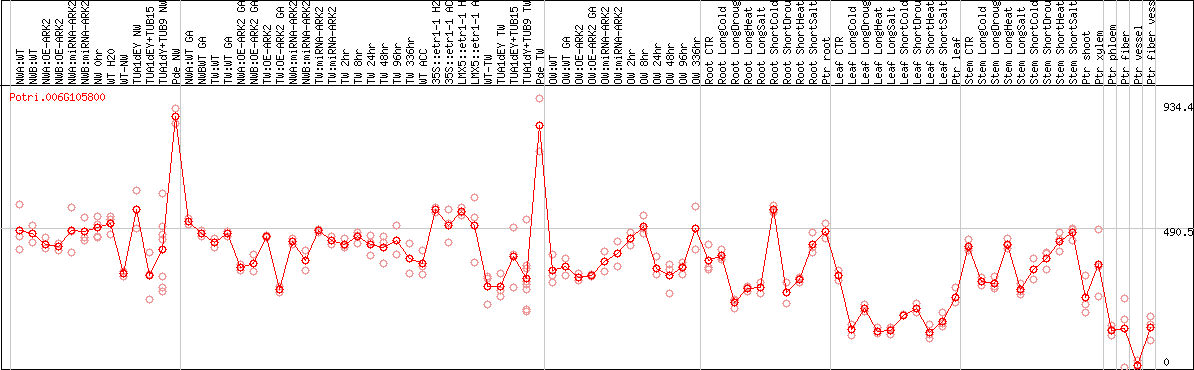
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G105800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
