Potri.006G106600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G106600.1 pacid=42769615 polypeptide=Potri.006G106600.1.p locus=Potri.006G106600 ID=Potri.006G106600.1.v4.1 annot-version=v4.1
ATGGCATTGGGAGATTTAATGGCGTCAAGATTCTCCTCACAATCTCCAGTTGCTTTTGTATCGAACCATTATGATCATTATCCGTCAAGTCACGAAGATG
ATGCTATTGATGTAGCCAGAAGAGATGACAATAATAATAGTAATAATAATAGAGATAGAGATTCTGATACTGCTAGCACTAGTAATTACGGCGGAGGCAA
TGCCACGGCAAGCACCACCGCCACAACCACTACTAGTACGGCGTATTTGCCGCAGACTGTTGTTTTGTGTGAGCTTAGACACGAGGCATTTGAAGCTTCT
GTGCCTACCGGTCCGTCTGATAGTGGACTCGTTTCGAAATGGCGGCCCAAGGACCGAATGAAGACTGGATATGTAGCTCTAGTTTTGTGTTTAAACATCA
GTGTTGATCCTCCGGATGTAATAAAGATATCCCCTTGTGCCAGAATGGAGTGCTGGACAGATCCACTTTCAATGGCACCTCAAAAAGCTCTTGAAACAAT
TGGAAAAAACTTAAGCATTCAGTATGAAAGGTGGCAACCTAAGGCTCGCTACAAGGTTCAACTTGATCCTACTGTTGATGAAGTGAAGAAACTTTGTAAT
ACATGCCGAAAATATGCCAAGTCTGAGAGAGTTTTGTTTCATTACAATGGACATGGTGTTCCAAAGCCAACTGCTAACGGTGAAATCTGGCTATTCAATA
AGAGTTATACACAATACATTCCCTTGCCAGTCAGTGATCTTGATTCCTGGCTGAGAACACCGTCAATCTATGTTTTTGATTGCTCTGCTGCTGGGATGAT
AGTCAATGCATTTCTGGAGCTTCATGACTGGAATGCTTCTGGCTCTGCTGGATCCACGCGGGATTGCATTTTGCTTGCAGCCTGTGAAGCACACGAGACT
TTGCCACAAAGTGATGAATTTCCTGCTGATGTGTTCACTTCATGCCTCACCACTCCTATCAAGATGGCATTGAAATGGTTCTACAGACGTTCACTACTTT
GTGACTCCCTTGACTACTCATTAATTGATAAAATTCCTGGCCGGCAAAATGATCGTAAAACGCTTCTGGGAGAATTGAACTGGATCTTTACTGCAGTGAC
AGACACAATTGCTTGGAATGTTCTTCCTCGTGATCTTTTCCAGAAGTTGTTCAGGCAAGATTTGCTAGTTGCCAGTCTGTTCAGAAATTTTTTGCTTGCT
GAAAGAATTATGCGTTCGGCAAATTGTTCTCCAATTTCTCATCCAATGTTGCCACCAACCCATCAGCATCATATGTGGGATGCATGGGATATGGCTGCTG
AAATTTGCCTCTCTCAGCTTCCATCTCTGGTCGAGGATCCAAATGCAGAATTCCAGCCGAGTCCTTTTTTTACTGAACAACTGACGGCTTTTGAGGTGTG
GTTGGACCATGGATTTGAACATAAGAAGCCACCCGAGCAGTTGCCTATTGTTCTTCAGGTTCTGCTAAGTCAATGTCATCGGTTCCGAGCACTGGTGCTT
CTAGGGAGATTCCTTGATATGGGACGCTGGGCAGTGGATCTGGCTTTATCTGTTGGAATATTTCCATATGTTTTGAAGCTGTTGCAAACAACAACACCAG
AACTAAGGCAAATTCTTGTATTCATATGGACAAAAATTCTAGCGCTGGATAAGTCATGCCAGGTTGATCTTGTTAAGGATGGGGGGCATACATATTTTAT
TAGGTTTCTCGACAGCTTGGAAGCATATCCAGAACAGCGTGCAATGGCTGCATTTGTTTTGGCTGTTATTGTTGATGGACACAGACGGGGCCAGGAAGCC
TGTATTAAAGCTGGTTTGATTCATGTGTGCTTGAAGCACCTTCAAGGTTCAGTGCCAAATGACACACAAACTGAACCCTTGTTTCTTCAGTGGCTATGCC
TCTGCCTGGGAAAGCTGTGGGAGGATTTCACAGAGGCTCAGATACTAGGTTTGCAGGCTGACTCCCCAGCAATATATGCTCCTCTACTGTTGGAGCCCCA
ACCAGAGGTTAGGGCTTCTGCTGCATTTGCACTGGCCACCTTGCTTGATGTTGGGGGTGATGTCTGCAGGGATGGTGCTACTGGAGATGATGAGTTTGAT
GATGATGAAAAAATTAGAGCTGAAATTAGTATTGTTAGAAGTCTATTAAGTGCTGTTTCAGATGGAAGTCCTCTGGTCAGGGCGGAGGTTGCTGTAGCTC
TGGCACGCTTTGCATTTGGCCACAAACAGCACCTCAAGTCAATTGCTGCCTCCTATTGGAAACCTCAGTCTAATTCTCTACTCAGCTCATTACCTTCTTT
GGCCCATATAAAAGCTACAGGTAGTGGACATGCCAACCCTAATCAGTATGTGCCACATGCAAGTATTGTTTCATCTCAATTTGGCCCTCTGACAAGAGTT
GGTAGTGACAGCCCATCCGTGGTTCGAGATGGAAGGGCTTCCACTAGCAGTCCTACTACAGCTGGAATTATGCATGGATCACCATTATCTGATGATTCAT
CTCTGCATTCTGATTCTGGTATATTGAATGACATTGTCAGCAATGGGGAAGTACTCCATTCAAGGCCAAAGCCTCTGGATAATGCATTATATTCCCAGTG
TGTGCTGGCTATGTGCACTTTGGCCAAGGATCCTTCTCCACGCATAGCAAGCCTTGGTCGAAGGGTACTGTCGATTATTGGGATTGAACAAGTGGTGACA
AAATCTGTAAATTCCACTGGTAGCAGTGGTCCCAAAACTTCATCTCCAAGTCTTGCTGGACTGGCTCGTTCATCATCATGGTTAGACATGCATGCAGGTC
ACATTCCTTTGACATTCAGAACTCCTCCTGTCAGTCCTCCACGATCAAGCTACTTAACAGGAATGCGTAGGGTGTGTTCTTTGGAGTTCAGGCCGCACCT
TATGAATTCTCCAGATTCAGGATTAGCTGATCCTCTTTTAGCTTCTGTTGGATCTTCTGGAGGCACAGAGCGGAGTTTGCTTCCTCAGTCAACTATCTAC
AATTGGAGTTGTGGTCACTTCTCAAAGCCGCTTCTCACCACACCGGATGATACTGAAGAAATATTAGTTAGAAGGGAAGAAAGGGAAAAATTTGCACTGG
AACATATTGCAATGTGTCAGCACTCTTCTGTTAGCAACCTTAATAACAGAATTGCTAGCTGGGATACAAAGTTTGAAACTGGTACAAAGACAGCCTTGCT
GCAGCCTTTCTCTCCTATTGTGGTTGCTGCTGACGAGAATGAACGAATTAGGGTGTGGAATTATGAGGAAGCTACCCTTCTGAATGGGTTTGATAATCAT
GATTTTCCTGACAGAGGAGTTTCGAAGCTGTGTCTTGTAAACGAGCTAGATGACAGTTTGCTCCTTGTTGCTTCATGTGATGGAAATATACGGATTTGGA
AAGACTACACTGTGAAGGGTAAGCAAAAGCTTGTCACTGCATTTTCTTCAATCCAAGGTCATAAACCTGGTGTGCGGAGTTTGAATGCTGTTGTGGACTG
GCAACAACAATCTGGATATCTGTATGCTTCGGGTGAGATATCATCCATCATGCTTTGGGACCTGGATAAAGAGCAGCTTATTCATTCTATTCCTTCATCA
TCAGATTGTAGCGTCTCAGCAATGTCTGCTTCTCAAGTTCATGGAGGTCAATTTACTGCTGGTTTTGTGGATGGATCTGTCAAACTCTATGATGTCAGAA
CACCTGAAATGCTTGTTTGTGCTACTCGGCCGCACACTGAGAATGTAGAAAAAGTTGTGGGGATTGGCTTTCATCCTGGGCTTGATCCTGGAAAGATTGT
CAGTGCATCCCAGGCTGGTGACATGAAGTTCCTGGATATGAGAAATTATAGAGATCCCTATCTCACGATCAAGGCCCACAGGGGCTCACTCACAGCTTTA
GCTGTTCATAGGCATGCCCCTATTATTGCCAGTGGATCGGCAAAACAAATTATTAAATTATTCAGTCTAAATGGTGAACCATTAGGCTCCATTAGATACC
ACCTAACCATAATGGCCCAGAAGATCGGTCCTGTAAGCTGCCTCACCTTCCATCCTTACCAGGTACTCCTTGCTGCTGGTGCTACTGATGCTTTATTCTC
AATCTACGCCGATGACAACACTCAAGCAAGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G106600.1 pacid=42769615 polypeptide=Potri.006G106600.1.p locus=Potri.006G106600 ID=Potri.006G106600.1.v4.1 annot-version=v4.1
MALGDLMASRFSSQSPVAFVSNHYDHYPSSHEDDAIDVARRDDNNNSNNNRDRDSDTASTSNYGGGNATASTTATTTTSTAYLPQTVVLCELRHEAFEAS
VPTGPSDSGLVSKWRPKDRMKTGYVALVLCLNISVDPPDVIKISPCARMECWTDPLSMAPQKALETIGKNLSIQYERWQPKARYKVQLDPTVDEVKKLCN
TCRKYAKSERVLFHYNGHGVPKPTANGEIWLFNKSYTQYIPLPVSDLDSWLRTPSIYVFDCSAAGMIVNAFLELHDWNASGSAGSTRDCILLAACEAHET
LPQSDEFPADVFTSCLTTPIKMALKWFYRRSLLCDSLDYSLIDKIPGRQNDRKTLLGELNWIFTAVTDTIAWNVLPRDLFQKLFRQDLLVASLFRNFLLA
ERIMRSANCSPISHPMLPPTHQHHMWDAWDMAAEICLSQLPSLVEDPNAEFQPSPFFTEQLTAFEVWLDHGFEHKKPPEQLPIVLQVLLSQCHRFRALVL
LGRFLDMGRWAVDLALSVGIFPYVLKLLQTTTPELRQILVFIWTKILALDKSCQVDLVKDGGHTYFIRFLDSLEAYPEQRAMAAFVLAVIVDGHRRGQEA
CIKAGLIHVCLKHLQGSVPNDTQTEPLFLQWLCLCLGKLWEDFTEAQILGLQADSPAIYAPLLLEPQPEVRASAAFALATLLDVGGDVCRDGATGDDEFD
DDEKIRAEISIVRSLLSAVSDGSPLVRAEVAVALARFAFGHKQHLKSIAASYWKPQSNSLLSSLPSLAHIKATGSGHANPNQYVPHASIVSSQFGPLTRV
GSDSPSVVRDGRASTSSPTTAGIMHGSPLSDDSSLHSDSGILNDIVSNGEVLHSRPKPLDNALYSQCVLAMCTLAKDPSPRIASLGRRVLSIIGIEQVVT
KSVNSTGSSGPKTSSPSLAGLARSSSWLDMHAGHIPLTFRTPPVSPPRSSYLTGMRRVCSLEFRPHLMNSPDSGLADPLLASVGSSGGTERSLLPQSTIY
NWSCGHFSKPLLTTPDDTEEILVRREEREKFALEHIAMCQHSSVSNLNNRIASWDTKFETGTKTALLQPFSPIVVAADENERIRVWNYEEATLLNGFDNH
DFPDRGVSKLCLVNELDDSLLLVASCDGNIRIWKDYTVKGKQKLVTAFSSIQGHKPGVRSLNAVVDWQQQSGYLYASGEISSIMLWDLDKEQLIHSIPSS
SDCSVSAMSASQVHGGQFTAGFVDGSVKLYDVRTPEMLVCATRPHTENVEKVVGIGFHPGLDPGKIVSASQAGDMKFLDMRNYRDPYLTIKAHRGSLTAL
AVHRHAPIIASGSAKQIIKLFSLNGEPLGSIRYHLTIMAQKIGPVSCLTFHPYQVLLAAGATDALFSIYADDNTQAR
|
DESeq2's median of ratios [POPLAR]
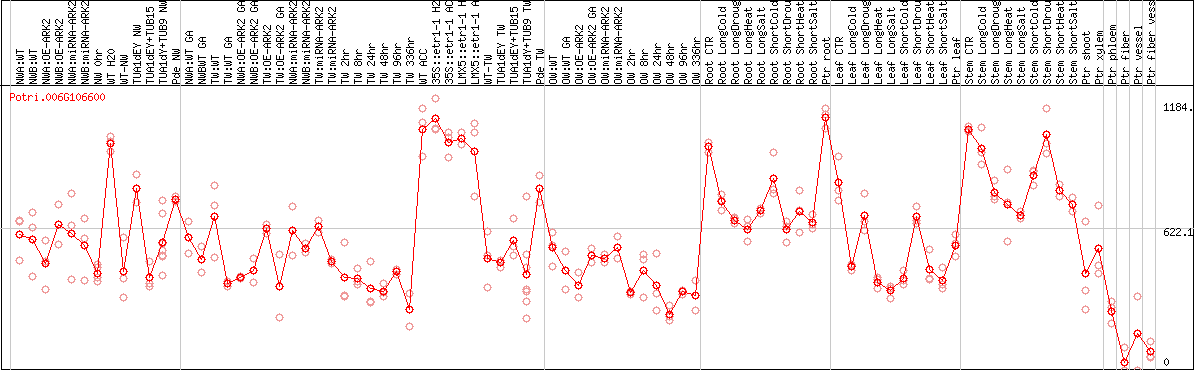
Coexpressed genes
Potri.006G106600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
