Potri.006G115000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G115000.1 pacid=42770175 polypeptide=Potri.006G115000.1.p locus=Potri.006G115000 ID=Potri.006G115000.1.v4.1 annot-version=v4.1
ATGGCGGCAGTTTTAGCCGGAGATGATCTGGCAAGATCAATGAGCAGCAGCAGCAGGAGGAGCTTTAGTTACAGGAGCTGGGCGTCTGCTAGCATAAGAG
AAGCATGGACTGCACCGGTGGACGTTTTCAGCCAGAATAGCGGGAGGAGGCAGCAGCAGATGGACGACGAGGAGGAGCTGCGATGGGCAGCTATCGAGAG
ATTGCCGACTTATGATCGGATGAGGAAAGGGGTTCTAAGGCAGGTACTTGATAATGGAAGAATGGTTCAAAGTGAGGTTGATGTAACAAGGCTTGGAATG
CAAGATAAGAAGCAGTTGATGGAGAACATACTTAGAGTTGTTGAAGAAGACAATGAAAAGTTTCTGAGGAGAGTTAGAGACAGGACTGATAGGGTGGGGA
TTGAGATTCCAAAGATTGAAGTTCGATTTCAGCACTTATCAGTGGAGGGTGAAGTTTTTGTTGGGAGCAGAGCACTTCCTACCTTACTTAATGCCACGCT
GAATGCAGTAGAGAGTATTCTTGGATTGGTTGGGCTTGCTCCATCAAAGAAGAGAACAGTTCAGATTCTTCAAGATATCAGTGGAATTGTCAAACCATCA
CGGATGGCCCTACTTCTGGGTCCTCCAAGCTCGGGGAAAACAACAATGCTAATGGCACTTGCAGGCAAACTCCACCGGGAGCTAAGATCATCCGGGAAGA
TCACCTATTGTGGGCATGAATTGAAAGAGTTTGTTCCTCAGAGGAGTTGTGCTTACATTAGTCAGCATGATCTTCATTATGGAGAAATGACAGTCAGAGA
GACATTGGATTTTTCTGGACGCTGCTTAGGTGTGGGCACGAGGTATGAACTGCTGGCGGAACTCTCTAGGCGTGAGAAAGAAGCAGGAATTAAGCCAGAT
CCAGAAATTGACGCGTTCATGAAAGCCACAGCAATGTCAGGACAGGAACATAGTTTGGTCACTGATTATACTCTCAAGATACTCGGATTGGATATTTGTG
CTGATATTTTGGTTGGGAATGACATGAAAAGGGGTATTTCTGGAGGGCAGAAGAAGCGTGTGACTACCGGAGAGATGTTGGTGGGACCAGCAAAGGTTCT
TTTAATGGATGAAATATCTACAGGGTTGGACAGTGCCACTACTTTCCAGATTTGCAAGTTCATGAGGCAAATGGTCCATACCATGGATGTAACTATGATA
GTTTCTCTCCTACAGCCAGCACCAGAGACATTTGAGCTTTTTGATGACATTATCCTGCTTTCAGAGGGTCAGGTTGTCTATCAAGGTCCACGCGAGCATG
TCCTTGAATTCTTTGAACACATGGGTTTCAGATGCCCTGATAGGAAAGGAGCTGCTGACTTTTTGCAAGAAGTAACTTCCAAGAAGGACCAGGAACAATA
TTGGTTCAGAAAGAATATACCTTACAGATTTATTTCAGTTCTTGAGTTTGTGCGAGGTTTCAACTCTTTTCATGTCGGCCAACAACTTGCTTCAGATCTT
AGGACTCCTTATGATAAATCAAGGGCCCATCCGGCTGCATTGGTCACAGAGAAGTATGGCATCTCAAATTGGGAGCTGTTCAGGGCATGTTTCTCAAGAG
AGTGGCTGCTGATGAAGCGGAACTCTTTTTTGTATATATTCAAAACAACCCAGATAACAATCATGTCAATAATTGCCTTCACAGTGTTTTTTCGGACAGA
AATGAAAGTGGGGACTGTACTAGGTGGACAAAAATTTTTTGGAGCATTGTTTTTCAGTCTAGTCAATGTGATGTTTAATGGGATGGCAGAACTTTCAATG
ACAGTTTTCAGGCTCCCTGTCTTCTATAAACAGAGGGATTTCTTGTTCTTTCCTGCGTGGGCTTTCGGCTTACCAATATGGGTACTCAGGATTCCATTAT
CGTTAATGGAATCAGCGATATGGATTATCCTAACATATTACACAATTGGTTTCGCTCCTTCTGCTAGCAGGTTCTTCAGACAGTTCCTTGCATTTTTTTG
CATACATCAGATGGCTCTGGCCCTCTTTCGTTTCATTGCTGCTGTTGGAAGAACACAAGTCGTTGCAAACACACTAGGGACCTTCACCTTGCTTTTGGTT
TTCGTGCTCGGAGGATTCATTGTTGCCAAAGACGACATTGAGCCATGGATGATCTGGGGCTACTATTCCTCTCCTATGATGTATGGACAGAATGCCATTG
TCATGAATGAATTCCTTGACGAAAGATGGAGTGTGAATAATACGGACTCTAATTTCGCTGGAGAAACTGTTGGGAAAGTGCTTCTGAAGGCTAGAGGCTT
CTTTACTGATGATTATTGGTTTTGGATTTGTATCGGAGCACTCTTTGGGTTTTCGCTTTTATTCAACGTTTTATTTATTGTAGCATTGACTTTCTTGAAC
CCTTTGGGTGACTCAAAAGCTGTCGTTGTGGATGACGATGCGAAGAAGAATAAGAAGACATCATCTGGACAACAGAGAGCAGAAGGTATTCCTATGGCAA
CGAGAAATTCTACAGAAATTGGTGGCGCAGTGGATAATTCAACAAAAAGAGGAATGGTTCTGCCATTCCAGCCCCTTTCACTTGCATTCAACCATGTGAG
CTACTATGTGGATATGCCTGATGAAATGAAGAGTCAAGGAATTGATGAAGAGCGTCTTCAACTGCTCAGAGATGTTAGTGGTGCTTTTAGACCAGGGATA
TTGACTGCACTTGTGGGCGTAAGTGGTGCTGGGAAGACCACCTTGATGGATGTGTTAGCTGGAAGGAAGACCGGTGGTTACATTGAAGGAAGTATTAACA
TCTCAGGCTATCCAAAGAATCAAGAGACATTTGCTCGTGTGAGCGGTTATTGTGAACAGAATGACATTCATTCACCTCGTGTTACTGTTTATGAGTCTCT
TCTGTACTCAGCCTGGCTCCGTCTTTCTAAAGATATAGATACAAAAACCCGAAAGATGTTTGTTGAAGAAGTTATGGAATTGGTTGAGCTAAATCCCCTC
AGAGATGCCTTAGTTGGACTTCCAGGTTTAGACGGCCTATCAACAGAACAAAGGAAAAGATTGACCATAGCAGTAGAATTGGTAGCTAATCCCTCTATCA
TATTCATGGATGAGCCAACTTCTGGTCTTGATGCTAGAGCTGCTGCTATCGTGATGCGTACTGTCAGAAACACTGTGGACACAGGTAGAACTGTTGTGTG
CACCATTCATCAGCCTAGCATAGACATTTTTGAAGCTTTTGATGAGCTGTTGTTGATGAAAAGAGGAGGTCAAGTCATTTATGCCGGATCTCTTGGTCAC
CGTTCTCACAAGCTCATAGAATATTTTGAGGCTGTCCCAGGTGTTCCCAAGATAAGAGATGCGTATAACCCTGCAACATGGATGCTCGAAATTAGTGCTC
CTTCAATGGAGGCACAACTGGATGTAGACTTCGCAGAACAATATGCCAACTCCTCCCTCTATCAGAGGAATCAGGAAATTATCAAAGAGCTAAGCACACC
AGCACCAGGATCCAAGGATCTCTACTTCCGAACCCAATACTCCCAAACTTTTCTTACTCAATGCAAAGCTTGTTTCTGGAAACAGCATTGGTCTTACTGG
AGAAACCCTCGTTATAATGCCATTCGTCTGTTCATGACATTAGCCATTGGAATAATATTCGGTCTCATCTTTTGGGACAAAGGACAGAAGACATTTAGCC
AGCAAGATCTATTGAATGTTTTCGGAGCCATGTACGCAGCGGTGCTTTTCCTTGGTGCAACCAATGCTGCGGGAGTGCAGTCTATAATTGCTATTGAGAG
AACAGTTTTCTATCGCGAAAGAGCAGCTGGAATGTACTCCCCATTGCCTTACGCATTTGCGCAGGTGGCCATAGAAGCAATTTATGTAGCAGTACAAACT
ATTGTTTATAGCATTCTCCTTTTCTCCATGATGGGCTTCGAGTGGACGGCTGCAAAGTTCTTGTGGTTCTACTACTTCATATTCATGTGCTTTGTTTACT
TCACATTGTTTGGGATGATGGTTGTTGCATTGACCCCTGCCCCACAAATTGCTGCCATTTGCATGTCCTTCTTCACGTCCTTCTGGAACTTGTTCTCTGG
TTTCCTTCTTCCTAGGCCTCAAATACCTATTTGGTGGAGGTGGTACTACTGGTGTTCTCCAGTGGCTTGGACATTATACGGCCTCGTGACATCTCAAGTG
GGTGATAAAACCAATACAATCAGCGTACCTGGCGAGTCTGAAGATGTGCCAATAAAGGAATTCCTCAAAGGATACTTGGGATTTGAATATGACTTTCTTC
CTGCCGTGGCCGCCGCCCATCTTGGATGGGTTGTCCTCTTCTTCTTTCTGTTTTCCTATGGTATCAAGTTCCTCAACTTCCAGAAGAGATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G115000.1 pacid=42770175 polypeptide=Potri.006G115000.1.p locus=Potri.006G115000 ID=Potri.006G115000.1.v4.1 annot-version=v4.1
MAAVLAGDDLARSMSSSSRRSFSYRSWASASIREAWTAPVDVFSQNSGRRQQQMDDEEELRWAAIERLPTYDRMRKGVLRQVLDNGRMVQSEVDVTRLGM
QDKKQLMENILRVVEEDNEKFLRRVRDRTDRVGIEIPKIEVRFQHLSVEGEVFVGSRALPTLLNATLNAVESILGLVGLAPSKKRTVQILQDISGIVKPS
RMALLLGPPSSGKTTMLMALAGKLHRELRSSGKITYCGHELKEFVPQRSCAYISQHDLHYGEMTVRETLDFSGRCLGVGTRYELLAELSRREKEAGIKPD
PEIDAFMKATAMSGQEHSLVTDYTLKILGLDICADILVGNDMKRGISGGQKKRVTTGEMLVGPAKVLLMDEISTGLDSATTFQICKFMRQMVHTMDVTMI
VSLLQPAPETFELFDDIILLSEGQVVYQGPREHVLEFFEHMGFRCPDRKGAADFLQEVTSKKDQEQYWFRKNIPYRFISVLEFVRGFNSFHVGQQLASDL
RTPYDKSRAHPAALVTEKYGISNWELFRACFSREWLLMKRNSFLYIFKTTQITIMSIIAFTVFFRTEMKVGTVLGGQKFFGALFFSLVNVMFNGMAELSM
TVFRLPVFYKQRDFLFFPAWAFGLPIWVLRIPLSLMESAIWIILTYYTIGFAPSASRFFRQFLAFFCIHQMALALFRFIAAVGRTQVVANTLGTFTLLLV
FVLGGFIVAKDDIEPWMIWGYYSSPMMYGQNAIVMNEFLDERWSVNNTDSNFAGETVGKVLLKARGFFTDDYWFWICIGALFGFSLLFNVLFIVALTFLN
PLGDSKAVVVDDDAKKNKKTSSGQQRAEGIPMATRNSTEIGGAVDNSTKRGMVLPFQPLSLAFNHVSYYVDMPDEMKSQGIDEERLQLLRDVSGAFRPGI
LTALVGVSGAGKTTLMDVLAGRKTGGYIEGSINISGYPKNQETFARVSGYCEQNDIHSPRVTVYESLLYSAWLRLSKDIDTKTRKMFVEEVMELVELNPL
RDALVGLPGLDGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELLLMKRGGQVIYAGSLGH
RSHKLIEYFEAVPGVPKIRDAYNPATWMLEISAPSMEAQLDVDFAEQYANSSLYQRNQEIIKELSTPAPGSKDLYFRTQYSQTFLTQCKACFWKQHWSYW
RNPRYNAIRLFMTLAIGIIFGLIFWDKGQKTFSQQDLLNVFGAMYAAVLFLGATNAAGVQSIIAIERTVFYRERAAGMYSPLPYAFAQVAIEAIYVAVQT
IVYSILLFSMMGFEWTAAKFLWFYYFIFMCFVYFTLFGMMVVALTPAPQIAAICMSFFTSFWNLFSGFLLPRPQIPIWWRWYYWCSPVAWTLYGLVTSQV
GDKTNTISVPGESEDVPIKEFLKGYLGFEYDFLPAVAAAHLGWVVLFFFLFSYGIKFLNFQKR
|
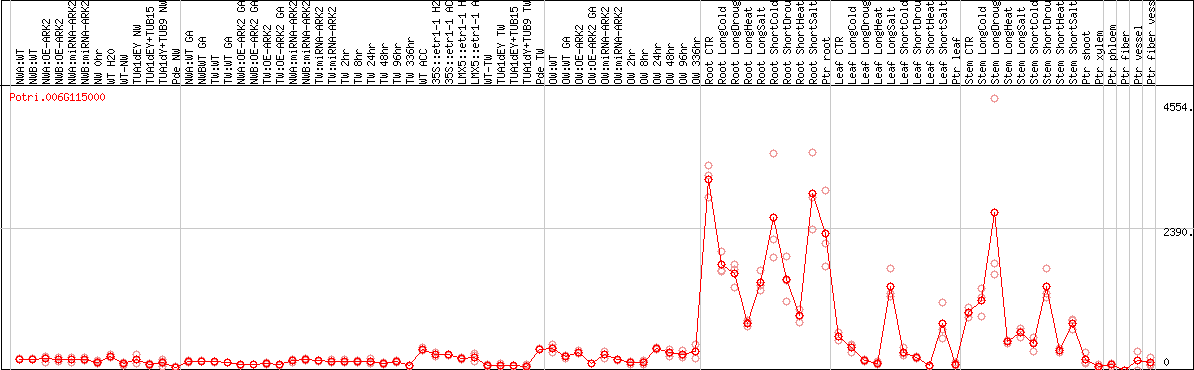
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G115000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
