AGO909 (Potri.006G118600) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | AGO909 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G118600.1 pacid=42767780 polypeptide=Potri.006G118600.1.p locus=Potri.006G118600 ID=Potri.006G118600.1.v4.1 annot-version=v4.1
ATGTACGGGAGAGGTCGACGCGGTGGCTCGCCTGCTCCGACAAAAGGAGGAGGGCGTGGCAGAGGACGCGGAGCTCCACTTCCTTCTCCTATGGCTTCGT
CCGAGGCCGACTCTATCTCATCCGTGAGCCAGCTCGGTGGTGAGATGGAGCGGCTCAGTGTTCAAACTGAACCACCTGCTCCTACTCAGGCACCAGCGGC
TATTCCTGCACCGCAACAACAGAAGCAGCAGCAGCAGCAACTCGTACCGGCTTCCTCTGTGAAGTTCGCTCAAAGACCTGATCACGGTACGGTGGGATCC
CGGTGTCTCATTAGAGCTAACCACTTTCTTGTTGAGCTCGCTGACCGAGACTTGCATCACTACGATGTTTCTATAACTCCTGAGGTTGCATCTCGAGGGG
TCAACAGAGCAATAATGAGAGAGCTGCTTGCTTCGAATAGCACACACTTCCAGAGTAGAAAACCTGCTTATGATGGAAGAAAGGGATTCTACACTGCTGG
ACCTTTAACATTCACCTCAAAGGATTTTGTGGTTACCCTTGTAGACAAAGATGATCAAGGATCTGTGAGAAAAGAGAGGAAATTCAAAGTCACTATCCGG
TTGGCATCTAAAACAGACCTTTATCATCTCAAGGAGTTCCTGCAGGGAAGACAAAGGGGTGCGCCGCATGATACCATACAAGTACTCGATGTAGTCCTAA
GGGAACCACCATCCAACAATTGCACCATCGTTGGGAGGTCTTTTTTCACAGCAGGTCTGGGTGGCCAAAATGAGATTGGTAATGGTATAGAATGCTGGAA
GGGATTCTACCAGTCTCTACGCCCAACGCAGATGGGAATGTCTCTTAACATAGATGTATCAGTCGCTGCCTTCTACGAGCCGATTCTTGCTGTTGACTTT
GTTGCAAAACTATTAAATTTAGGAGATCCAATTAGAGCAGCAACCAGGCCTTTGTCAGATAGTGATCGAGCAAAGTTGAAAAAAGCTTTGAGAGGAGTCA
GAGTAAAAGTTACCCATGGAGAGGAGAAGCGTTACAAAATCACTGGAATATCTCCTTCGGCAACAAACCAACTAAGGTTTGCTGCTGAAGATGGAAAACA
AAAATCAGTTGTCCAATACTTCCTGGAAAAATACAATATAAGGCTTCGTTTTGCATCCTGGCCTGCTCTTCAATCAGGAAATGATTCAAGACCAATATTC
CTGCCTATGGAGTGTTGCAAGATTATCGAAGGACAAAGGTACTCAAAGAAGTTGAATGAGAAGCAGGTGACAGCCTTATTGAGAGAGGCCTGCAGGCGCC
CTGTTGAGAGAGAACATAGTATTGAGCAGATTGTTCATTTTAATGACGTTGCACAAGATGATCTAGCCAAAGAATTTGGAGTCAGTGTCAAAAAGGAGCT
GACTTGCATCGATGCCAGAGTCTTGCCACCTCCGGTGCTCAAGTACCATGATTTGGGAAAAGCTAGAACTGTAAGACCCCGAGTGGGGCAGTGGAACATG
ATTAATGCTAAATTGTTCAATGGGGCCACAGTGAACTTTTGGATGTGTGTCAACTTCTCAAGTCTTGGTGAGCAAATGGCTGCTAGCTTTTGTCGAGCCC
TGGTAGGCATGTGCAACAACAAGGGGATGGTCATAAATCCTGCACCTGTATTCCCAATACGGTCTGGCCATCCTAACCAGCTGGAGAAAACATTGGCCGA
AGTTCACAGCATGTGTAATAATGAAAGAAAACAGCTTCAAATCTTAATTATCATTCTCCCTGATGTCAGTGGAAGTTATGGCACAATAAAAAGAGTATGC
GAAACTGAACTTGGGATAGTTTCTCAATGCTGTCAACCCAAGCAAGCAAGAAAGTGTAGTCCTCAATACTTGGAAAATGTTGCGCTGAAAATTAATGTGA
AGGCTGGAGGGCGAAACACAGTATTAGAGGATGCCCTGAATAGGAGAATACCTCTTCTAAGTGACACCCCAACTATAATCTTTGGTGCTGATGTAACCCA
TCCACAACCAGGGGAGGATTCTAGCCCTTCAATAGCTGCGATTGTGGCATCAATGGACTGGCCTGAAGTAACCACCTACAGGGGCCTGGTATCTGCTCAG
AAACATCGTCAAGAGATTATTCAAGATTGTGCTGGAATGATCAGGGAACTTATGATTGCTTTCAGAAGAACAACTAATCAGAAACCTAGCAGAATAATTT
TCTATAGGGATGGTGTTAGTGAGGGCCAGTTCAGCCAAGTTCTCCTGTATGAGATGGATGCTATCCGAAAGGCTTGTGCATCTCTGGAACCAAACTATTT
GCCACCAGTTACTTTTATTGTAGTGCAGAAGAGACACCATACTCGGCTCTTTGCTACAAATCCTAACCAAACAGACAAGAGTGGAAACATCCTTCCTGGA
ACGGTTGTTGATACAAAGATATGCCATCCTTCAGAGCACGACTTTTATCTCTGCAGTCATGCAGGAATTCAGGGGACTAGCAGGCCGGTGCACTATCATG
TGTTGTGTGACATGAACAAATTCACTGCTGATTGCCTGCAGATGTTGACTAACAACCTGTGCTACACGTATGCAAGGTGCACCCGCTCTGTTTCTGTGGT
TCCTCCTGCATACTATGCACACTTGGCAGCATTCAGGGCAAGATACTACATAGAGGGGGACATTGCATCTGATAGTGGCGGCGGCGGCACAGGACCTCCT
GTGAGGAGGGAAGCTGCCCCTGTCCGCCCACTTCCAGCCATCAGCCCCAACGTGAAGAATGTTATGTTTTACTGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G118600.1 pacid=42767780 polypeptide=Potri.006G118600.1.p locus=Potri.006G118600 ID=Potri.006G118600.1.v4.1 annot-version=v4.1
MYGRGRRGGSPAPTKGGGRGRGRGAPLPSPMASSEADSISSVSQLGGEMERLSVQTEPPAPTQAPAAIPAPQQQKQQQQQLVPASSVKFAQRPDHGTVGS
RCLIRANHFLVELADRDLHHYDVSITPEVASRGVNRAIMRELLASNSTHFQSRKPAYDGRKGFYTAGPLTFTSKDFVVTLVDKDDQGSVRKERKFKVTIR
LASKTDLYHLKEFLQGRQRGAPHDTIQVLDVVLREPPSNNCTIVGRSFFTAGLGGQNEIGNGIECWKGFYQSLRPTQMGMSLNIDVSVAAFYEPILAVDF
VAKLLNLGDPIRAATRPLSDSDRAKLKKALRGVRVKVTHGEEKRYKITGISPSATNQLRFAAEDGKQKSVVQYFLEKYNIRLRFASWPALQSGNDSRPIF
LPMECCKIIEGQRYSKKLNEKQVTALLREACRRPVEREHSIEQIVHFNDVAQDDLAKEFGVSVKKELTCIDARVLPPPVLKYHDLGKARTVRPRVGQWNM
INAKLFNGATVNFWMCVNFSSLGEQMAASFCRALVGMCNNKGMVINPAPVFPIRSGHPNQLEKTLAEVHSMCNNERKQLQILIIILPDVSGSYGTIKRVC
ETELGIVSQCCQPKQARKCSPQYLENVALKINVKAGGRNTVLEDALNRRIPLLSDTPTIIFGADVTHPQPGEDSSPSIAAIVASMDWPEVTTYRGLVSAQ
KHRQEIIQDCAGMIRELMIAFRRTTNQKPSRIIFYRDGVSEGQFSQVLLYEMDAIRKACASLEPNYLPPVTFIVVQKRHHTRLFATNPNQTDKSGNILPG
TVVDTKICHPSEHDFYLCSHAGIQGTSRPVHYHVLCDMNKFTADCLQMLTNNLCYTYARCTRSVSVVPPAYYAHLAAFRARYYIEGDIASDSGGGGTGPP
VRREAAPVRPLPAISPNVKNVMFYC
|
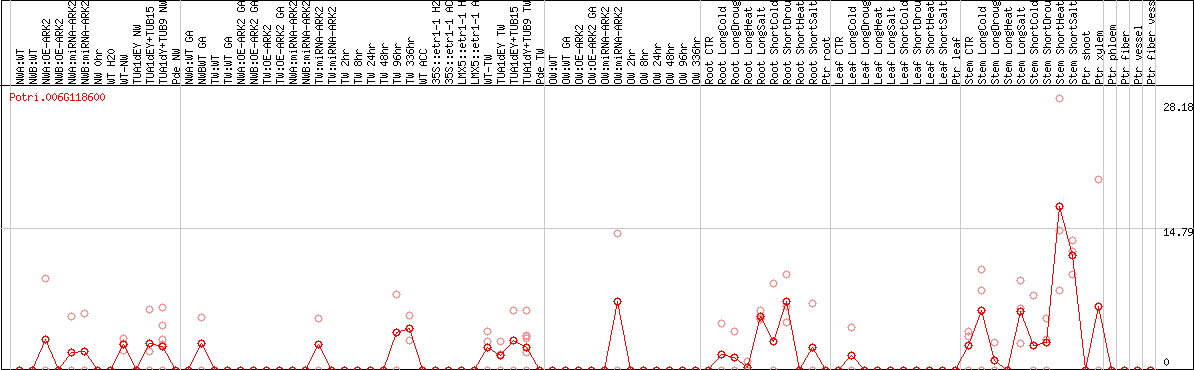
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G118600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
