Potri.006G119600 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.006G119600.3 pacid=42769967 polypeptide=Potri.006G119600.3.p locus=Potri.006G119600 ID=Potri.006G119600.3.v4.1 annot-version=v4.1
ATGGCATCCCCAAAACAACTCCTCTCCACCATCGAATCCACACTTCTAAATCCTTCGCCACCGTCAGCGGCGGAGAGAGTCGAACTAATGCACGCTATTC
GCAGCTCTCTCCCGTCTCTGCAAGCGCTCCTCTTCTACCCGCCTCCCAAACCATCGGACCGCGCACAAGTCGTCCAGTCGAAAGAAGTTAGGCTGCCGGA
TTCACCAGCAATCTCACTAGATGATCAGGATGTGCAAATTGCATTGAAATTGAGTGATGACCTCCATCTTAACGAAATAGAGTGTGTTCGATTGCTGGTT
TCTGCCAATCAAGAGTGGGGTTTAATGGCACGGGAGCCATTAGAAATTCTGCGGCTTGCAGCAGGACTTTGGTACACCGAGAGACGAGATCTAATTACAG
CCCTGCATATGCTATTAAGGGCTGTAGTGCTTGATCGAGGACTTGAAGATGATATCGTGTCAGACATCCAGAAGTATCTTGAAGATCTAATCAATGGTGG
GCTTCGACAGCGATTGATTTCTCTAATAAAGGAACTCAATCTGGAAGAACCAGCTGGTTTTGGTGGGCCTCTTTGTGAGCATTATGTCCTTGATTCTAGA
GGTGCTCTTGTTGAGCGTCAAGCTGTGGTTTGTAGAGAGAGATTGATACTTGGACATTGCCTTGTCCTCTCTGTTCTGGTTGTGCGGACAAGTCCAAAAG
ATGTAAAGGATATTTTTAATTGTTTAAAAGATAGTGCAGCTGAACCTATGGAGGGCACCAACACATTAAAACATCAGATTACATTCAGTCTGCTTTTCTC
TCTTGTAATTGCCTTCATATCAGATGCTCTGAGTGCCGTTCCTGATAAAGGATCTATCCTGTCTCATGATGCTTCTTTTAGAAAAGAATTTCATGAAATT
GTGATGGCTGTTGGAAACAATCCAAACGTGGAGGGCTTTGTTGGTGTTATAAGACTTGCTTGGTGTGTGCATTTGATGTTAATAAATGATGGCGTTTCAG
CTTCTTCATCAAATAATTCAGGCTATGTAAATTCGTGCCTTGAATTCATTTTCTCACACAATGTTTTTCAGTTTTTACTGGATAACATTCTTCGAACAGC
AGCCTACCAGAATGATGACGAGGATATGATCTACATGTACAATGCTTATATGCACAAGCTGACTACTTGCTTACTCTCTCACCAACTTGTGAGGGATAAG
GTAAAAGAATCTAAGGACAAGGCAATGAGCACGCTCAACTCATACCGTTTGGCTGTTTCACAAGATCTTATGCATGATAGCAACCCGGATTCACAACAGG
CCACCGAAACTGGTCCTCTGCTGTTTGTCTCTCTGCTGGAGTTTGTGAGCGAAATTTATCAGAAAGAACCAGAGCTATTGTCTGGCAATGATGTTTTGTG
GACATTTGTCAATTTTGCTGGAGAGGATCATACTAATTTTCAAACACTTGTAGCCTTTTTAAAAATGCTGAGTGCTCTGGCTTCAAGTCAAGAAGGTGCT
GCAAAGGTTTATGAGTTACTTCAGGGAAACGCCTTCCGTTCTGTTGGGTGGAGCACTTTGTTTGATTGCTTAACTATATATGACGAGAAGTTCAAACAGT
CAGTTCAGACAGCTGGAACCATGCTACCAGAGTTTCAGGAAGGGGATGCAAAAGCACTGGTTGCATATCTGGATGTTCTTCAAAAGGTTATTGAAAATGG
GAATCCTGTGGATAGAAAAAACTGGTTTCCTGACATTGAACCATTGTTCAAGCTCCTTAGTTATGAAAATGTGCCTCCATATTTGAAGGGTGCTCTGCGG
AATGCTATAGCTACTTTTGTTCATGTCTCTCCAGTCCTAAAAGATGCTATTTGGAGTTATCTTGAGCAGTATGATTTACCTGTGGTTGTTGGGGCTCAGG
TTGGAAACATTGCACAGCCGACAGGAACACAGGTTTATGATATGCAATATGAGTTGAATGAAATAGAAGCAAGGAGAGAACGATATCCTTCGACAATATC
TTTCCTTAACTTGTTAAATGCTCTCATTGGTGAGGAAAAAGATGTGAGCGATAGAGGGCGTAGATTTATAGGCATCTTTAGGTTCATATGTGATCATGTC
TTTGGGCCATTCCCTCAGAGAGCCTATGCAGATCCTTGCGAGAAATGGCAGTTAGTTGTTTCTTGTCTTCAACATTTTCACATGATGTTGAGCATGTATG
AAATTGAAGATGAGGACATTGACAGTGTTGTTGACCTGTCACAGCTTTCAACAGGCACGCAACCATCTTCGCTTCAGATGCAGCTTCCGGTCTTAGAGTT
GTTAAAAGATTTCATGAGTGGCCGGATTGTTTTTCGAAATATCATGGGCATCCTTCTGCCAGGGGTAAATTCTATTATCACTGAACGGACTTCTCAAATT
TATGGGCAACTGCTTGAAAAGGCTGTACAACTTTCTTTGGAGATTATAATTCTTGTCCTTGAAAAGGATTTACTAGTTTCTGATTATTGGCGTCCTCTAT
ACCAGCCTTTGGATGTTATTCTTTCACAAGATCATAATCTAATTGTGGCATTGCTGGAGTATGTCAGATATGATTTTCTACCCAAAATTCAGCAATGCTC
AATCAAAATCATGAGTATTCTAAGTTCCCGTGTGGTGGGGCTCGTACAGTTACTGTTGAAATCCAATGCTGCAAACTCTTTGGTTGAGGATTATGCAGCC
TGCCTTGAGGTACGATCAGAAGAATGTCAGATTATAGAGAACAAGGTTGATGATCCAGGGGTTCTGATAATGCAGCTCCTAATAGATAACATTAGTCGGC
CAGCTCCAAATGTTACACATTTACTGCTTAAATTTGATATTGACCACGCAGTTGAGCGGACAGTTCTGCAACCAAAGTTTCACTACAGTTGCTTGAAGGT
TATTCTTGAAATATTGGACAGACTTTTAAAACCTGAAAAGAATGCAATGCTTCATGAGTTTGGTTTTCAGCTCCTTTATGAGTTGTCCGTGGACCCGCTT
ACATGTGGCCCTACCATGGATTTGCTGAGCAAGAAAAAGTATCAATTCTTTGTAAAGCACCTTGATACTATTGGTATTGCTCCGCTTCCTAAACGGAATA
GCAATCAGCCACTTCGCATCAGCTCCCTTCACCAGAGAGCATGGCTCTTGAAGCTTTTAGCTGTAGAATTGCACTCTGGTTATGTGGGTGGCCCCTCTCA
TCGAGAAGCATGTCAGACCATACTTGCTCATCTTTTTGGACGGGATGTAATTGAAAGTGGGCCTGATCATGTAGTTTATGATTCCGTTATACTTCGAAAC
GGCACTGAGCATGCTGGAACCCAGACAATAAGCAAGAATAAGGTTTTGGAGTTGCTCGAGGTTATCCAATTTCGATCTCCTGATACTACCATGAAGCTTT
CCCAGATTGTTTCCAATATGAAGTATGATTTGATGGCAGAGGAAATACTTGGAGATCCAAGAACTTCAGGAAGGGGCGGTATCTACTACTATTCAGAGCG
CGCTGATCGTCTGATTGACCTTGCTTCTTTCCGTGATAAGCTTTGGCAGAAATTCAATTCAGTCTATCCCCAGTTAAGTAACTTTGAAAATGAAGCTGAA
CTCAATGATGTAAGAGAGACAATTCAACAGTTATTGAGATGGGGGTGGAAGTATAATAAGAACCTTGAGGAACAAGCTGCCCAACTTCATATGTTAACTG
GCTGGTCACATATTGTTGAGGTGTCTGCATCCAGAAGAATACATTCACTTGAAAATCGATCTGACATTTTATATCGGGTTCTTGATGCCTCTCTAAGTGC
TTCTGCTTCTCGAGACTGTTCATTGAGGATGGCATTTGTATTGAGTCAGGTTGCACTAACATGCATGGCTAAACTACGTGATGAAAGGTTCTTGTGCTCT
GGTGGTTTAAATTCAGATAATATGACATGTCTTGATGTTATAACGGCAAAGAAACTATCAAATGGTGCCTGTCACTCAATATTATTTAAGCTTATAATGG
CAATTCTGAGAAATGAATCATCAGAATCTCTGAGAAGACGTCAATATGCATTGCTGCTCGGCTATTTTCAGTATTGTCAGCATATGCTTGATCCAAATAT
CCCAACATCAGTTATGCAATTTTTGATGCTCGAGGAGCAAGATAGTGAAGATCTAGATTTCCAGAAAATTGACAAAGATCAGGCAGAACTAGCTCGTACC
AATTTTTCAATTATGAGGAAAGAAGCTCAGGCCATCTTGGATTTGGTAATAAATGATGCAACTAAAGGAAGCGAACCTGGGAAGACAATAGCGCTGTATG
TTCTGGTTGCATTAATATGTATTGACCATGAGAGATACTTCTTAAGCCAACTTCAAAGTAGAGGATTTTTAAGGACTTGCCTAACGAGCATCAGCAATTT
TTCAAACCAGGATGGCGGGCATTCATTGGATTCATTGCAGCGGGCAAGTACCCTTGAGGCTGAATTGGCATTACTGTTGAGGATTAGCTACAAATATGGG
AAATCTGGGGCCCAGGTTTTGTTTTCCATGGGTGCCTTGGAGCATCTTGCATCATGCAGGGCAGTCAGTTTACAGGGATGTTTGAGGCGGTTTGACCGAA
AGCTTCGTCGAGATGTTTCTGTAGATTTTGACAAACAATGCATGATTGTAACACCTATGTTGAGATTGTTGTTTTCTTTGACGTCATTGGTTGATACATC
TGATATATTCGAGGTGAAGAATAAAATTGTTCGTGAAGTTATAGATTTTGTAAAAGGACATCAGATGCTGTTTGATCAAATTCTTCGTGAAGATATTTCC
ACAGCTGATGAATTGACTGTGGAGCAGATCAATCTTGTTGTTGGCATACTTTGTAAGGTATGGCCCTATGAAGAGAGTGACGAGTTTGGTTTTGTTCAAG
GTCTTTTCAGTATGATGCGTGCTCTTTTTTCTTGTGACTCTGGAGCTCCTACTGTTGGTAAATTGGCCCAATCTTCTGAGGTATCTCAGAATAAAAGCAA
GGTAGAACTGAACTCGTTTCGGCTATGCTTCAGTCTGGGCTCTTACCTATATTTCTTAGTCACAAAAATGTCTCTGAGACTCCAGGTCTCAGACAGCTCC
ATAGATTACCATTCCCCTGCTATGCTGCAGCAGCCCACATTGATGTTACTTGACTCTCTCCTCCGTTCAGTTGCAACTTCCCTTGAAATGGCTGCTGAGG
AGAAATCCTTACTTCTTAATAAGATTCAAGATATAAATGAACTGTCAAGGCAGGAGGTGGATGAAATTATCAACATGTGTGTTATGCAAGAGTCTGTTTC
TTCATCCGATGACATACAGAAGAGGAGGTATATCGCAATGATGGAAATGTCCCGTGTAGCTGGCGAAAGAGATCAGCTAATTACATTGTTACTTCCACTT
GCAGAACATGTGCTGGATATCATCCTTGTCCATTTTCGTGAAGGGTCCATGGCATCTGATAATAGTGGAGCTACAAAAGCAGTTACATTTGGCACACACT
CTGACCCCAGACAGGATCTATCTTGGATGTGCGGGATGTTGGTTCCCACGTTAGAACGACTTGAACTGTTGAGTGAGGATAAGGTCGGCCACAATCTGAA
GGTGTTTCGAAGATTGGTGACATCCCTTAAAGAAATGGCAATCCAAAACCTGGCTTTATGA
|
|||||||||||||||
|
AA sequence
|
>Potri.006G119600.3 pacid=42769967 polypeptide=Potri.006G119600.3.p locus=Potri.006G119600 ID=Potri.006G119600.3.v4.1 annot-version=v4.1
MASPKQLLSTIESTLLNPSPPSAAERVELMHAIRSSLPSLQALLFYPPPKPSDRAQVVQSKEVRLPDSPAISLDDQDVQIALKLSDDLHLNEIECVRLLV
SANQEWGLMAREPLEILRLAAGLWYTERRDLITALHMLLRAVVLDRGLEDDIVSDIQKYLEDLINGGLRQRLISLIKELNLEEPAGFGGPLCEHYVLDSR
GALVERQAVVCRERLILGHCLVLSVLVVRTSPKDVKDIFNCLKDSAAEPMEGTNTLKHQITFSLLFSLVIAFISDALSAVPDKGSILSHDASFRKEFHEI
VMAVGNNPNVEGFVGVIRLAWCVHLMLINDGVSASSSNNSGYVNSCLEFIFSHNVFQFLLDNILRTAAYQNDDEDMIYMYNAYMHKLTTCLLSHQLVRDK
VKESKDKAMSTLNSYRLAVSQDLMHDSNPDSQQATETGPLLFVSLLEFVSEIYQKEPELLSGNDVLWTFVNFAGEDHTNFQTLVAFLKMLSALASSQEGA
AKVYELLQGNAFRSVGWSTLFDCLTIYDEKFKQSVQTAGTMLPEFQEGDAKALVAYLDVLQKVIENGNPVDRKNWFPDIEPLFKLLSYENVPPYLKGALR
NAIATFVHVSPVLKDAIWSYLEQYDLPVVVGAQVGNIAQPTGTQVYDMQYELNEIEARRERYPSTISFLNLLNALIGEEKDVSDRGRRFIGIFRFICDHV
FGPFPQRAYADPCEKWQLVVSCLQHFHMMLSMYEIEDEDIDSVVDLSQLSTGTQPSSLQMQLPVLELLKDFMSGRIVFRNIMGILLPGVNSIITERTSQI
YGQLLEKAVQLSLEIIILVLEKDLLVSDYWRPLYQPLDVILSQDHNLIVALLEYVRYDFLPKIQQCSIKIMSILSSRVVGLVQLLLKSNAANSLVEDYAA
CLEVRSEECQIIENKVDDPGVLIMQLLIDNISRPAPNVTHLLLKFDIDHAVERTVLQPKFHYSCLKVILEILDRLLKPEKNAMLHEFGFQLLYELSVDPL
TCGPTMDLLSKKKYQFFVKHLDTIGIAPLPKRNSNQPLRISSLHQRAWLLKLLAVELHSGYVGGPSHREACQTILAHLFGRDVIESGPDHVVYDSVILRN
GTEHAGTQTISKNKVLELLEVIQFRSPDTTMKLSQIVSNMKYDLMAEEILGDPRTSGRGGIYYYSERADRLIDLASFRDKLWQKFNSVYPQLSNFENEAE
LNDVRETIQQLLRWGWKYNKNLEEQAAQLHMLTGWSHIVEVSASRRIHSLENRSDILYRVLDASLSASASRDCSLRMAFVLSQVALTCMAKLRDERFLCS
GGLNSDNMTCLDVITAKKLSNGACHSILFKLIMAILRNESSESLRRRQYALLLGYFQYCQHMLDPNIPTSVMQFLMLEEQDSEDLDFQKIDKDQAELART
NFSIMRKEAQAILDLVINDATKGSEPGKTIALYVLVALICIDHERYFLSQLQSRGFLRTCLTSISNFSNQDGGHSLDSLQRASTLEAELALLLRISYKYG
KSGAQVLFSMGALEHLASCRAVSLQGCLRRFDRKLRRDVSVDFDKQCMIVTPMLRLLFSLTSLVDTSDIFEVKNKIVREVIDFVKGHQMLFDQILREDIS
TADELTVEQINLVVGILCKVWPYEESDEFGFVQGLFSMMRALFSCDSGAPTVGKLAQSSEVSQNKSKVELNSFRLCFSLGSYLYFLVTKMSLRLQVSDSS
IDYHSPAMLQQPTLMLLDSLLRSVATSLEMAAEEKSLLLNKIQDINELSRQEVDEIINMCVMQESVSSSDDIQKRRYIAMMEMSRVAGERDQLITLLLPL
AEHVLDIILVHFREGSMASDNSGATKAVTFGTHSDPRQDLSWMCGMLVPTLERLELLSEDKVGHNLKVFRRLVTSLKEMAIQNLAL
|
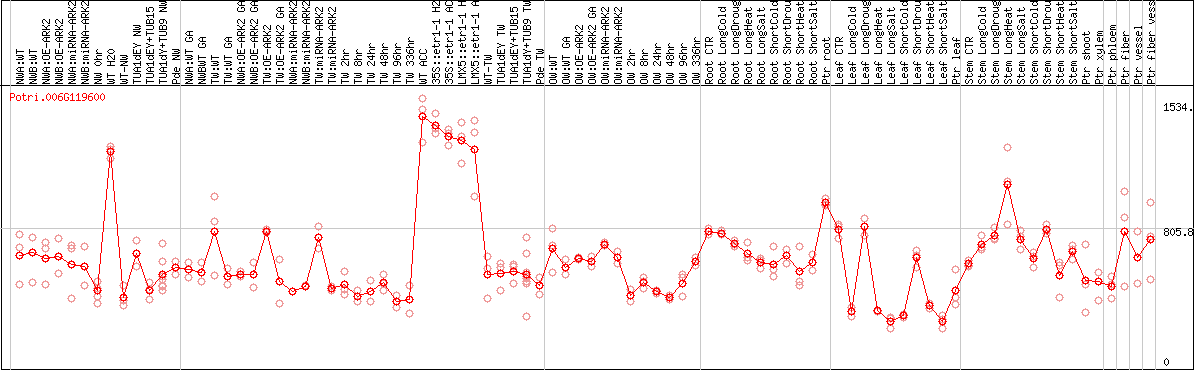
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G119600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
