Potri.006G120800 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.006G120800.3 pacid=42769840 polypeptide=Potri.006G120800.3.p locus=Potri.006G120800 ID=Potri.006G120800.3.v4.1 annot-version=v4.1
ATGGCTTCTTCAAGCTCCGGAAACTCAATAGCTGCATCAGAGGCTGTTCAGGTTTTGGTCTCTTCTCTGGCAGATGAATCCCCCACTGTTCGACAATCCT
CCATGGCTTCTCTGAAGGACATCGCTAGCTTGAATCCGCTTCTGGTTCTTGATTGTTGCTATACGGTTTCTCGAGGCGGACGTCGACGATTTGGGAACAT
GGCTGGGGTGTTTCAAGTGATGGCACTGGGAGTTAGTGCATTAAACGAAGGAGATGTTGATCCTTCTTTCATGGCAAAGCTTGCAAAGATTGCCACCACA
GAAATGATCTCGTCTAAGGAACTAAATGCTGACTGGCAAAGGGCTGCAGCAGGCCTATTGGTTTCAATTGGTTCCCATTTGCCTGACCTTATGATAGAAG
AAATTTTTCTTCATTTGTCAGGGCCGAGTTCAGCTTTGCCAGCTATGGTTCAAATACTCGCAGACTTTGCTTCGGCTAATGCTATACAGTTCATTCCACG
ACTGAAAGGTGTGCTTTCCAGAGTTTTACCTATTCTTGGAAATGTGCGAGATGTCCATCGGCCAATATTTGCAAATGCATTTAGATGTTGGTGTCAGGCT
GCTTGGCAATATAACATGGATTTCCCCTCACTTTCTCCTCTTGATGCTGATGTCATGTCGTTTTTAAATTCTGCCTTTGAGCTTTTATTGAGAGTATGGG
CAACTTCAAAGGATCTTAAGGGACCTCAACTGCCTTGTCAGCTTCTTCGAAAAGCAAGGACTTCACAATTAAAGAATGTTCGCACGTCCTCTGTGGAAGC
ATTGGGTCAGATGTTTGGTCTTGTCACCCGGACACAACTAAAGACAGCTTTACCCAGGCTTGTCCCTACTATACTGGAATTGTATAAAAAAGATCAAGAT
ATCGCCCTACTTGCAACTTGTAGTCTTTACAATCTCCTAAATGCCTCGTTACTGTCAGAAACTGGTCCGCCTTTGCTTGATTTTGAGGATCTCACAGTTA
TTCTATCAACACTCCTTCCGGTTGTTTGCTTTAATAATGATATCAAAGAGAATTCAGATTTTTCAGTGGGATTGAAGACCTATAATGAAGTTCAACGTTG
TTTTTTAACAGTTGGCTTGGTATACCCTGATGATCTGTTTACATTCCTCTTAAATAAATGCCGGTTGAAGGAAGAGTCTTTGACTTTTGGTGCGCTATGT
GTTTTAAAGCATCTCTTACCAAGGTCGTCTGAAGCTTGGCATAGTAAAAGGCCTTTGCTTGTTGAAGCTGTAAAGTCGTTGCTAGATGAGCAAAATTTTG
GTGTTCGGAAGGCGTTGTCTGAGTTGATTGTAGTAATGGCCTCACACTGCTATTTGGTTGGTCCATCTGCGGAGTTGTTCATTGAATACCTTGTATGTCA
TTGTGCTCTATCAGATCACAATAGAAATGATCCTGAGAACTCCAAGGTGAGGATAGGAGCCTTTTGTCCTACCAAGTTAAGAGCTGTGTGTGAGAAGGGT
CTTCTTCTACTAACTATAACAATTCCTGAAATGGAGCATATTCTATGGCCTTTTCTATTGAAGATGATTATTCCACGATCTTACACCGCTGCTACTGCGA
CTGTTTGCAGATGCATCTCAGAATTATGCAGAAACAGATCTTCTAATAGTAATAGTATGGTTAGTGAGTGCAAAGCTCGTGCTGATGTCCCGAGTCCAGA
GGAGCTTTTTGCTCGCTTGTTGGTGCTTTTACATGATCCACTATCAATGGAGCAGCTGGCAACTCAGATTTTGACAGTCCTTTGTTATTTGGCACCTCTC
TTCCCGAAGAATATCAACTTGTTTTGGCAAGACGAGATTCCTAAAATGAAGGCTTATGTCAGTGACACGGATGACCTAAAGTTGGATCCTTCATACCAGG
AGACTTGGGATGACATGATAATCAATTTTCTTGCAGAATCATTGGATGTGATCCAAGACACTAATTGGGTGATCTCTCTCGGAAATGCTTTCACACACCA
ATACGAGCTCTATACGTCGGATGATGAACATTCTGCTCTTCTCCACAGGTGCCTTGGAATGCTTCTACAGAAAGTTGATGACAGGGCTTATGTTCGAAAT
AAAATTGATTGGATGTATAAACAAGCTAGCATTGCTAATCCAGCCAACAGGCTTGGTTTGGCAAAAGCCATGGGATTGGTTGCAGCTTCTCACTTGGATA
CCGTGTTGGAAAAACTGAAAGTCATTCTGGATAATGTCGGGCAGAGTATTTTTCAAAGGTTGTTGTCTCTTTTCTCTGATATTTATAGAACAGAAGAGTC
AGATGATATACATGCTGCCTTGGCTTTGATGTATGGTTATGCTGCACGATATGCTCCATCAACAGTTATTGAAGCCAGAATAGATGCCCTTCTTGGGACT
AATATGCTTTCTCGGCTTCTTCATGTACGTCACCCTACAGCAAAGCAGGCTGTAATAACCGCTATTGATTTACTAGGCCGTGCTGTGATTAATGCTGCGG
AAAGTGGTGCATCATTCCCTCTGAAGAAAAGAGACCAAATGCTTGACTATATTCTAACTTTGATGGGGAGAGATGATGATGGTTTTGTTGATTCCAGTCT
TGAACTTCTGCGTACTCAGGCTCTGGCTTTAAGTGCCTGTACTACCTTGGTCTCTGTGGAGCCAAAGCTGACAATAGAAACAAGAAATTACATCATGAAG
GCAACCTTGGGGTTCTTTGCTTTGCCTAATGAGCCAGTGGATGTTGTCAATCCCCTCATAGAAAACCTTATCACTCTCTTATGTGCAATTCTTCTAACAA
GTGGAGAGGATGGAAGAAGCCGGGCCGAGCAGCTATTGCACATTATGAGACATACTGATCAGTATGTTTCTTCCTCTGAGGAGCATCAGAGGAAGAGAGG
CTGTCTTGCAGTTTATGAGATGCTTCTCAAGTTTCGGATGCTTTGCATCACTGGACATTGTGCCCTAGGTTGCCATGGAAGTTGCACTCACAGAAAACAA
ACTGACCGTACTCTTCATAGCACCATTTCCAATTTACCATCTGCATTTGTATTGCCAAGTCGTGAAGCCCTTTGTTTGGGAGAGAGGGTCATTAAATATC
TTCCACGTTGTGCAGACACTAATTCTGAAGTTAGAAAAGTTTCTGCTCAGATTCTTGATCAACTATTCAGCCTCGCTCTTTCACTTCCAAAGCCTTCAGG
TTTTAGCTTAAATGTGGATATAGAATTGCCCTACAGTGCCTTATCCTCTCTAGAGGATGTTATAGCTATCTTGAGAAGTGATGCTTCCATTGATCCATCA
GAAGTTTTTAACAGAATCGTGTCCTCAATCTGTCTTTTATTGACAAAAGATGAGCTTGTTGCCACCCTGCAAGGCTGCTCAGCAGCTATATGTGATAAGA
TCAAGCCATCAGCTGAAGGGGCTATACAAGCTATTATTGAGTTTGTTATGAAAAGGGGGAAAGAGCTAAGTGAAACGGACGTTTCAAGGACAACTCAATC
ATTGCTCTCTGCTGTGGTTCATGTTACAGAGAAGCATTTGCGTCTGGAGACTCTTGGAGCTATTGCTTCTCTAGCTGAAAGTACTAGTTCAAATATTGTC
TTTGATGAAGTATTGGCCACAGCTGGAAAGGATGTAGTTACGAAGGATATATCTAGACTACGAGGTGGCTGGCCAATGCAGGACGCATTCTATGCCTTTT
CTCAACACGCAGTTCTTTCTTTTCAATTCCTGGAGCATCTGATATCATTCCTTAACCAGACTCCTGTTGTTAAGAGTGATTTGGAGAAAGGAGACAACTC
TAGTCATTTAGCTGATGGTCAGATAGAGGATGACATTCTGCAAGCTGCTATGATTGCTCTCACTGCCTTTTTCAGAGGGGGTGGTAAAGTTGGAAAGAAA
GCAGTAGAGCAAAGCTATGCCTCTGTTGTTGTTGCACTCACACTCCAATTTGGAAGTTGTCATGGTCTAGCTAGTTCTGGTCAGCATGAGCCATTGCGAG
CTCTTCTAACAGCATTTCAAGCCTTTTGTGAGTGTGTCGGAGACCTTGAAATGGGAAAGATTTTGGCTAGAGATGGAGAGCAAAATGAAAAGGAGAGGTG
GATCAATCTTATCGGAGAATTAGCTGGCAGCATATCTATAAAGAGGCCAAAAGAAGTTCGGACTATTTGTGTAATTTTAACGGAATCTTTAAACCGACGT
CAAAAATTTCAAAGGGAAGCTGCTGCTGCGGCATTGTCGGTGTTTGTTCCCTATAGTGGGGGGTTTGATTCCCTGTTGGAGCAGATGGTTGAAGCATTGT
GCCGGCATGTTTCGGATGAGTCTCCAACAGTTAGACGTCTTTGTTTAAGAGGGCTTGTCCAGATACCATCGCTTCATATTTATCAGCACACAATCCAGAT
TCTTGGCATAATAGTGGCATTGCTTGATGATTTGGATGAATCTGTGCAATTAACTGCTGTTTCGTGCCTACTGATGATTCTTGAATCATCCCCAGATGAT
GCAGTAGAGCCCATTTTGCTCAATCTTTCTGTTCGGCTCCGCAATCTTCAAATATCCATGGATGTGAAAATGCGAGCTGATGCCTTTGCAGCATTTGGAG
CCCTGAGTAAATATGGTGTTGGGGCACAGCGTGAAATATTTCTTGAGCAGATACACGCTGCCATTCCTCGCCTGGTTTTGCACCTTCATGATGATGATCT
TAGTGTGAGACAGGCTTGCCGAAACACTCTGAAAAGGCTTGCCCACTTGATGGAGATGGAAGAATCAACTGCTCTATTCAACTCGCACTATTTTACTTCT
GATCACCGAAGTGACTATCAGGACTTTGTAAGAGATCTGACTAAGCAGTTTATCCAGCATCTTCCTTCCAGAGTTGATACTTACATGGCATCAACCATAC
AGGCATTTGATGCACCTTGGCCAATAATACAGGCTAATGCTATTTACTTGGTCAGCTGCTTGGTATCACTTTCAGATGATCAGCGCATTTTAGCCCTTTA
CCAAACACAGGTGTTTGGTACGTTGATGGGGAAGATGAGTCGATCACCAGACGCAATCGTGCGAGCTGCATGCTCTTCAGCACTTGGCCTATTACTAAAA
TCAACCAATTCTCTAGTATGGAGAACAGCTCGACTAGATTTAAAGTGA
|
|||||||||||||||
|
AA sequence
|
>Potri.006G120800.3 pacid=42769840 polypeptide=Potri.006G120800.3.p locus=Potri.006G120800 ID=Potri.006G120800.3.v4.1 annot-version=v4.1
MASSSSGNSIAASEAVQVLVSSLADESPTVRQSSMASLKDIASLNPLLVLDCCYTVSRGGRRRFGNMAGVFQVMALGVSALNEGDVDPSFMAKLAKIATT
EMISSKELNADWQRAAAGLLVSIGSHLPDLMIEEIFLHLSGPSSALPAMVQILADFASANAIQFIPRLKGVLSRVLPILGNVRDVHRPIFANAFRCWCQA
AWQYNMDFPSLSPLDADVMSFLNSAFELLLRVWATSKDLKGPQLPCQLLRKARTSQLKNVRTSSVEALGQMFGLVTRTQLKTALPRLVPTILELYKKDQD
IALLATCSLYNLLNASLLSETGPPLLDFEDLTVILSTLLPVVCFNNDIKENSDFSVGLKTYNEVQRCFLTVGLVYPDDLFTFLLNKCRLKEESLTFGALC
VLKHLLPRSSEAWHSKRPLLVEAVKSLLDEQNFGVRKALSELIVVMASHCYLVGPSAELFIEYLVCHCALSDHNRNDPENSKVRIGAFCPTKLRAVCEKG
LLLLTITIPEMEHILWPFLLKMIIPRSYTAATATVCRCISELCRNRSSNSNSMVSECKARADVPSPEELFARLLVLLHDPLSMEQLATQILTVLCYLAPL
FPKNINLFWQDEIPKMKAYVSDTDDLKLDPSYQETWDDMIINFLAESLDVIQDTNWVISLGNAFTHQYELYTSDDEHSALLHRCLGMLLQKVDDRAYVRN
KIDWMYKQASIANPANRLGLAKAMGLVAASHLDTVLEKLKVILDNVGQSIFQRLLSLFSDIYRTEESDDIHAALALMYGYAARYAPSTVIEARIDALLGT
NMLSRLLHVRHPTAKQAVITAIDLLGRAVINAAESGASFPLKKRDQMLDYILTLMGRDDDGFVDSSLELLRTQALALSACTTLVSVEPKLTIETRNYIMK
ATLGFFALPNEPVDVVNPLIENLITLLCAILLTSGEDGRSRAEQLLHIMRHTDQYVSSSEEHQRKRGCLAVYEMLLKFRMLCITGHCALGCHGSCTHRKQ
TDRTLHSTISNLPSAFVLPSREALCLGERVIKYLPRCADTNSEVRKVSAQILDQLFSLALSLPKPSGFSLNVDIELPYSALSSLEDVIAILRSDASIDPS
EVFNRIVSSICLLLTKDELVATLQGCSAAICDKIKPSAEGAIQAIIEFVMKRGKELSETDVSRTTQSLLSAVVHVTEKHLRLETLGAIASLAESTSSNIV
FDEVLATAGKDVVTKDISRLRGGWPMQDAFYAFSQHAVLSFQFLEHLISFLNQTPVVKSDLEKGDNSSHLADGQIEDDILQAAMIALTAFFRGGGKVGKK
AVEQSYASVVVALTLQFGSCHGLASSGQHEPLRALLTAFQAFCECVGDLEMGKILARDGEQNEKERWINLIGELAGSISIKRPKEVRTICVILTESLNRR
QKFQREAAAAALSVFVPYSGGFDSLLEQMVEALCRHVSDESPTVRRLCLRGLVQIPSLHIYQHTIQILGIIVALLDDLDESVQLTAVSCLLMILESSPDD
AVEPILLNLSVRLRNLQISMDVKMRADAFAAFGALSKYGVGAQREIFLEQIHAAIPRLVLHLHDDDLSVRQACRNTLKRLAHLMEMEESTALFNSHYFTS
DHRSDYQDFVRDLTKQFIQHLPSRVDTYMASTIQAFDAPWPIIQANAIYLVSCLVSLSDDQRILALYQTQVFGTLMGKMSRSPDAIVRAACSSALGLLLK
STNSLVWRTARLDLK
|
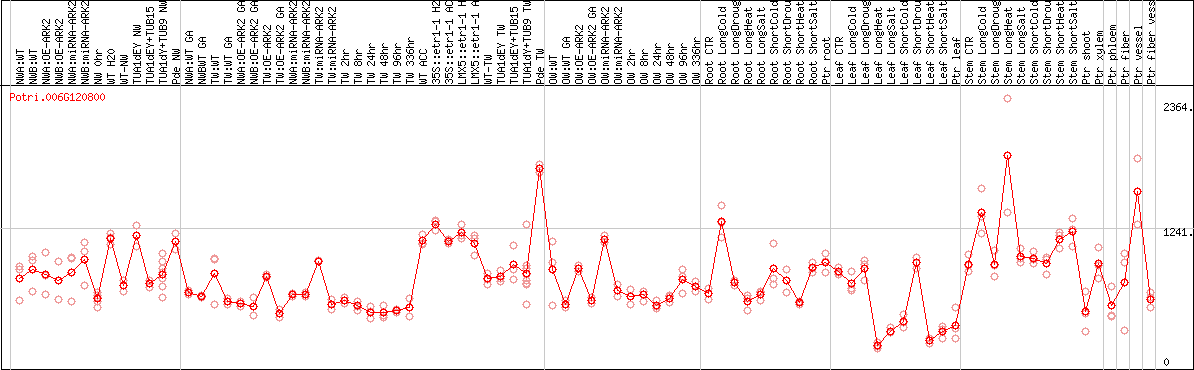
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G120800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
