External link
Symbol
Pt-TIP2.7
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G36830 345 / 6e-121
TIP1;1, GAMMA-TIP1, GAMMA-TIP
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
AT4G01470 325 / 4e-113
ATTIP1.3, GAMMA-TIP3, TIP1;3
tonoplast intrinsic protein 1;3 (.1)
AT3G26520 320 / 3e-111
TIP1;2, SITIP, GAMMA-TIP2, TIP2
SALT-STRESS INDUCIBLE TONOPLAST INTRINSIC PROTEIN, tonoplast intrinsic protein 2 (.1)
AT1G73190 245 / 2e-81
ALPHA-TIP, TIP3;1
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
AT1G17810 239 / 3e-79
BETA-TIP
beta-tonoplast intrinsic protein (.1)
AT3G16240 238 / 4e-79
DELTA-TIP1, ATTIP2;1, AQP1, DELTA-TIP
delta tonoplast integral protein (.1)
AT5G47450 224 / 1e-73
ATTIP2;3, DELTA-TIP3
DELTA-TONOPLAST INTRINSIC PROTEIN 3, ARABIDOPSIS THALIANA TONOPLAST INTRINSIC PROTEIN 2;3, tonoplast intrinsic protein 2;3 (.1)
AT4G17340 220 / 6e-72
TIP2;2, DELTA-TIP2
tonoplast intrinsic protein 2;2 (.1)
AT2G25810 216 / 2e-70
TIP4;1
tonoplast intrinsic protein 4;1 (.1)
AT3G47440 141 / 6e-41
TIP5;1
tonoplast intrinsic protein 5;1 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014411
373 / 7e-132
AT2G36830 390 / 4e-139
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10023913
372 / 2e-131
AT2G36830 391 / 3e-139
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10022611
370 / 8e-131
AT2G36830 403 / 5e-144
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10021510
366 / 2e-129
AT2G36830 402 / 9e-144
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10005885
335 / 7e-117
AT4G01470 395 / 1e-140
tonoplast intrinsic protein 1;3 (.1)
Lus10040863
330 / 3e-115
AT4G01470 394 / 2e-140
tonoplast intrinsic protein 1;3 (.1)
Lus10003288
308 / 3e-106
AT4G01470 388 / 6e-138
tonoplast intrinsic protein 1;3 (.1)
Lus10018256
246 / 8e-82
AT1G17810 370 / 7e-131
beta-tonoplast intrinsic protein (.1)
Lus10038324
244 / 6e-81
AT1G73190 380 / 1e-134
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
Lus10036187
243 / 7e-81
AT1G73190 382 / 4e-135
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00230
MIP
Major intrinsic protein
Representative CDS sequence
>Potri.006G121700.1 pacid=42770087 polypeptide=Potri.006G121700.1.p locus=Potri.006G121700 ID=Potri.006G121700.1.v4.1 annot-version=v4.1
ATGCCGATCAGAAACATCGCCGTAGGCCACTACCATGAGGCAACTCAGCCCGATGCCTTGAGGGCAGCCTTGGCTGAGTTCATCTCCACCCTTATTTTTG
TCTTTGCCGGTGAAGGGTCCGGCATGGCCTTCGCCAAACTTACGGACGGTGCTGCCAACACACCCGCTGGACTTATCGCTGCAGCTATCGCCCATGCTTT
CGCTCTTTTCGTTGCTGTGTCTGTCGGTGCCAACATCTCCGGTGGCCATGTCAACCCTGCTGTTACTTTCGGAGCTTTTATTGGTGGCAACATCACCCTC
TTGCGTGGCATCCTTTACTGGATCGCTCAGCTCCTGGGCTCCACTGTCGCTTGCTTGCTTCTCAAGTTCACCACTGGTGGCCTGGAAACCTCTGCTTTCG
CTCTGTCCTCTGGGGTTGGTGTCTGGAACGCTTTCGTTCTCGAGATCGTGATGACCTTCGGGCTTGTATACACAGTGTATGCCACAGCTGTTGATCCAAA
GAAGGGTAACTTGGGGATCATAGCTCCAATTGCAATTGGTTTCATTGTTGGTGCTAACATTTTAGCTGGAGGTGCATTTGATGGAGCCTCAATGAACCCA
GCTGTGTCATTCGGCCCTGCTTTGGTGAGCTGGACCTGGACCAACCACTGGGTCTACTGGGCAGGACCTTTGATTGGCGGTGGACTTGCCGGGCTCATTT
ATGAGTTCTTCTTCATTGGCTTCGGCAACCACGAGCAGTTGCCGACCGCTGACTACTAA
AA sequence
>Potri.006G121700.1 pacid=42770087 polypeptide=Potri.006G121700.1.p locus=Potri.006G121700 ID=Potri.006G121700.1.v4.1 annot-version=v4.1
MPIRNIAVGHYHEATQPDALRAALAEFISTLIFVFAGEGSGMAFAKLTDGAANTPAGLIAAAIAHAFALFVAVSVGANISGGHVNPAVTFGAFIGGNITL
LRGILYWIAQLLGSTVACLLLKFTTGGLETSAFALSSGVGVWNAFVLEIVMTFGLVYTVYATAVDPKKGNLGIIAPIAIGFIVGANILAGGAFDGASMNP
AVSFGPALVSWTWTNHWVYWAGPLIGGGLAGLIYEFFFIGFGNHEQLPTADY
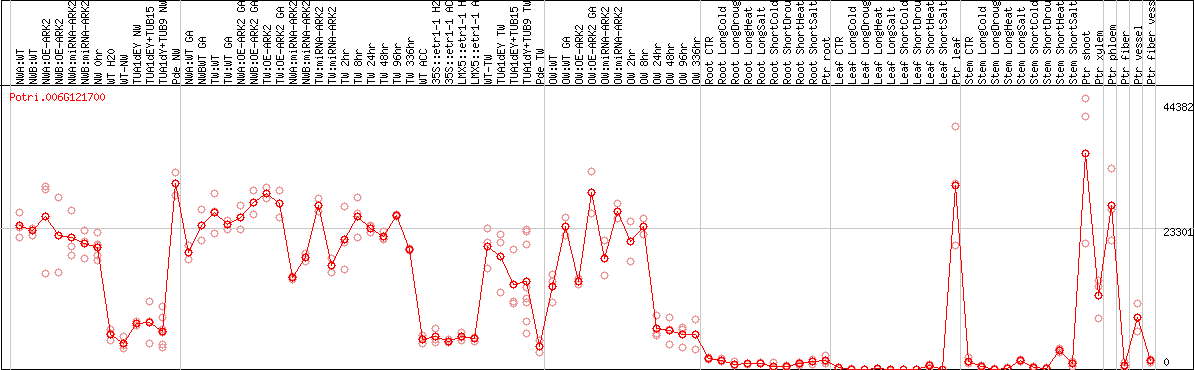
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G121700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.