Potri.006G137300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G137300.1 pacid=42769953 polypeptide=Potri.006G137300.1.p locus=Potri.006G137300 ID=Potri.006G137300.1.v4.1 annot-version=v4.1
ATGAAATCAACTTCGTCGTTTGGGAGGACGAGCTCCAGGCTATCCAATTACAGCCGCACGTTTGACTTACCAGACGATGATCTCGACGGTTCAGAAACCG
CCAGCGACTGCGAGATTGGCGGTGCAATGTTGCCGATTTTCTTGAACGACCTGAGAAGAAACAACCAAGATGAGTTAGTTGAGGTCACGCTCGAGCTCGA
GAAAGACTCCATTGTTGTTTGCAGTGTTAATCCGAACGGTAGAGGTATGTCAACTCCGAGTACGCCTGGCGGTGGTGCTGCGGCGGGGATTTTAGAGAGG
AGTTTGTCGGCAACGTCGAGGATTAGACGGAAGTTCGGGTGGTTAAGATCAAGATCTTCGAGAACGACATGGTCGGAGAATGAAGAGAGAGTAATCTCGG
CGAGAGATGCACGGAAATTGAAAGCCAAGCTTGACAGGTCAAAACTGAGCGCTCAAAGAGGACTGAAAGGGTTGAGGTTTATAAATAAGACAACCGGTAC
GACGTCTGAGTCGAATGAGCTGTGGAAACGAGTCGAGTCCAGGTTCAAGTTGCTTTCTAAAGACGGTTTGCTTGCTAGAGAAGATTTTGGTGAATGTATA
GGGATGGTGAATTCGAAGGAGTTTGCGGTGTGTATATTTGATGCACTAGCGAGGAGGAAGAGGCAGAGAATAACGAAAATAAGCAAAGAAGAGCTTCATG
ATTTCTGGTTACAGATTTCTGACCAGAGTTTTGACGCGCGCCTTCAAATTTTCTTTGACATGGCAGATAGCAATGAAGACGGAAGGATCATAAGAGAAGA
AGTGCAGGAGCTTATAATGCTCAGTGCTTCTGCAAATAAACTGTCAAAGTTGAAAGAACAAGCAGAAGAATATGCTTCGTTGATAATGGAAGAACTTGAT
CCTGAAAACCTCGGATACATTGAGTTATGGCAGTTAGAGACACTTCTTCTACAAAGGGACACATACATGAACTACAGCAGACCATTGAGTACAGCAAGTG
TAAGCTGGAGTCAGAACATAAGTTCAATCAAACCCCGGAACGTGATGCATAGGCTAAGCTTTAAATTGAGGAATTTAATTCTAGAGAAATGGCAAAGGGC
ATGGATTTTATCGCTGTGGGTGATGATAATGGTTGGCCTATTCGTCTGGAAATTCCTTCAATACAAGAACAAGGCTGCATTTCACGTCATGGGCTATTGC
TTGGCGAGTGCCAAAGGTGCTGCAGAGACTCTCAAATTCAACATGGCTCTAATTCTATTACCTGTTTGTCGAAACACACTGACTTGGCTTCGGTCCACTC
GAGCAAGGTCATTTGTTCCTTTTGATGAAAATATTAATTTCCACAAGATGGTTGCAGGCGCTATAGTAATTGGAGTCATCTTGCATGCAGGAAACCATCT
ATTATGTGACTTTCCCCGTCTGATAAACTCGTCTCCTGAAAATTTTGCCTTGATTGCGTCTGATTTCAATAATAAAAAGCCTACATATAAAGAACTTGTG
ACTGGTATTGAAGGCGTGACTGGGATTTCTATGGTGGTGTTATTGACCATTGCATTCACTCTGGCAACAGGCCGTTTCCGGAGAAATGGGGTGAGACTAC
CTGCACCCTTCAACAAATTGACAGGTTTCAATGCGTTCTGGTACTCTCATCATCTTACTGGTGTGGTCTATATTCTGCTGCTCGTCCATGGAACCTTCTT
GTTCTTAGCCCACAAGTGGTATCAGAAAACAACGTGGATGTATATCTCTGCTCCCTTGCTGCTCTACATGGTTGAACGAAATGTGAGAACACGTAGATCA
GAACATTATTCTGTAAAATTATTGAAGGTGTCAGTGCTACCGGGAAATGTCCTAAGCTTAATCTTGTCCAAGCCACAGGGATTTAAGTACAAAAGTGGGC
AGTACATTTTTCTACAATGCCCAGCAATTTCCTCCTTTGAATGGCACCCATTCTCTATAACCTCAGCACCAGGGGATGACTATCTCAGTGTTCACATCCG
GATAGTCGGAGACTGGACGGAGGAATTGAAGCGAGTTTTCACCGAGGAAAATGATTCACCATCTGTTATTGGTCGTGCCAAATTTGGTCAACTTGGGCAC
ATGGATCAAACAAGACAACCAAAGCTGTATGTTGATGGTCCATATGGGGCTCCAGCACAAGACTACAGAAATTACGATGTCTTGCTTCTCGTGGGGCTTG
GCATAGGAGCTACCCCTTTCATAAGCATCCTTAGAGATCTTTTGAATAATACACGAACAGCAGATAATCAAATGGATTCAAACACTGAAAACAGTCGGTC
AGATGACAGTTCAAATAGTTATGCATCTTCAAGTATGACACCAGTTAGCAAGAAGAGAACACAGAGGACTACCAATGCTCATTTCTACTGGGTTACTAGA
GAGCCTGGATCTTTTGAATGGTTTAAAGGAGTCATGGATGAAGTTGCAGAGATGGATCACAAAGGTCAAATCGAACTGCACAACTACCTTACAAGCGTTT
ACGAAGAAGGTGATGCAAGGTCAACTTTGATCACCATGGTCCAAGCTCTCAATCATGCGAAACATGGTGTTGACATCGTGTCAGGCACTCGAGTGAGGAC
ACACTTTGCAAGACCAAATTGGAAAGAAGTGTTCAACAAAATTGCTTCAAAGCATCCGTTTGGCACAGTAGGGGTGTTCTATTGCGGGATGCCAGTGTTG
GCAAAGGAGCTGAAGAAGCTATGCCAGGAGCTGAGTCACAAGACAACCACAAGGTTTGAGTTTCACAAGGAGTATTTCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G137300.1 pacid=42769953 polypeptide=Potri.006G137300.1.p locus=Potri.006G137300 ID=Potri.006G137300.1.v4.1 annot-version=v4.1
MKSTSSFGRTSSRLSNYSRTFDLPDDDLDGSETASDCEIGGAMLPIFLNDLRRNNQDELVEVTLELEKDSIVVCSVNPNGRGMSTPSTPGGGAAAGILER
SLSATSRIRRKFGWLRSRSSRTTWSENEERVISARDARKLKAKLDRSKLSAQRGLKGLRFINKTTGTTSESNELWKRVESRFKLLSKDGLLAREDFGECI
GMVNSKEFAVCIFDALARRKRQRITKISKEELHDFWLQISDQSFDARLQIFFDMADSNEDGRIIREEVQELIMLSASANKLSKLKEQAEEYASLIMEELD
PENLGYIELWQLETLLLQRDTYMNYSRPLSTASVSWSQNISSIKPRNVMHRLSFKLRNLILEKWQRAWILSLWVMIMVGLFVWKFLQYKNKAAFHVMGYC
LASAKGAAETLKFNMALILLPVCRNTLTWLRSTRARSFVPFDENINFHKMVAGAIVIGVILHAGNHLLCDFPRLINSSPENFALIASDFNNKKPTYKELV
TGIEGVTGISMVVLLTIAFTLATGRFRRNGVRLPAPFNKLTGFNAFWYSHHLTGVVYILLLVHGTFLFLAHKWYQKTTWMYISAPLLLYMVERNVRTRRS
EHYSVKLLKVSVLPGNVLSLILSKPQGFKYKSGQYIFLQCPAISSFEWHPFSITSAPGDDYLSVHIRIVGDWTEELKRVFTEENDSPSVIGRAKFGQLGH
MDQTRQPKLYVDGPYGAPAQDYRNYDVLLLVGLGIGATPFISILRDLLNNTRTADNQMDSNTENSRSDDSSNSYASSSMTPVSKKRTQRTTNAHFYWVTR
EPGSFEWFKGVMDEVAEMDHKGQIELHNYLTSVYEEGDARSTLITMVQALNHAKHGVDIVSGTRVRTHFARPNWKEVFNKIASKHPFGTVGVFYCGMPVL
AKELKKLCQELSHKTTTRFEFHKEYF
|
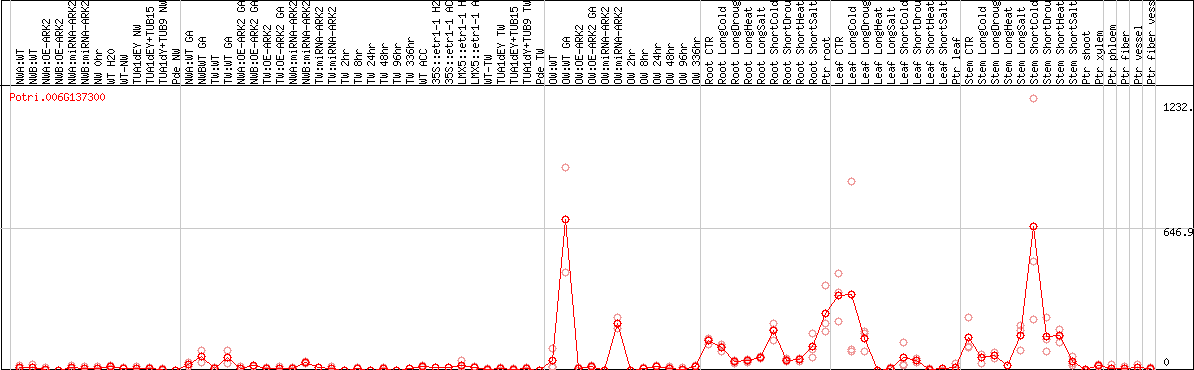
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G137300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
