Potri.006G141900 [POPLAR]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Flax homologues |
|
||||||||||
| PFAM info |
| ||||||||||
|
Representative CDS sequence |
>Potri.006G141900.3 pacid=42769718 polypeptide=Potri.006G141900.3.p locus=Potri.006G141900 ID=Potri.006G141900.3.v4.1 annot-version=v4.1
ATGACCAAAATACAAGCTTACGAGGACTTCGTGAAGGTCCACGGCATCCTTTTAGCAGCCTCAGGACTACCTCGTACACTGCACCGCAAACTGTTCGACA
AACTTACTTCCGAGACTTTCGATGGCGGCGCCTACTTCCAAGTCGACCCATGCCAGGATGGCCGTCAGAGGCGCCTCCTCCTTACCTCTGCCGCCTCCAT
GCCCAAAGATTCCAACGTTTTTCTCATTGACCACGCCTGGACTTTTCGCCTCTCCGATGCTTATAAACAGCTTCAAGAAGTGCCAGGGTTAGCCCAGAGA
ATGGCTGCTTTGATGTGTGTGGATATTGATTCAAATTCCGATGTTGAAGAGATAGATGGGGATGGAGTTTCACGAGACACCTATTCAAAATTGAATGTCA
CTGACATTGTTGAGAATGAAATTGGTTATGCGAAAGAGAGAGGCTATGACACTGTTAAGTGGTTGGAGCTTGAGGAACTTGACATTGACGATGACATGCT
TCTCTCTCTTGATTTGTCTAGTAAATTTCCGGATTTACTAGCACTTAGCCTATGTGGAAACAAGCTCGAGAATGTTGAAATAGTTGTGCAGGAAGTTACC
AAATTGAAAAACCTAAAAGCTCTCTGGCTGAATAACAATCCAGTTCTAGAAAACTGTGATGGTTGTATGGCAGATACAATTTTCAAGGGATGTCCAGGGC
TCGAAATCTACAACTCATGTTTCACTAGTAATTTTGGTGAGTGGGCCTTGGGATTTTGCGGAGGAGTTTATGAGAAGGATAATCCTTGCCCTATCCATCA
GGACAACCATCCATTGCAGAGTGTGACATCTCTCGACCTTTCAAACAGAAGTATTCATTCTCTCATTAACAAGGCATTTTCTCCTGTTGAGATGCCTTCT
CTTTCACATCTGAATATTCGAGGAAATCCTCTGAAGCAGAATTCAGTTAGTGAGCTGTTCAAAGTATTGAAGGGGTTTACTAGTTTGCAAACTTTAGAGG
TGGATTTACCTGGACCCCTAGGAGAGAGTGCTATAGAAATTCTTGAATCTGTTCCAAATCTTTCTCAGTTGAATGGCGTCAATGTATCAAAAATATTAGA
AACTGGAAACCATGTAATCGATGCTGTACTTCAGCCACGCCTTCCTGAATGGACAGCTGAAGAGCCTCTTGCTGATCGTGTTATTAATGCAATGTGGTTA
TATTTGATGACATATAGACTTGCAGACGAGGAAAAGATTGATGAAACCTCTGTGTGGTATGTAATGGATGAACTAGGTTCAGCTTTAAGGCACAGTGATG
AACCAAATTTCAGGGTGGCTCCTTTTCTCTTCATGCCAGAGGGAAATCTGGATTCTGCTGTGAGCTATTCCATATTATGGCCAATCCAGAATGTTCAAAA
TGGAGATGAATGCACTCGTGATTTCCTATTTGGTATTGGGGAGGACAAGCAACGTTCTGCCAGGCTTACAGCATATTTCCATACACCACAATATTATTTT
ATTCAGGAGTATGAGAAATTCCATCAGAAGTTGCAATCAAAAAGTTCAACTCCCTTGCCAGTGAAGTCATCTTCCAGCAGAACCTTACGTCGTACTGATG
GATGTGCTTTGCGTGTGTACACTGACTTACCTCAAGTGGAAGGATTTTTGACACGTACTGAATTTATAATCACCACAGAGTTGAAGGATGCAGACATTAT
ATGGACAGGCATGCAGGTGGATGATGATGTCAAAAGAGCTGCTGGAATTACAGATCAGCAATATATAAACCAATTTCCTTTCGAAGCTTGTCTTGTCATG
AAACATCATTTGGCAGAGACTATTCAAAAGGCACATGGATCTCCTGATTGGTTGCATCCTACTTACAATTTAGAATCACATCTCTCTCAACTTATTGGCG
ACTATTATGCACGCAAAAGAGATGGAATGAACAACCTATGGATCTTAAAACCATGGAACATGGCCCGGACTATTGATACAACTGTAACAGATAATTTGTC
TGCTATTATCCGCCTGATGGAAACTGGTCCAAAAATATGCCAGAAATATATCGAGCATCCTGCTCTGTTCGAAGGCAAGAAATTTGATATCCGTTATATA
GTACTGGTTCGCAGTGTGAAGCCTCTGGAACTGTTCCTTGCAGATGTTTTCTGGGTTAGGTTGGCAAACAACCAATACACTCTTGACAAGCACAGCCTTT
TTGAGTACGAGACCCACTTTACTGTTATGAACTATCGTGGAATATTGAATCACAAGAACACTCCAGAGTTTGTGAAGGAATTTGAACAGGAGCACCAAGT
TAAATGGTTGGATATCCATGAGAGAGTGAGAAATATGATTCGATCTGTTTTCGAGGCAGCTGCAACAGTACATCCAGAGATGCATAGTCCAATGTCCAGG
GCAATGTATGGCGTGGATGTCATGCTTGATAGCTCTTTCCAGCCAAAGTTATTGGAGGTCACTTACTGTCCAGACTGTACAAGAGCATGCAAGTATGATA
CACAAGCTATAGGTGGAGGGGGAGAATTATTGAAAGGTAGTGATTTCTACAATTATGTGTTTGGTTGTCTCTTTCTTGATGAAACTAGACATGTGTGCCC
ATTGTAA
|
||||||||||
|
AA sequence
|
>Potri.006G141900.3 pacid=42769718 polypeptide=Potri.006G141900.3.p locus=Potri.006G141900 ID=Potri.006G141900.3.v4.1 annot-version=v4.1
MTKIQAYEDFVKVHGILLAASGLPRTLHRKLFDKLTSETFDGGAYFQVDPCQDGRQRRLLLTSAASMPKDSNVFLIDHAWTFRLSDAYKQLQEVPGLAQR
MAALMCVDIDSNSDVEEIDGDGVSRDTYSKLNVTDIVENEIGYAKERGYDTVKWLELEELDIDDDMLLSLDLSSKFPDLLALSLCGNKLENVEIVVQEVT
KLKNLKALWLNNNPVLENCDGCMADTIFKGCPGLEIYNSCFTSNFGEWALGFCGGVYEKDNPCPIHQDNHPLQSVTSLDLSNRSIHSLINKAFSPVEMPS
LSHLNIRGNPLKQNSVSELFKVLKGFTSLQTLEVDLPGPLGESAIEILESVPNLSQLNGVNVSKILETGNHVIDAVLQPRLPEWTAEEPLADRVINAMWL
YLMTYRLADEEKIDETSVWYVMDELGSALRHSDEPNFRVAPFLFMPEGNLDSAVSYSILWPIQNVQNGDECTRDFLFGIGEDKQRSARLTAYFHTPQYYF
IQEYEKFHQKLQSKSSTPLPVKSSSSRTLRRTDGCALRVYTDLPQVEGFLTRTEFIITTELKDADIIWTGMQVDDDVKRAAGITDQQYINQFPFEACLVM
KHHLAETIQKAHGSPDWLHPTYNLESHLSQLIGDYYARKRDGMNNLWILKPWNMARTIDTTVTDNLSAIIRLMETGPKICQKYIEHPALFEGKKFDIRYI
VLVRSVKPLELFLADVFWVRLANNQYTLDKHSLFEYETHFTVMNYRGILNHKNTPEFVKEFEQEHQVKWLDIHERVRNMIRSVFEAAATVHPEMHSPMSR
AMYGVDVMLDSSFQPKLLEVTYCPDCTRACKYDTQAIGGGGELLKGSDFYNYVFGCLFLDETRHVCPL
|
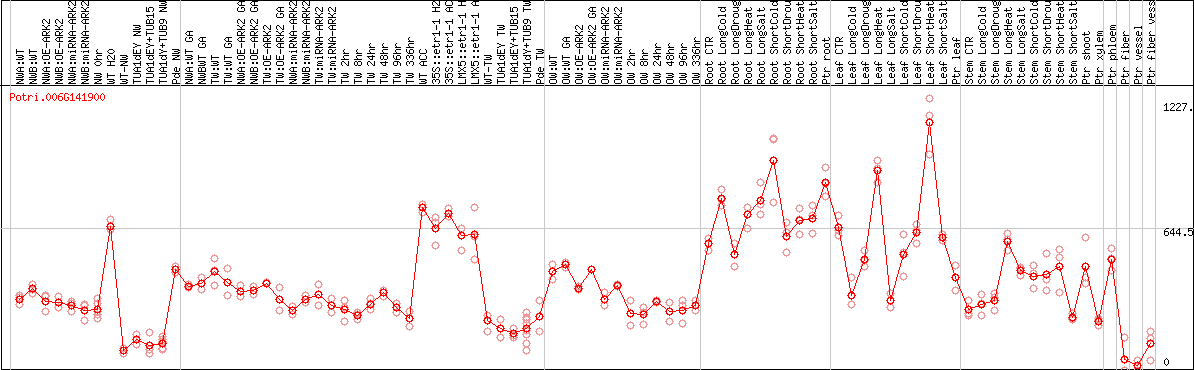
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G141900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
