Potri.006G146600 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G146600.1 pacid=42769657 polypeptide=Potri.006G146600.1.p locus=Potri.006G146600 ID=Potri.006G146600.1.v4.1 annot-version=v4.1
ATGGCAACATCTAGTAAGTTCGATATGTCTTCTGATAGCCCGGATAGGCCGATATACTCTAGTGGCCATCGTGGATCCCACTTAGCTGCTCAGATGGACA
GATCGAGTAGCTTTCGTGAGAGTATGGAGAATCCAATCCTATCTTCTGTCCCGAACATGGCTCGAAGTAGTGCTGTGGTAGCGCAGGGAGACGTGGTGAA
CTTCTTCCAATGCATGCGTTTTGATCCAAAGGTGGTGGCTGCAGACCATAAATCTAGTCGTCAAGGGGACTTTAAGCGACACATGAATGCTGCTCTTGGA
ATTTCAGCAGATGACTCTTCTGGTTCTTTGAAAGGCAAGGTGGTATTGTCTCCCTCACCTGAAGAAATCAAGCGAGTAAGAGATGGTCTGCGTGGAAGCT
CTGCCAAGGGCAGGGAACGTGTGAAGATTTTCACTGAAGCCTTGTCTGCATTCAACAAGCTCTTCCCAACCATACCGTCAAGGAAGAGGTCTAGATTAGA
AGGTTATTCTAATGATAGGCCTAATGCTTCAGTATCAAGCGATCGGTCGGTCCTTGTTCCAAGCCTGGGTAAAATGGGAATTCAAAATCATTCTGCCACC
AGTGGTTTTGAACTTGAACAGCAAAAGTCAGAAGAACGGACCAAAAATATTGTTCCAAACAAGCGCACACGAACTTCTTTGGTGGATGTGCGGGGTAATG
CTCTGGTCAGGCCATCTGGTACTGTTGACAGAGACAGGGAAATGCTAAGGCTTGCAAATAGTGGTGCTGTTCAAGGTGAGGATCGAAGTTTATCAATTGG
TGTTGATGGTTGGGAAAAGACAAAAATGAAGAAGAAGCGTTCTGGGATAAAGCCAGATGTTGCTTCAAACATGGTGTCAACTAAACCCTCAGATGGCTAT
CGGGAATCAAAACAGGGAGCGCTGCAAAGACCTGGTACTGATGCTCGTTCAAGATTGAATATTGATTCCCATGGGTTCAGGCCTGGAGTTTCTAATGGTG
CTGTTGGAGTTGGAAAAATAGATGGTATCTCACAACCAACAGGCTTGAGCGTGCGATCGATGACCCCTAGGACTGACCTGGAGAACAGTTCTCTTCTAAA
TGATAGGAGAGAACGCCCTCTTGGTTCAGATAAGGAAAGGGTGAATATCAGAGCTGTTACCAAGGCTGTTCGGGATGATTTTAATTCTGCTAGTCCCACT
TCCAGTGCAAAAATGAATCCGTCCATTCGGGCTCCCCGATCAGGTTCAGGCATTATGCCCAAGTTGTCCCCAGTTGTCCATAGAGCAACTGCTCCCAATG
ATTGGGAGCTCTCTCATTGTACGAACAAGCCGCCTGCTGTTGGAGCTAACAACCGCAAACGCACAGCATCTGCTCGGTCTTCGTCTCCACCTGTTGCTCA
CTGGGCTGGTCAGAGGCCACAGAAGATCTATCGAACAGCTAGAAGAACAAATTTGGTTCCCATTGTTAACAATGATGAAAGTCCCACTTTGGATAGTGTA
TCTGATGTTTCAGGTAATGAAATTGGGGTGGGATTTGCCAGGCGCTTGAGCGGCAATTCTCCTCAGCAAGTTAAATTAAAAGGTGACACTTTATCGTCAG
CTGTTTTATCAGAAAGTGAAGAATCAGGGGCTACAGAGGTTAAATCTAAAGATAAGAGCAGAAAGTCTGATGAGATAGATGAGAAAGCTGGGCAGAATGT
TCAAAAGATATCACCTTTGGGCCTTCCATCAAGAAAGAACAAGCTGGTTTCAGGTGAAGATATTGGAGATGGTGTTCGCAGGCAAGGGAGGACTGGGAGG
GGTTTTACTTCAACAAGGTCTCTTGTGCCAACAGCAGTTGAGAAACTGGGCAATGTTGGAACAGCAAAACAGCTTAGAAGTGCCAGACTTGGTTTTGATA
AAAATGAAAGCAAGACAGGCCGTCCTCCAACCAGAAAACTCTCTGATCGTAAGGCTTATACACGCCAAAAGAATACAACAGTCAATGCAACAGCAGATTT
TCTTGTTGGTTCAGAAGATGGGCATGAAGAGCTATTGGCTGCTGCAAGTGCTGTTATTAATCCTGGCCTAGCCCTTTTGAGCTCATTTTGGAGGCAAATG
GAGACATTCTTTGGTTTTATATCTGATGTAGACATTGCTCACTTGAAACAACAGGGAAGTATTGTGTTTACTGCACCGTCAGCAACACCAGTCCATTCTG
ATGCAAACAACTACAGTACTGTCCCTAATGGGTATGGGTTATTTGAACATGATAGGGAGGTTGAGCTTGAGCTTGCTGCTGAAACTAGGACTTCAGAACT
TCTTCCAGATCAGTTAATGCCTGTAGATAGAGAAATTCCCCTTTCTCAGTTACTCTTGGCAGCCCTGACCTCAGAGGAAGATTGTACCCTTGGAAATGCA
GACCTGGAATTTGATGCATATGGAACTGATTTCGAGCTACATGAGGAGTTGGAATCAAATTGTGTGAATCACCTAGATAATTTTCAGTTTTCTGGACATG
TTGCATTTAGTGGTTGCAAGGTGAGTGGGAAGCCAGACCACGACGAGACAGATAATGATATTTCAGGCATCCCAAACATGGGTATTGACTCAAGCTTCAG
AAATACCATAAATGGTGTCCTTTCAGACCATGCATTGGTACCAGGGATGGCCTGTTCCAAATTTCAATATGATAATATGAAAATAGAAGAAAAACTTCGT
TTGGAGGTTCTTAGTCTGGGAATTTTTCCTGAATCAATGCCTGACATGCCGATGGATGATGAAGGAATCTGTGGGCACATCAGCAAATTAGAGGAGAATC
AACATGGGCAGGTTTCAAGGAAGAAAGGTTTATTAGACAAGCTGTTGAAACATGCCTCAGAAATGAAAGAACTCCAAGAAAAGGAATTTGAACAACGTGC
TCATGACAAACTCGTCACAATGGCTTATGAAAAGTACATGACTTGTTGGGGTCCTAATGCCACTGGTGGGAAGAGTTCCAGTAGCAAAATGGCCAAGCAA
GCTGCCTTAGCATTTGTTAAACAAACTTTAGAGCGATGTCATAAATTTGAAGTTACAGGAAACAGCTGCTTCAGTGAGCCCTCCTTTAGGGATATGTTCC
TTTCTGGGACTGCCCGCCTAAATGGTGCACAATCAGTGGATACTCCTACTGATGGTGAATCTGCAAAGTTGTATGGCAATACCTCAACCCGCTCTTTGGA
AGCCAGAGTTTCAGCTTCCATGGGTTCACAGCCAAGTCCTCGAACTTTACATGTGGGTCAGAATGGGGATAGCCACATCTCAAATCCTTCTGATCTACTT
CCACCTGTTAATCGCTTATCGGAACAAATTACTGGTAAGGAAGATACGTGGTCAAACAGGATGAAGAAGAGGGAGTTGTTGCTTGATGATGTTGTTGGTA
GCCCATCAAGTGCTCCATCAGGAATAGGAGGTTCGCTATCGAGCAGTACAAAAGGAAAGAGGAGTGAGAGAGACAGGGAAGGAAAGGGGCACAACAGAGA
GGTTTTATCTAGAAACGGATCTAATAAAATTGGTCGACCAACATTATCTAATCAAAAGGGGGAGCGGAAAACTAAAACGAAGCCCAAGCAAAAAACAACT
CAACTGTCTGTTTCTGTGAATGGCCTTGTTGGCAAGATATCGGAGCAACCTAAAACAACTTTGCCTTCCAAGGCAAAGTCAAGTGAAAACAACAGTAATA
GCAAAGCAAAGGAAAAGGATAGGTTTGGCTTGGATGTGTTGGATGATGCTATTGATTTGTCCAACCTGCAGCTGCCAGGAATAGATGTATTAGATGACAG
CCAAGGGCAGGATCTGGGTTCTTGGCTGAACATTGATGATGATGGATTACAAGAGCATGGTGACATTGATTTCATGGGCCTTGAAATTCCAATGGATGAC
CTTTCAGATCTAAATATGATGGTTTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G146600.1 pacid=42769657 polypeptide=Potri.006G146600.1.p locus=Potri.006G146600 ID=Potri.006G146600.1.v4.1 annot-version=v4.1
MATSSKFDMSSDSPDRPIYSSGHRGSHLAAQMDRSSSFRESMENPILSSVPNMARSSAVVAQGDVVNFFQCMRFDPKVVAADHKSSRQGDFKRHMNAALG
ISADDSSGSLKGKVVLSPSPEEIKRVRDGLRGSSAKGRERVKIFTEALSAFNKLFPTIPSRKRSRLEGYSNDRPNASVSSDRSVLVPSLGKMGIQNHSAT
SGFELEQQKSEERTKNIVPNKRTRTSLVDVRGNALVRPSGTVDRDREMLRLANSGAVQGEDRSLSIGVDGWEKTKMKKKRSGIKPDVASNMVSTKPSDGY
RESKQGALQRPGTDARSRLNIDSHGFRPGVSNGAVGVGKIDGISQPTGLSVRSMTPRTDLENSSLLNDRRERPLGSDKERVNIRAVTKAVRDDFNSASPT
SSAKMNPSIRAPRSGSGIMPKLSPVVHRATAPNDWELSHCTNKPPAVGANNRKRTASARSSSPPVAHWAGQRPQKIYRTARRTNLVPIVNNDESPTLDSV
SDVSGNEIGVGFARRLSGNSPQQVKLKGDTLSSAVLSESEESGATEVKSKDKSRKSDEIDEKAGQNVQKISPLGLPSRKNKLVSGEDIGDGVRRQGRTGR
GFTSTRSLVPTAVEKLGNVGTAKQLRSARLGFDKNESKTGRPPTRKLSDRKAYTRQKNTTVNATADFLVGSEDGHEELLAAASAVINPGLALLSSFWRQM
ETFFGFISDVDIAHLKQQGSIVFTAPSATPVHSDANNYSTVPNGYGLFEHDREVELELAAETRTSELLPDQLMPVDREIPLSQLLLAALTSEEDCTLGNA
DLEFDAYGTDFELHEELESNCVNHLDNFQFSGHVAFSGCKVSGKPDHDETDNDISGIPNMGIDSSFRNTINGVLSDHALVPGMACSKFQYDNMKIEEKLR
LEVLSLGIFPESMPDMPMDDEGICGHISKLEENQHGQVSRKKGLLDKLLKHASEMKELQEKEFEQRAHDKLVTMAYEKYMTCWGPNATGGKSSSSKMAKQ
AALAFVKQTLERCHKFEVTGNSCFSEPSFRDMFLSGTARLNGAQSVDTPTDGESAKLYGNTSTRSLEARVSASMGSQPSPRTLHVGQNGDSHISNPSDLL
PPVNRLSEQITGKEDTWSNRMKKRELLLDDVVGSPSSAPSGIGGSLSSSTKGKRSERDREGKGHNREVLSRNGSNKIGRPTLSNQKGERKTKTKPKQKTT
QLSVSVNGLVGKISEQPKTTLPSKAKSSENNSNSKAKEKDRFGLDVLDDAIDLSNLQLPGIDVLDDSQGQDLGSWLNIDDDGLQEHGDIDFMGLEIPMDD
LSDLNMMV
|
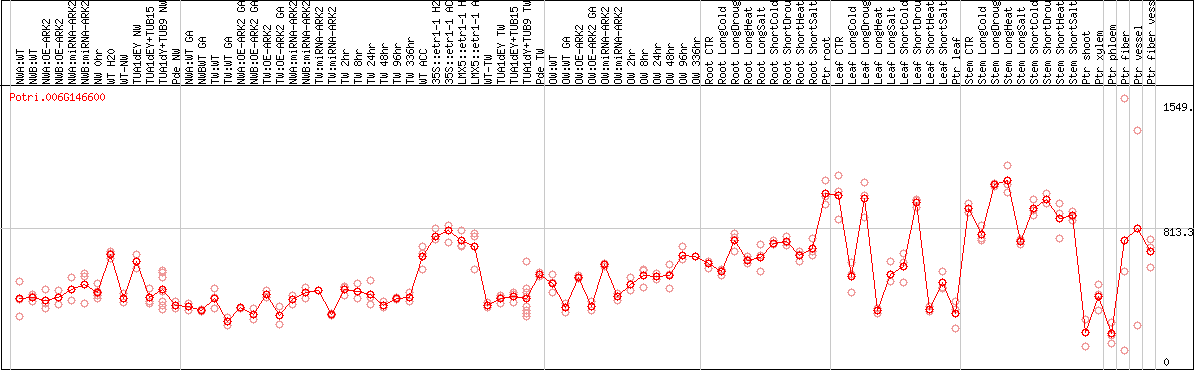
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G146600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
