Potri.006G149800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G149800.4 pacid=42766997 polypeptide=Potri.006G149800.4.p locus=Potri.006G149800 ID=Potri.006G149800.4.v4.1 annot-version=v4.1
ATGCTTCATCGGAGTTTCAAGCCTGCCAAATGCAAAACATCGTTGAAGTTAGCGTCTTCGCGAATAAAGCTGTTGAAGAATAAGAGAGAAGCACAGGTGA
AGCATCTCAAGAGAGAATTGGCTCAATTGCTGGACGCTGGCCAAGAACGGACTGCCAGAATCCGGGTTGAACATGTGGTTAGGGAAGAGAAGACTATGGC
AGCATATGAACTTATCGAGATATATTGCGAGCTTATTGTGGCGCGTTTGCCCATCATTGAGTCGCAGAAAAACTGTCCCATTGACTTGAAGGAAGCGGTC
TCAAGTGTAATTTTTGCCTCTCCAAGATGTGCTGATGTACCAGAATTGATGGACATTCGGAAGCATTTAACAGCAAAATATGGGAAAGAATTTGTATCTG
CAGCAGTTGAATTACGTCCAGACTGTGGTGTTAGCCGCCTATTGGTGGAAAAATTATCTTCCAAATCACCTGATGGCCCTACCAAAATTAAAATTTTGAC
TGCAATCGCTGAGGAACACAATATCAAGTGGGATCCTATGTCATTTGAAGAGAAAGACACCAAACCTCCTGAGGACATGCTGAAGGGACCCGCTACATTT
GAGCAGGCTAGTAGAGTCCATGTGGAACCTACTAATGCCCAAGCCTCACCGAATAGGGTTGATCAGGGATCTCATAATTTCCATGATCCGTCGCAACACT
ATGTAAAGCATGATGTTCCTGCAAACTCTCATGGCCCCGATTTACAATCTTCACCTCATTCCTATCCAGATCATAGGCCTTCAGGGAACCATAGTGAGGT
CTTATCTGGTCCACAGAATTGGAATATGGAATTCAAGGATGCCACAGCTGCTGCACAGGCAGCTGCTGAATCTGCTGAACGAGCAAGCATGGCTGCCAGA
GCTGCTGCAGAACTTTCAAGTCAAGAAAGGATAACCAGGCAGCATTTAGCAGAATCACGGAAGGCTTTTGCTTTTGAATCAAGAGATGTAGGACCTCAAA
ACTATACTGGCTCAAAGCTGCAAGGTGAAGATGTTGACAAAGATCAAATGAGTAACAACGTGTACCAAAGACATCCAGGGTTACATCGTGAGGAGAGAGA
AGGCAATGAGCAAGATGACCTGGCTGGATTAACTAAAAGGTTTTACAATCTCAAGAGCCCTAATAAACCTAGTCAGTCAGCTTCCTCAAAGTCCAGTAAT
TCTTTTGTGGATGATTATCCATTAATTGATGATCTCCCAATGCCTGATAGGCTTTCTCAAAAGAGGTCTTCCGAATTGGGTGAATCTAGTGTAAAGCTAG
AGTCTAGGGAATCTGAGGTAAGCTTTGTGAGTAAACTGGAAGATGGAATGACATCAGAGAATGTCAGTCACTTTGAGGAAGCAAGAATCAGAAAACAGTC
TAGCAGTGTTTCCTCACATTCGCATTCACAAACTTTCAGTGATGATTACAATGTGTTTTCCAATACCAACCAGCAGAGAATGGGAGATGAGACTGATAAG
GAGCAAAGAGATGCAAAAGGAGCAAATTCTTATGACAATGCCATGGTTTTTGATGATTCTTCTTCAGACAAGGAAATTAAGTTCGATGTGGAGGATGAAC
ATAATGACCAAGTCTATGATTCGGATTTTTCATCAGAAGGTAGAAAGTCATCTTCTCATTTGTTAGCAAATGCAGATGCTTGGGGCCGTACGGAAAACAT
GGATGAGTTTCGAGGAAAGTCCAGTTCGCAGACACCGTTGACCTCGGCATTCTTTTCTCAAGATTTTACCACTGACCCTGTTCCTTCCCAACCACATGAG
ACACCGTTGACCTCGGCATTCTTTTCTCAAGATTTTACCACTGACCCTGTTCCTTCCCAACCACATGACTTGTTACCCATGACTTTTGATGATTCTGACA
GCGTAAGTTCAGAGAGGGAAGTGGATCTGGATACCTATGAGGTGGTTGGTGGTTCAAGTACTGGCATTTTTGCTCATACAAAGAGTGTTAGCACCAGAAA
CTCTGACCCCATTCATAGCGGGAGTCCTCACTCAATAAGATTTTCACTTGCAGACAAGGAAAATTTGGGATCCAACCGTAAAACGCATTTACAGACAGCT
TCACTTGATTCAGATGTCCAAGAAGTATTCTCTATGAAAAATCAGAGAACTGGAGTTGATGTTGAAATGGATAATAAGTTTGCTTATGGTAAATTGGACA
CGAGTCAATCTTCTCCCATACCTGTGAAATCTTGCACAAGTTCTAATGACTTGAAGGACAACCTGCAAACTTCAGGGCATCCAGTAGTGAAAAATGTTCA
AAATTATGAATTACCAATCACCACAAAGAATGCTGATCCTATTGAAGAATCCAATTTGGAAACTGGTACAGAGTTGAACTTGGGAATATTGACAGGTGGC
TTTCGAAACAAGGGTTATAGGCATCCGCCATACCACAGGAATGTATCAAATAATTCCTCATCATCTGAACAAGCAATAGATAATATTCATTCAAGGACTG
GTCGGACCTCTTCTCCTGTCAAGGTTGATATTGGTTCTGGAGCTCGTGATCAGGAGACAAACAATCAGAGACTGCACCCAAAAGTAGATGTAAAAGCAAG
CTCAAGAACCCCAGCTACATATTATGATGACGATTCTGATGAGGAAGTTCCACAACAACATTTTAGCAATAGTCCAAAGTCATATGGCCGAAAATCAGTT
ATTGAAGGGAATGATAGGTCAAGCACAAAGACATCAGATAATCGTGATTCTGAGTTTTCTAAGCCGAATCCATCTAGCAAAACTCATTTAAGCACTAGGT
TTTCTCGACGAACAAAGGCATCTCCTTCTGATAAAGATTATTCTTCGAAGCCTCCAGTGCTTTCCAAGTCTCCAGTGAGTGCAGACTCTTTTGTGGAAAG
AACACCATCCTCAAGTAGCTCCTACACTGCTGACGCTGAATCAATCCCCCAGAGCAGAAGCTCGGGTTACCAGGGAAGTTCTGAGCAATGTAGGTCAACA
GAAGAAGCAGCTTCTAAGCGAATTCAGCAGTCCAAGAGGTTCTCATATGAAGAATCATCCCCAAGGAGTTCAGATGCTATTTCTAGTCAGCAAAAACCAC
CATCTCAGTCAAAGAGTTCAGATTACTGGGCAAGTTCTAGGCAGCCACCTAGATCAGCAGAACAAGCAGCTTCTAAGCAAATTTCAGAATCCAAGAGGTC
TTCGCGTGAGGAAACTCTTAAGTCATCAGCTCGGGAACAGCCATTTAGTTCTCCTCCTAAGCCAGTCGCAACAGACAGTGCTCAAAGTTCAAAGACTTCC
AGTACCCATGGTGAGACGCCATCCAGGGAGAATTCCATTAATAAACCCAGCCATGTTCACCCAAAACTTCCTGACTATGACACCTTTGCAGCGCATCTAC
TGTCTCTCCGCCAGAATCAGCAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G149800.4 pacid=42766997 polypeptide=Potri.006G149800.4.p locus=Potri.006G149800 ID=Potri.006G149800.4.v4.1 annot-version=v4.1
MLHRSFKPAKCKTSLKLASSRIKLLKNKREAQVKHLKRELAQLLDAGQERTARIRVEHVVREEKTMAAYELIEIYCELIVARLPIIESQKNCPIDLKEAV
SSVIFASPRCADVPELMDIRKHLTAKYGKEFVSAAVELRPDCGVSRLLVEKLSSKSPDGPTKIKILTAIAEEHNIKWDPMSFEEKDTKPPEDMLKGPATF
EQASRVHVEPTNAQASPNRVDQGSHNFHDPSQHYVKHDVPANSHGPDLQSSPHSYPDHRPSGNHSEVLSGPQNWNMEFKDATAAAQAAAESAERASMAAR
AAAELSSQERITRQHLAESRKAFAFESRDVGPQNYTGSKLQGEDVDKDQMSNNVYQRHPGLHREEREGNEQDDLAGLTKRFYNLKSPNKPSQSASSKSSN
SFVDDYPLIDDLPMPDRLSQKRSSELGESSVKLESRESEVSFVSKLEDGMTSENVSHFEEARIRKQSSSVSSHSHSQTFSDDYNVFSNTNQQRMGDETDK
EQRDAKGANSYDNAMVFDDSSSDKEIKFDVEDEHNDQVYDSDFSSEGRKSSSHLLANADAWGRTENMDEFRGKSSSQTPLTSAFFSQDFTTDPVPSQPHE
TPLTSAFFSQDFTTDPVPSQPHDLLPMTFDDSDSVSSEREVDLDTYEVVGGSSTGIFAHTKSVSTRNSDPIHSGSPHSIRFSLADKENLGSNRKTHLQTA
SLDSDVQEVFSMKNQRTGVDVEMDNKFAYGKLDTSQSSPIPVKSCTSSNDLKDNLQTSGHPVVKNVQNYELPITTKNADPIEESNLETGTELNLGILTGG
FRNKGYRHPPYHRNVSNNSSSSEQAIDNIHSRTGRTSSPVKVDIGSGARDQETNNQRLHPKVDVKASSRTPATYYDDDSDEEVPQQHFSNSPKSYGRKSV
IEGNDRSSTKTSDNRDSEFSKPNPSSKTHLSTRFSRRTKASPSDKDYSSKPPVLSKSPVSADSFVERTPSSSSSYTADAESIPQSRSSGYQGSSEQCRST
EEAASKRIQQSKRFSYEESSPRSSDAISSQQKPPSQSKSSDYWASSRQPPRSAEQAASKQISESKRSSREETLKSSAREQPFSSPPKPVATDSAQSSKTS
STHGETPSRENSINKPSHVHPKLPDYDTFAAHLLSLRQNQQ
|
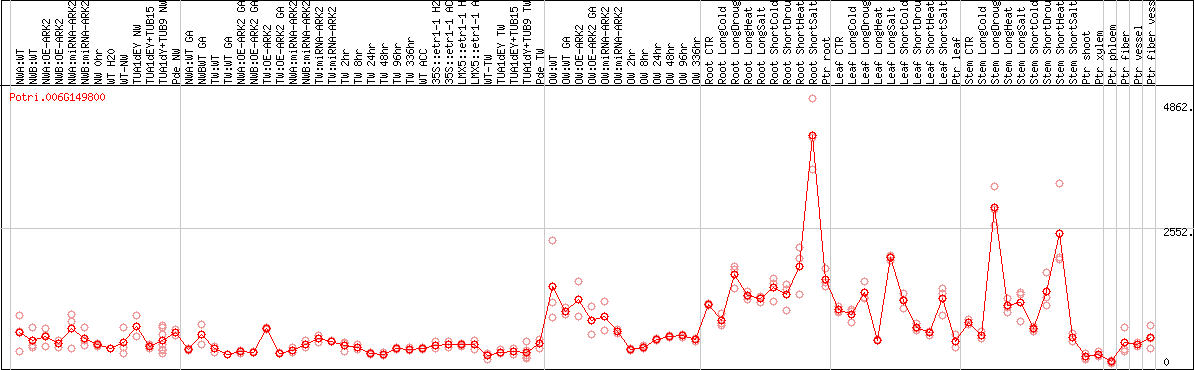
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G149800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
