Potri.006G180100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G180100.1 pacid=42768617 polypeptide=Potri.006G180100.1.p locus=Potri.006G180100 ID=Potri.006G180100.1.v4.1 annot-version=v4.1
ATGGCTCCCAAGACTGGAAAGGCCAAGCCTCATAAAGCTAAAGGAGAGAAAAAGAAGAAGGAAGAAAAAGTTTTGCCTACTGTCATAGAAGTCACCGTTG
AAACGCCGGACGATTCTCAAGTGTCTCTCAAGGGCATCTCCACGGACAGGATCTTGGATGTAAGAAAGCTTTTGGGTGTGCATGTCGAGACATGCCACTT
GACCAATTTCTCCTTATCTCACGAGGTACGGGGACCCAGACTCAAAGATTCAGTGGACATCATCTTGCTCAAACCCTGCCACCTCACCATTACAGAAGAG
GACTACACAGAAGAACAATCCATAGCTCACATCCACCGACTCCTCGACATCGTCGCCTGCACCACCTCATTTGGCGCCTCTTCCACTTCTCCGACAAAAA
CACCGGGTCGAACCGGAGGTTCAAAGGAGTCCGGTTCGACTGAGACGGGCGGAGATAACAAGAAAATCGTGAACAAGAGTGGCAAGGATGCTTGTACGGA
CGCGATGGAGAAAGCGGATGCGGCGGTTTCCATGTGCCCGCCGCCACGGCTTGGCCAGTTTTATGAATTCTTCTCTTTCTCGCACCTCACGCCGCCGGTT
CAATATATCAGGAGATCGAGTCGTCCTTTCCTTGAAGATAAAACGGAAGATGATTTCTTCCAGATAGACGTTAGGGTTTGCAGTGGGAAGCCAATGACAA
TTGTTGCTTCGCGGGAAGGATTCTATCCAGCTGGAAAACGCGCTCTTCTTTGTCGCTCTTTGGTTAGTTTGTTGCAGCAGATTAGCAGAGTTTTTGATTC
TGCATACAAAGCTCTTATGAAAGCTTTCACTGAGCATAATAAATTTGGTAACCTTCCTTATGGTTTTCGAGCGAACACGTGGGTTGTTCCACCGCTTGTT
GCTGATAATCCTTCTGTTTTTCCGCCACTTCCTGTGGAGGATGAGAATTGGGGAGGTAATGGAGGAGGACAAGGAAGAGATGGGAAACATGATTACAGGC
CATGGGCGAAGGAGTTTGCGATTTTGGCAACAATGCCTTGTAAAACAGCAGAAGAGAGGCAAATTCGGGACCGGAAAGCCTTTCTGCTTCATAGTTTATT
TGTCGATGTTTCTGTTTTTAAAGCTGTTGCAGCAATCAAAAGTATCATTGAAAATCAGTGCTTTTTAAGTGATACTGTGAAGTCGTTTTTGCATGAGGAA
AGAGTTGGAGATTTGATTATTATCATTACGAGAGATGTGTCAGATGCTAGCACCAAGTTGGATTGTAAAAATGATGGGTGCCAGGTTCTTGGAGTGTCAC
AGGAAGAACTTGCTCGGAGAAATTTACTAAAGGGAATAACTGCTGATGAAAGTGCAACTGTTCATGATACTCCCACTTTAGGTGTAGTGGTTGTTCGACA
TTGTGGCTTCACAGCTGTTGTTAAAGCTTCTTCTGAGGTGAACTGGGAAGGGGATCCTATTCCCCAGGACATTTCCATTGAGGAACACCCTGAAGGAGGC
GCCAATGCTTTAAATGTTAATAGCTTGAGGATGCTATTGCATAAGTCATCGACGCCCCAGTCATCTAATACACTCCAGCGACTACAAGGTGGAGATCTCG
AAATTTTACACTCTGCCAGGTCTTTGGTGAGGAAAATACTAGAGGATAGTTTGTTGAAGCTACAGGAAGAATCCAGTAGATACACGAAATCTATCAGATG
GGAGCTAGGAGCATGTTGGGTTCAACATTTGCAAAATCAGGCTGCGGGGAAAACTGAGGCTAAGAAAAATGAAGAAACTAATCCTGAGCCTGCTGTCAAG
GGGCTTGGAAAGCAAGGTGCATTGTTAAGGGAGATAAAGAAGAAAACTGATGTCAAAACCGGCAAAACTGAAGAGGGAAAGGATGTTTATGCTGGTAACA
ACCTTGACATGAGCAAGAAACCTGATAGCACTAATCAGGAAGAAATGGAGAAAAAAGATGAGGAAATGAAAGTAATATGGAAAAAGCTGCTTCCTGAAGC
AGCATATCTGCGACTTAGAGAATCAGAAACCGGTCTTCACCTCAAGACACCTGATGAGTTGATTGAAATGGCATATAAATACTATGCTGACACTGCTCTT
CCAAAACTGGTGGCAGATTTTGGCTCCCTGGAACTTTCACCAGTTGATGGAAGAACATTGACAGATTTTATGCATACACGGGGTTTGCAGATGTGTTCTT
TGGGGCGTGTGGTTGAACTTGCAGACAAGCTCCCTCACGTGCAATCACTATGTATACATGAAATGATTGTCCGAGCCTACAAGCACATTTTACAAGCTGT
TGTGGCATCAGTTAATGACGTTGCTGACTTGGCTGCTTGCATTGCATCATGTCTAAATATGTTGTTAGGAACGCCTTCAACTGAAACTGAGGACTCGGAT
ATCATAAATGATGAGAAACTAAAGTGTAAGTGGGTGGAAACATTCGTTGGAAAGAGGTTTGGGTGGCAGTGGAAGCATGAGAGCTACCAGGATCTAAGAA
AGTTTGCCATTCTTCGTGGACTGTCCCACAAGGTTGGCCTTGAGCTTCTTCCCAGGGACTATGATATGGACAATGCTTTTCCTTTCAAGAGGTCTGATAT
TATAAGCATGGTCCCTGTATACAAGCATGTTGCATGTTCATCTGCGGATGGACGTACCCTGCTAGAATCATCCAAAACTTCCTTAGATAAAGGTAAATTG
GAGGATGCTGTAAACTATGGAACTAAGGCACTGTCAAAACTCGTGTCAGTCTGTGGCCCTTACCATCGAATGACTGCGGGAGCATACAGTCTTCTGGCTG
TAGTACTCTACCATACTGGGGATTTTAATCAGGCTACCATTTATCAACAAAAAGCATTGGATATTAATGAGAGGGAGCTTGGACTTGATCATCCTGATAC
AATGAAAAGTTACGGGGATCTAGCTGTCTTCTACTATCGTCTTCAACATACGGAATTGGCCTTGAAGTATGTCAATCGTGCACTGTATCTTTTGCATCTA
ACGTGCGGGCCATCTCATCCAAATACTGCTGCGACGTATATCAATGTAGCAATGATGGAAGAAGGATTAGGGAACGTGCATGTTGCTCTGAGGTACCTTC
ACGAGGCTCTAAAGTGTAACCAGAGACTTCTTGGAGCGGATCATATACAGACTGCTGCCAGCTACCATGCCATAGCAATTGCTCTGTCATTGATGGAAGT
TTACTCCTTAAGTGTTCAGCATGAACAGACTACTCTGCAAATACTGCAGGCCAAACTTGGACCTGAGGATCTACGCACTCAGGATGCAGCTGCATGGCTT
GAATATTTTGAGTCAAAGGCTCTGGAGCAGCAAGAAGCTGCGCGCAACGGTACTCCAAAGCCAGATGCCTCGATCTCCAGTAAAGGTCATTTAAGTGTAT
CAGATCTTTTGGATTATATAACACCAGATGCAGATATGAAAGCAAGAGAAGCACAAAAGAAAGCTCGTGCAAAGGTTAAAGGCAAACCAGGCCAAAATGG
GGAGACAGTCTCAGATGAATACCAAAAGGATGAAATATTGTCACCAACTTATCCTATTGTAGAAAATTCAAGTGACAAGGAAAACAAATCTGAAACTCAA
TTTGCAGAGCCTGGGAATGAAAAATCTGATTCTGGTCTTCCAGATCAGTCATTGTTGAAGACTGATGATAAGACCCAAGAGGAAGACTCAGATGAAGGAT
GGCAAGAAGCTGTTCCTAAAGGTCGTTCACCGACAAGTCGCAAATCTTCTGGTTCAAGGAGGCCAAGCCTAGCCAAATTGAATACCAACTTCATGAATTT
ACCCCAGTCATCAAGATTCCGAGGAAAGCCTAACAATTTTGCATCCCCTAAAACATCACCGAATGATCCTGCTGCTTCCACTGGACTGACAGTTCCTGTT
CCTAAAAAGTTTGCCAAGAGTGCCAGCTTCAGTACAAAAGTAAATAACTCTGGTGCCTCTACTGGTGGAGCAGAGAAATCAAGTACCCCTAAATCAGCCC
CAGCTACCCCTGCCTCAACTGAACAAGTTGCCAAAGCAGCCCCAACAGCAAGTCCAATCAGCGTTCAGTCAGCTGGAAAAATCTTTTCCTATAAAGAAGT
TGCTTTGGCTCCTCCTGGGACAATTGTGAAGGCAGTAGCAGAACAGTTGCCGAAAGGAAATCTTCCAATGGAACCAAGTACTCAGGGTAGCAATGAGGCA
TCTGCAACTGATGTCACATCAGGAGAGGTGACAACATTAAAAGCTGCAGAGGTAGACAACTTTCTGAAACCTGAAGCAGTGAAACATCTTCCGGCTTCAG
AGGGAATGAAAAGTCCTGTCGATCAAAAGAAAGAAACAGAAGAAGGGGGTTTGGTGGCGACAGAGCAACTCGAAGGAAAGAAGTCTGCTGTTGAGGATCG
CACTGATAAAGAAGACAATGGTGCTGAAATTAAAATAGTTGCTGTTAAAGTTAATACCTCAGAAGCTGGAAACATTTCATTCTTGGGTAATGAGAATTTG
GATACTTCCAAGGATTCAAATACCATCTCTTCCCCAACTGAAGTGCCAGAAACACAAGTATCAGACGGTTTCCCAGCAGCTTCCCCTGATATGGAACCTC
AATCTACTTCAACTGAAAATTCTGGTTTGATGGAAAAAGATGCTTCAATCTCAAATGAAGGGGTAGAAGATGAGAATACTCTGGATCCATCTAGTGACAA
TACAAATGCTAAGGCATTGTCGACTGAAGGAGGAAAGCAAGATGAAACAGAAACTGGCAAGGAGACAGCCAAGAAACTTTCTGCAGCTGCACCACCATTT
AACCCATCCATAATAATTCCAGTTTTTGGCTCAGTTACCATTCCAGGATTCAAAGATCATGGAGGACTTCTTCCTTCACCAGTGAATATTCCTCCAATGC
TTACTGTCAATCCTGTTCGCAGATCACCTCATCAGTCAGCAACAGCTAGGGTTCCATATGGTCCGAGGCTGTCTGGTGGTTTTAACAGATCTGGAAATCG
CGTTCCACGTAACAAGCCGAGCTTCAACAATGGTGAGCATACAGGGGATGGGAATCATTTCAGCCCTCCTAGAATAATGAACCCACATGCAGCTGAATTT
GTGCCTGGCCAACCTTGGGTTCCCGATGGCTATTCAATACTGCAAAATGGTTATATGGCTACCACGAATGGTATGCCAGTATCTCCAAATGGCTTCCCCA
TTTCCCCAACCGGTATTCCTGTGTCACCAAATGGTTATCCAGCATTATTGAACGGTATTCAAGCAACTCAAAATGAGTTTCCAGCCTCTCCAGTCAGTTC
AGTAGAAAGACCCATGTTAGTTTCTGTTGATGTTCGTGTTGAAAACAAGAGTGAAGCTGAAGCAGAGAATGGTGTTGAAACTTCTGCAATTGAAGTAGGA
GTTGAAGACCAGTCAGGTGAGAAAGAACATCAAGAAGAAGATGTAAATCCCGAAATAAAAGAAAATCCTGCCGAGCTCCCAGAGACCAGTGATACTGTTG
TAGCAATAGAAACTTGTGATAGCTTGCCAATTGAGGAGAAACCAAGCAAGTGTTGGGCGGATTACAGCGATAATGAAGCTGATATTGTTGAGGTTGCAAG
TTGA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G180100.1 pacid=42768617 polypeptide=Potri.006G180100.1.p locus=Potri.006G180100 ID=Potri.006G180100.1.v4.1 annot-version=v4.1
MAPKTGKAKPHKAKGEKKKKEEKVLPTVIEVTVETPDDSQVSLKGISTDRILDVRKLLGVHVETCHLTNFSLSHEVRGPRLKDSVDIILLKPCHLTITEE
DYTEEQSIAHIHRLLDIVACTTSFGASSTSPTKTPGRTGGSKESGSTETGGDNKKIVNKSGKDACTDAMEKADAAVSMCPPPRLGQFYEFFSFSHLTPPV
QYIRRSSRPFLEDKTEDDFFQIDVRVCSGKPMTIVASREGFYPAGKRALLCRSLVSLLQQISRVFDSAYKALMKAFTEHNKFGNLPYGFRANTWVVPPLV
ADNPSVFPPLPVEDENWGGNGGGQGRDGKHDYRPWAKEFAILATMPCKTAEERQIRDRKAFLLHSLFVDVSVFKAVAAIKSIIENQCFLSDTVKSFLHEE
RVGDLIIIITRDVSDASTKLDCKNDGCQVLGVSQEELARRNLLKGITADESATVHDTPTLGVVVVRHCGFTAVVKASSEVNWEGDPIPQDISIEEHPEGG
ANALNVNSLRMLLHKSSTPQSSNTLQRLQGGDLEILHSARSLVRKILEDSLLKLQEESSRYTKSIRWELGACWVQHLQNQAAGKTEAKKNEETNPEPAVK
GLGKQGALLREIKKKTDVKTGKTEEGKDVYAGNNLDMSKKPDSTNQEEMEKKDEEMKVIWKKLLPEAAYLRLRESETGLHLKTPDELIEMAYKYYADTAL
PKLVADFGSLELSPVDGRTLTDFMHTRGLQMCSLGRVVELADKLPHVQSLCIHEMIVRAYKHILQAVVASVNDVADLAACIASCLNMLLGTPSTETEDSD
IINDEKLKCKWVETFVGKRFGWQWKHESYQDLRKFAILRGLSHKVGLELLPRDYDMDNAFPFKRSDIISMVPVYKHVACSSADGRTLLESSKTSLDKGKL
EDAVNYGTKALSKLVSVCGPYHRMTAGAYSLLAVVLYHTGDFNQATIYQQKALDINERELGLDHPDTMKSYGDLAVFYYRLQHTELALKYVNRALYLLHL
TCGPSHPNTAATYINVAMMEEGLGNVHVALRYLHEALKCNQRLLGADHIQTAASYHAIAIALSLMEVYSLSVQHEQTTLQILQAKLGPEDLRTQDAAAWL
EYFESKALEQQEAARNGTPKPDASISSKGHLSVSDLLDYITPDADMKAREAQKKARAKVKGKPGQNGETVSDEYQKDEILSPTYPIVENSSDKENKSETQ
FAEPGNEKSDSGLPDQSLLKTDDKTQEEDSDEGWQEAVPKGRSPTSRKSSGSRRPSLAKLNTNFMNLPQSSRFRGKPNNFASPKTSPNDPAASTGLTVPV
PKKFAKSASFSTKVNNSGASTGGAEKSSTPKSAPATPASTEQVAKAAPTASPISVQSAGKIFSYKEVALAPPGTIVKAVAEQLPKGNLPMEPSTQGSNEA
SATDVTSGEVTTLKAAEVDNFLKPEAVKHLPASEGMKSPVDQKKETEEGGLVATEQLEGKKSAVEDRTDKEDNGAEIKIVAVKVNTSEAGNISFLGNENL
DTSKDSNTISSPTEVPETQVSDGFPAASPDMEPQSTSTENSGLMEKDASISNEGVEDENTLDPSSDNTNAKALSTEGGKQDETETGKETAKKLSAAAPPF
NPSIIIPVFGSVTIPGFKDHGGLLPSPVNIPPMLTVNPVRRSPHQSATARVPYGPRLSGGFNRSGNRVPRNKPSFNNGEHTGDGNHFSPPRIMNPHAAEF
VPGQPWVPDGYSILQNGYMATTNGMPVSPNGFPISPTGIPVSPNGYPALLNGIQATQNEFPASPVSSVERPMLVSVDVRVENKSEAEAENGVETSAIEVG
VEDQSGEKEHQEEDVNPEIKENPAELPETSDTVVAIETCDSLPIEEKPSKCWADYSDNEADIVEVAS
|
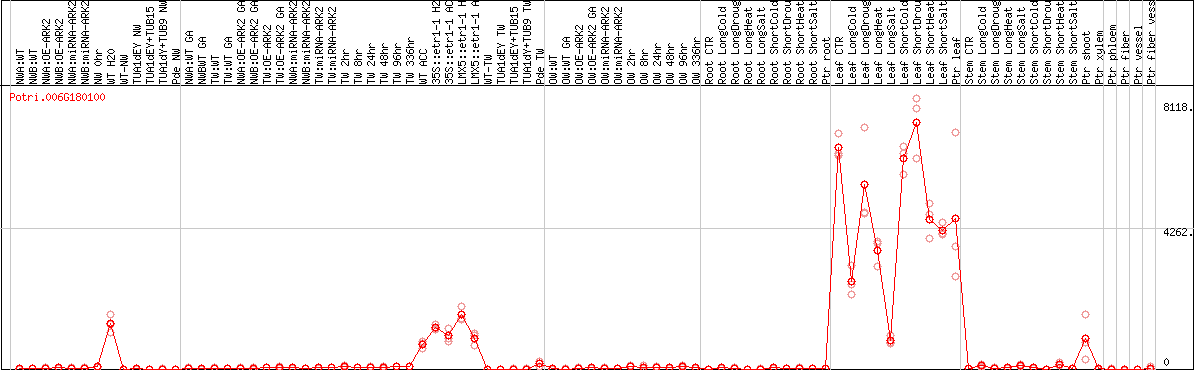
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G180100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
