External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25220 433 / 1e-151
HD
KNAT3
KNOTTED1-like homeobox gene 3 (.1.2)
AT5G11060 425 / 4e-149
HD
KNAT4
KNOTTED1-like homeobox gene 4 (.1)
AT4G32040 389 / 4e-135
HD
KNAT5
KNOTTED1-like homeobox gene 5 (.1)
AT1G62990 332 / 8e-114
HD
IXR11, KNAT7
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
AT1G70510 109 / 2e-27
HD
ATK1, KNAT2
ARABIDOPSIS THALIANA KN 1, KNOTTED-like from Arabidopsis thaliana 2 (.1)
AT1G23380 109 / 3e-27
HD
KNAT6S, KNAT6L, KNAT6
KNOTTED1-like homeobox gene 6 (.1.2)
AT4G08150 107 / 6e-26
HD
BP1, KNAT1
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
AT1G62360 102 / 2e-24
HD
WAM1, WAM, SHL, BUM1, STM
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
AT1G75430 57 / 6e-09
HD
BLH11
BEL1-like homeodomain 11 (.1)
AT5G02030 57 / 6e-09
HD
PNY, BLR, BLH9, RPL, HB-6, VAN, LSN
VAAMANA, REPLUMLESS, PENNYWISE, LARSON, BELLRINGER, BEL1-LIKE HOMEODOMAIN 9, POX (plant homeobox) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G114100
585 / 0
AT5G25220 441 / 6e-155
KNOTTED1-like homeobox gene 3 (.1.2)
Potri.006G259400
446 / 6e-157
AT5G25220 539 / 0.0
KNOTTED1-like homeobox gene 3 (.1.2)
Potri.018G022700
437 / 3e-153
AT5G25220 513 / 0.0
KNOTTED1-like homeobox gene 3 (.1.2)
Potri.001G112200
343 / 3e-118
AT1G62990 466 / 2e-167
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Potri.005G014200
123 / 2e-32
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.013G008600
122 / 4e-32
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.011G011101
118 / 3e-30
AT1G62360 400 / 7e-139
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.006G270401
117 / 5e-30
AT1G62360 399 / 3e-138
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.010G043500
108 / 4e-27
AT1G23380 371 / 3e-129
KNOTTED1-like homeobox gene 6 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006291
473 / 3e-167
AT5G25220 489 / 3e-172
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10001941
387 / 2e-134
AT5G25220 439 / 2e-153
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10001934
387 / 6e-134
AT5G25220 435 / 9e-152
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10003183
361 / 2e-126
AT5G25220 381 / 2e-133
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10024485
290 / 2e-97
AT1G62990 406 / 4e-144
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Lus10017824
198 / 7e-63
AT1G62990 269 / 9e-92
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Lus10001195
140 / 1e-40
AT1G62990 189 / 2e-60
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Lus10029125
110 / 2e-27
AT1G23380 417 / 3e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10013037
108 / 9e-27
AT1G23380 417 / 4e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10030003
90 / 8e-20
AT1G62360 315 / 5e-105
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03789
ELK
ELK domain
PF03790
KNOX1
KNOX1 domain
PF03791
KNOX2
KNOX2 domain
CL0123
HTH
PF05920
Homeobox_KN
Homeobox KN domain
Representative CDS sequence
>Potri.006G190000.1 pacid=42767968 polypeptide=Potri.006G190000.1.p locus=Potri.006G190000 ID=Potri.006G190000.1.v4.1 annot-version=v4.1
ATGGCTTTTCAAGACCACCACCATACACCACAAGAAATGACTTTTCAACTACCACAACACCACCACCTCTCAGCCTCACCTTCCACCGGTCCCACATGGC
TAAGCAACGCAGTCCTCCGTAGACACGACGACGTTTTAACCCAAACCCGCATCGAAAAACCAGAAAACAACACCAATAATGGAAGTGAAGAGGAATTGAT
AGATTCCGTTAGTGATAATTGGGAGAGAGCTAAATGCAAAGCTGAGATATTAGGGCATCCCTTATATGAACAGTTATTGGCTGCTCACGTGGCTTGTTTA
AGGATTGCCACGCCAGTTGACCAGTTGGCTAGGATTGATACACAGTTGGCTCAGTCTCAAGATGTTGTGGCTAAGTATTCTGGTGTTGGTAGGAGCCATG
TTGTTGATGAGAAAGAACTTGATCAGTTCATGACACATTACGTTATATTGCTCTGTTCCTTCAAAGACCAATTGCAACAACATGTCCGTGTTCATGCTAT
GGAGGCTGTAATGGCATGCTGGGAACTTGAACAGTCTCTACAAAGTTTGACAGGTGTATCTCCAGGGGAAGGCACTGGTGCAACTATGTCTGATGATGAT
GACGACCAAGCAGATAGTGATGCCAACTTGTATGATGGAAATCTTGATGGGTTGGATACCATGGGATTTGGTCCTCTAGTTCCCACCGAGACTGAGAGGT
CCTTGATGGAGCGTGTAAGGCAAGAATTGAAGCATGAGCTCAAACAGGATTACAAGGAGAAGATTGTGGATATTAGAGAGGAAATTCTACGTAAGAGAAG
AGCTGGAAAACTGCCAGGTGATACTACGTCCCTCTTAAAAGCTTGGTGGCAAACACATTCCAAGTGGCCATACCCAACTGAGGAAGACAAAGCAAGATTG
GTGCAGGAAACAGGTTTGCAATTAAAGCAGATAAATAATTGGTTCATAAACCAAAGGAAAAGGAACTGGCACAGCAGTCCTTCAGGTTCTACTTCGAAGA
GCAAACGAAAGAAGTAA
AA sequence
>Potri.006G190000.1 pacid=42767968 polypeptide=Potri.006G190000.1.p locus=Potri.006G190000 ID=Potri.006G190000.1.v4.1 annot-version=v4.1
MAFQDHHHTPQEMTFQLPQHHHLSASPSTGPTWLSNAVLRRHDDVLTQTRIEKPENNTNNGSEEELIDSVSDNWERAKCKAEILGHPLYEQLLAAHVACL
RIATPVDQLARIDTQLAQSQDVVAKYSGVGRSHVVDEKELDQFMTHYVILLCSFKDQLQQHVRVHAMEAVMACWELEQSLQSLTGVSPGEGTGATMSDDD
DDQADSDANLYDGNLDGLDTMGFGPLVPTETERSLMERVRQELKHELKQDYKEKIVDIREEILRKRRAGKLPGDTTSLLKAWWQTHSKWPYPTEEDKARL
VQETGLQLKQINNWFINQRKRNWHSSPSGSTSKSKRKK
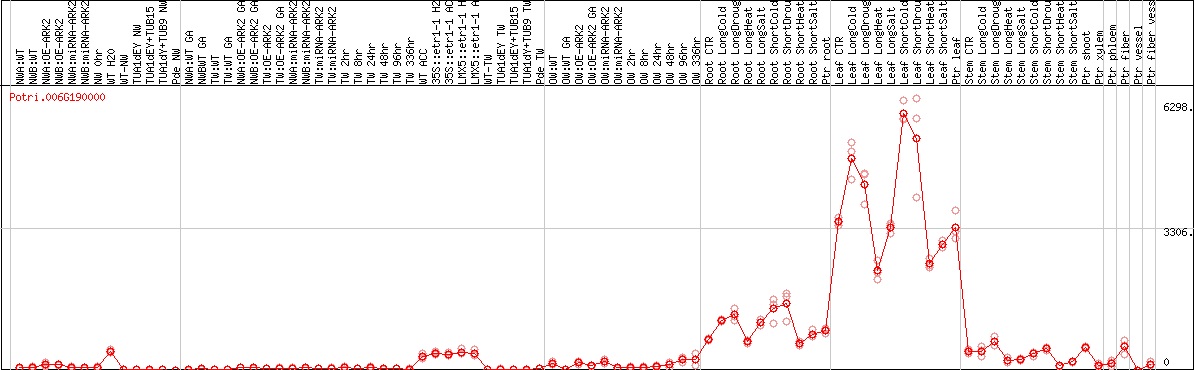
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G190000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.