External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G15010 180 / 3e-53
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT3G56860 178 / 9e-52
UBA2A
UBP1-associated protein 2A (.1.2.3.4.5)
AT2G41060 174 / 2e-50
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G22090 112 / 3e-28
UBA1A
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G19380 100 / 4e-23
RNA recognition motif (RRM)-containing protein (.1)
AT1G17640 98 / 8e-23
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G24770 96 / 3e-22
CP31, ATRBP33, ATRBP31, RBP31
ARABIDOPSIS THALIANA RNA BINDING PROTEIN, APPROXIMATELY 31 KD, 31-kDa RNA binding protein (.1)
AT5G47620 94 / 3e-21
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT1G76460 92 / 3e-21
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G22330 90 / 2e-20
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G048100
499 / 7e-179
AT3G15010 181 / 1e-53
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.004G055400
197 / 4e-60
AT3G15010 303 / 2e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.011G065200
196 / 1e-59
AT3G15010 295 / 1e-97
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.001G377400
187 / 2e-55
AT3G15010 342 / 2e-114
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.001G220600
177 / 1e-51
AT3G56860 232 / 1e-70
UBP1-associated protein 2A (.1.2.3.4.5)
Potri.006G025600
176 / 3e-51
AT3G56860 353 / 2e-117
UBP1-associated protein 2A (.1.2.3.4.5)
Potri.005G081600
174 / 5e-51
AT2G41060 213 / 8e-65
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.016G023800
168 / 6e-48
AT3G56860 348 / 1e-115
UBP1-associated protein 2A (.1.2.3.4.5)
Potri.016G090700
97 / 3e-23
AT2G37220 302 / 4e-103
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023817
420 / 2e-147
AT3G15010 188 / 5e-56
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10021035
402 / 1e-136
AT3G15010 182 / 4e-51
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10025418
184 / 1e-54
AT3G15010 336 / 8e-113
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10032516
183 / 3e-54
AT3G56860 405 / 6e-139
UBP1-associated protein 2A (.1.2.3.4.5)
Lus10015290
179 / 9e-53
AT3G15010 335 / 2e-112
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10043016
180 / 2e-52
AT3G56860 463 / 1e-160
UBP1-associated protein 2A (.1.2.3.4.5)
Lus10039054
99 / 1e-22
AT3G07810 428 / 4e-147
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10023723
95 / 6e-22
AT1G60000 259 / 1e-85
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10043168
97 / 7e-22
AT3G07810 456 / 2e-157
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10032579
96 / 1e-21
AT3G07810 447 / 8e-154
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.006G190900.2 pacid=42767375 polypeptide=Potri.006G190900.2.p locus=Potri.006G190900 ID=Potri.006G190900.2.v4.1 annot-version=v4.1
ATGGAAGACTTGAAGAAGAGAAAGATGGACGAGGCAATCATCAATGGTTCTGCCGAAACATTAACCACACAAGATTATCTTCGCTCACTGCTTGACCCTC
TTAGTAAACCCCAGCTTGTCGATCTTCTTTCTAGACTAGGGTCTCAATATCCTTCAATTGCAGAAGAAATTAAAAGTCTTGCCAGTGCGGATCCAGTCCA
TCGAAAGCTTTTTGTTCGTGGCTTGGCCTGGAATACCACTTCAGAAACCTTGTGTGCTGAATTTCGAATGCATGGTGAGATAGAAGAAGGGTCAGTGATC
TATGACAAAGCGACAGGAAAGTCACGTGGCTATGGTTTCATTACGTACAAGCATATGGAATCGGCACAAAGTGCACTAGGTGCACCTAGCAAATTGATTG
ATGGCCGAATGGCTGTCTGCAATTTAGCTTGTGAGGGATTAACTGGTGCAACTACCACACCGGATCTAACCCAGAGGAAACTCTACATTGGGGGTTTGTC
GCCAGAGATCTCAAGTGAGATGCTGCTTCATTTTTTTGGAAGATATGGTGAGATTGAAGAAGGCTCAGTTGCCTATAACAAAGATACTAATGAATCACGT
GGGTTTGGTTTTGTTACATACAAGACAGTGGAGGCTGCAAAGAAGGCCATAGATGATCCACACAAGTTGCTTGGGGGGAGAACTATCACTGTGAAGCTTG
CTGATACCCACAAGGGCAAGACGGTACAAATGCAGTCACCAGCGCCCATGGTTCCAGTACCTGTACCAATAGCAGCAGCTGGTTATGCGCAGCCAGGTAA
AGCACCAGTTGGCAGTGGTACTCCTGTTGGTTATCCTTATCATCAAACTGTAGCATCATATCCAGCTTCTTCCTACCCTAATCCTCCAGTTGCACCTGCT
CCATATCCAACACAATCTCAAGTTTCATATGCACCAGTTTCTGCAAAGAAAGAACTGTTAGGACTTTCATCAACACCACCTGTCGGAATGGGTGGATACC
CTTATTACTACCCTAAACAGTAA
AA sequence
>Potri.006G190900.2 pacid=42767375 polypeptide=Potri.006G190900.2.p locus=Potri.006G190900 ID=Potri.006G190900.2.v4.1 annot-version=v4.1
MEDLKKRKMDEAIINGSAETLTTQDYLRSLLDPLSKPQLVDLLSRLGSQYPSIAEEIKSLASADPVHRKLFVRGLAWNTTSETLCAEFRMHGEIEEGSVI
YDKATGKSRGYGFITYKHMESAQSALGAPSKLIDGRMAVCNLACEGLTGATTTPDLTQRKLYIGGLSPEISSEMLLHFFGRYGEIEEGSVAYNKDTNESR
GFGFVTYKTVEAAKKAIDDPHKLLGGRTITVKLADTHKGKTVQMQSPAPMVPVPVPIAAAGYAQPGKAPVGSGTPVGYPYHQTVASYPASSYPNPPVAPA
PYPTQSQVSYAPVSAKKELLGLSSTPPVGMGGYPYYYPKQ
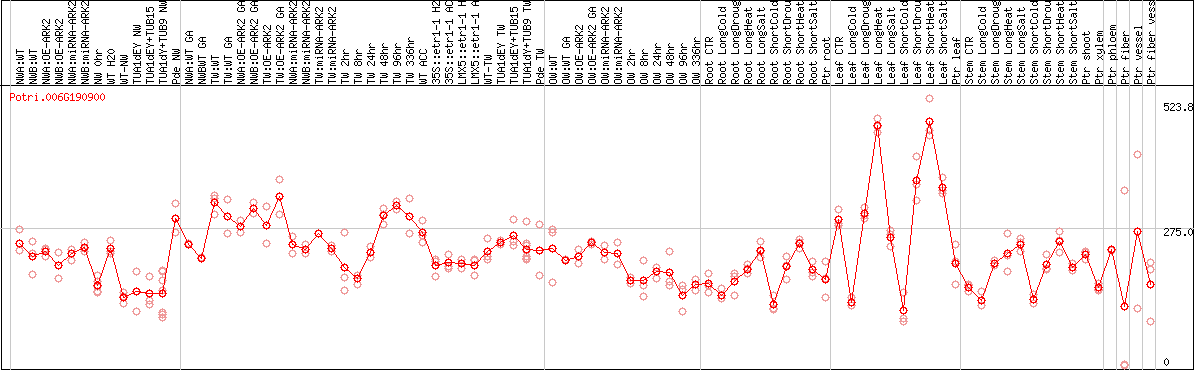
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G190900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.