Potri.006G194900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G194900.6 pacid=42769903 polypeptide=Potri.006G194900.6.p locus=Potri.006G194900 ID=Potri.006G194900.6.v4.1 annot-version=v4.1
ATGGCGACGAAACAGGGATCAAAATCAAGAATATCAGGTTTAATAAGTAATTCAAAGAAACCAGCAGCGAATTCACAGTCATCATCAACGGCATCTTCTA
CTAAGCAATTTTTAGAGAATTCAATGGACGGTCAAAGCTCACCTGCATCCTCGTCAGCTCGAAGCAAACCACAATATTTTTACTCCGAAAGTGTTAATCT
AGATACTGAGAGATCCAAAGAAAATGTAACTGTCACTGTTCGCTTTCGGCCCCTAAGTCCTAGGGAAATCAGGCAAGGAGAGGAGATTGCGTGGTATGCG
GATGGAGAAACTGTCGTGCGAAATGAACATAATCCATCTACTGCGTATGCGTATGATCGGGTGTTTGGTCCTACAACTACAACGCGCCATGTATATGATG
TTGCTGCTCAGCATGTTGTCAATGGTGCTATGGAAGGGATCAATGGCACCATTTTCGCATATGGAGTGACTAGCAGCGGGAAAACCCATACCATGCATGG
GGATCAAAGGTCACCTGGAATTATACCATTAGCAGTGAAGGATGCTTTTAGTATCATTCAAGAGACTCCAAACCGTGAGTTCCTCCTCCGTGTTTCATAC
CTGGAGATCTATAATGAGGTTGTTAACGACTTGTTGAATCCAGCAGGACAAAATTTAAGAATCAGAGAAGATGCTCAGGGAACTTTTGTCGAGGGAATAA
AAGAAGAAGTTGTATTATCCCCTGCTCATGCACTTTCCCTTATAGCAGCTGGGGAAGAGCACAGACATGTTGGGTCCACAAACTTTAATTTACTCAGCAG
CAGGAGTCATACAATATTTACACTGACAGTAGAGAGTAGCCTGTATGGTGAAAATAGTGAAGGAGAGGCTGTAAATTTGTCCCAGCTGAGCCTCATCGAT
CTGGCAGGTTCTGAAAGCTCAAAGGCTGAAACTACTGGCGTGAGACGGAAAGAAGGATCCTATATTAATAAAAGTTTGCTGACACTTGGAACTGTTATAT
CGAAGTTGACGGATGGGAGAGCCGCTCATATTCCTTACAGGGACTCTAAATTGACAAGGCTACTTCAGTCTTCATTAAGTGGTCATGGGCGTGTATCACT
CATTTGCACTGTCACTCCCTCATCTAGTAGTTCGGAAGAGACACACAATACTTTAAAATTTGCCCACCGTGCAAAGCACATTGAAATTCAAGCAGCACAG
AACAAGATTATTGATGAGAAATCGCTTATCAAGAAGTACCAAAATGAGATACGCTCTTTAAAGGAAGAGTTAGAACAATTGAAAAGGGGTATTGTGACAA
TTCCTCGATTGAAAGACATTGTAGAAGACGACATTGTCCTCCTCAAGCAGAAGCTAGAAGATGGTCAAGTGAAACTGCAATCAAGATTGGAACAAGAAGA
AGAAGCTAAAGCAGCTCTATTGAGCAGAATACAGCGGCTGACAAAATTAATTTTGGTGTCAACAAAAGCTTCACAACCATCAAGGATCTCTCACCGCCCA
GGTCCAAGGAGGAGGCATTCCTTTGGAGAAGAAGAGCTTGCATACCTACCGTACAAGAGGCAGGACTTGATATTGGATGACGAAAATATTGACTTGTATG
TTTCTCTTGAAGGAAATACTGAAAGTGCGGATGAGACACTGAAAGAGGAGAAGAAAACCCGAAAGCATGGATTGCTGAACTGGTTAAAGCTACGGAAACG
AGATAGTGGATTGGGAATGAGCACCAGTGACAAATCAAGTGGAGTTAAATCTAATAGTACACCTTCAACACCTCAAGCAGAAAACAGCAATTATTATGCA
GAATCCAGACTTTCACATCCTTCACTTGCAGAAAGCTCTCCATCAGCTGATCTTCTATCTGAGGTCAGGCAGGATAGAGAGGTACCTGAGGACAATTTCC
TTGAGCAGGAAACTCCTTTGAATGGCATAAAAACAAGTGATCAGATTGATCTTCTGAGGGAGCAGCAGAAAATTTTATCTGGAGAGGTGGCACTCCATTC
AAGTATTTTGAAGCGATTGTCTGAGGAGGCTTCAAGGAATCCCCTGAAGGAACACATACAGTTGGAGATGAAGAAGTTGAGTGATGAAATCAAGGTGAAG
AATGAACAAATAGCTTTGTTGGAAAAGCAAATTGCTGATTCCATCATGGCCTCTCACAACAGCTTGGCTAACTTGGAAGCATCGCAAACGATTGCTGAAC
TGACAGCTCAATTGAATGAGAAGTCATTTGAACTCGAGGTTAAAGCGGCAGATAATTGTATAATTCAAGACCAGCTCAGCCAAAAGATCTGTGAATGTGA
AGGATTGCAGGAAACAATTGTCTCTTTGAAGCAGCAGCTCTCAGATGCACTGGAGTCAAAAAATATTAGCCCTCTAGCTAGCTACTCTCAACGAATTTCT
GAACTAAAAAGCTTTCATGCACAACATCACATGAACAAGGAAACTGCAGCATCAAAAGATAGAAATGAAGATTTGCTTCTACAAGCACAGGCCACTGAGA
TGGAAGAACTGAAGCAAAAAGTTGATGCACTAACAGAATCAAAAGAGCAGTTAGAAACTCGGAACCAAAAATTGGCCGAGGAGAGTTCATACGCCAAAGG
GCTAGCATCAGCTGCTGCTGTTGAGCTCAAGGCATTATCAGAAGAAGTGGCGAAGCTTATGAATCATAATGAGAGATTAACTGCCGAGCTGATAGCATTG
AAGAACTCCCCCACTCAGCGTAGAAGTGGCAGTACTGTTCGAAATGGCCGGAGAGACAATCACATGAAACACCAAGACCAAGTTGGGGCAGCCTCAGAGC
TCAAGAGAGAGTTGGCCGTGAGTCGAGAAAGAGAAGTTCAATATGAAGCTGCTCTTATGGAGAAGGATCAAAGAGAGACTGATCTTCAAAGGAAAGTTAA
GGAATCCAAACAAAGAGAAGCTTATCTGGAAAATGAACTTGCTAACATGTGGGTTCTCGTTGCGAAGCTGAAGAAATCCCAAGGAGCTGAAATGGATGTC
TCCGAGGCAACAGGACATGACGGTTTAGGAATCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G194900.6 pacid=42769903 polypeptide=Potri.006G194900.6.p locus=Potri.006G194900 ID=Potri.006G194900.6.v4.1 annot-version=v4.1
MATKQGSKSRISGLISNSKKPAANSQSSSTASSTKQFLENSMDGQSSPASSSARSKPQYFYSESVNLDTERSKENVTVTVRFRPLSPREIRQGEEIAWYA
DGETVVRNEHNPSTAYAYDRVFGPTTTTRHVYDVAAQHVVNGAMEGINGTIFAYGVTSSGKTHTMHGDQRSPGIIPLAVKDAFSIIQETPNREFLLRVSY
LEIYNEVVNDLLNPAGQNLRIREDAQGTFVEGIKEEVVLSPAHALSLIAAGEEHRHVGSTNFNLLSSRSHTIFTLTVESSLYGENSEGEAVNLSQLSLID
LAGSESSKAETTGVRRKEGSYINKSLLTLGTVISKLTDGRAAHIPYRDSKLTRLLQSSLSGHGRVSLICTVTPSSSSSEETHNTLKFAHRAKHIEIQAAQ
NKIIDEKSLIKKYQNEIRSLKEELEQLKRGIVTIPRLKDIVEDDIVLLKQKLEDGQVKLQSRLEQEEEAKAALLSRIQRLTKLILVSTKASQPSRISHRP
GPRRRHSFGEEELAYLPYKRQDLILDDENIDLYVSLEGNTESADETLKEEKKTRKHGLLNWLKLRKRDSGLGMSTSDKSSGVKSNSTPSTPQAENSNYYA
ESRLSHPSLAESSPSADLLSEVRQDREVPEDNFLEQETPLNGIKTSDQIDLLREQQKILSGEVALHSSILKRLSEEASRNPLKEHIQLEMKKLSDEIKVK
NEQIALLEKQIADSIMASHNSLANLEASQTIAELTAQLNEKSFELEVKAADNCIIQDQLSQKICECEGLQETIVSLKQQLSDALESKNISPLASYSQRIS
ELKSFHAQHHMNKETAASKDRNEDLLLQAQATEMEELKQKVDALTESKEQLETRNQKLAEESSYAKGLASAAAVELKALSEEVAKLMNHNERLTAELIAL
KNSPTQRRSGSTVRNGRRDNHMKHQDQVGAASELKRELAVSREREVQYEAALMEKDQRETDLQRKVKESKQREAYLENELANMWVLVAKLKKSQGAEMDV
SEATGHDGLGI
|
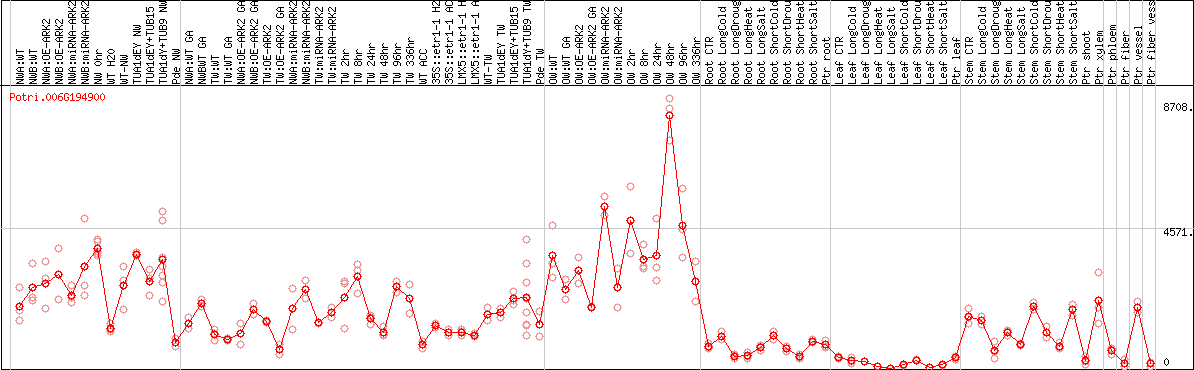
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G194900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
