External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06210 160 / 1e-51
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G27330 90 / 3e-24
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G73530 91 / 6e-24
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G23830 90 / 7e-24
AtGRP4, GR-RBP4, GRP4
glycine-rich RNA-binding protein 4 (.1.2)
AT3G46020 89 / 7e-24
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G37510 86 / 2e-22
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G13860 85 / 2e-22
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G13850 86 / 4e-22
ATGRP2, GR-RBP2
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
AT1G74230 89 / 6e-22
GR-RBP5
glycine-rich RNA-binding protein 5 (.1)
AT4G39260 83 / 1e-21
ATGRP8, CCR1, GR-RBP8
glycine-rich RNA-binding protein 8, GLYCINE-RICH PROTEIN 8, cold, circadian rhythm, and RNA binding 1 (.1.2.3.4)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029426
143 / 6e-41
AT5G27720 190 / 2e-58
SM-like protein 4, embryo defective 1644, Small nuclear ribonucleoprotein family protein (.1)
Lus10004224
101 / 7e-29
AT5G06210 96 / 1e-26
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10013306
99 / 3e-27
AT5G06210 94 / 3e-25
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10023758
95 / 1e-25
AT2G37510 164 / 9e-53
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10003980
92 / 3e-24
AT2G37510 169 / 1e-54
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10034685
92 / 2e-23
AT5G61030 177 / 7e-54
glycine-rich RNA-binding protein 3 (.1)
Lus10021154
92 / 3e-23
AT5G61030 188 / 5e-58
glycine-rich RNA-binding protein 3 (.1)
Lus10017852
91 / 4e-23
AT5G61030 178 / 7e-55
glycine-rich RNA-binding protein 3 (.1)
Lus10040519
92 / 2e-22
AT5G61030 189 / 2e-53
glycine-rich RNA-binding protein 3 (.1)
Lus10015146
85 / 6e-22
AT3G08000 145 / 5e-46
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.006G208500.2 pacid=42769591 polypeptide=Potri.006G208500.2.p locus=Potri.006G208500 ID=Potri.006G208500.2.v4.1 annot-version=v4.1
ATGGCTGCGGCACTTAGAAGAATCGCAGGAGAGAAAAGCTTTATGCTTTCAAGTATTCGTGCTAAACTTGTTGCTGCTCCAACTGCCCTTTTCATTTGGC
AGCGAGGCATAACCACTAAACTCTTTGTTGGAGGCTTGTCAATTTACACCACAGAGAATGGATTATCAGAAGCATTCTCTCAATATGGTCAGGTTGTTGA
AGCTAAAATTGTGATGGACCGAGCCTTGGACAGATCTAAAGGTTTTGGTTTTGTAACTTATGCATCTGAAGATGAAGCTCAGAAAGCTCTGGATGAGATG
AATGGGAAGGCACTGAATGGACGAGTCATCTATGTGGATTATGCAAAGCTCAAAACTAATTTTGGTGGTGGGATACCAATTGCTAGAGGACCTCCTGAAC
CAACAACATCTGAGAGTTGA
AA sequence
>Potri.006G208500.2 pacid=42769591 polypeptide=Potri.006G208500.2.p locus=Potri.006G208500 ID=Potri.006G208500.2.v4.1 annot-version=v4.1
MAAALRRIAGEKSFMLSSIRAKLVAAPTALFIWQRGITTKLFVGGLSIYTTENGLSEAFSQYGQVVEAKIVMDRALDRSKGFGFVTYASEDEAQKALDEM
NGKALNGRVIYVDYAKLKTNFGGGIPIARGPPEPTTSES
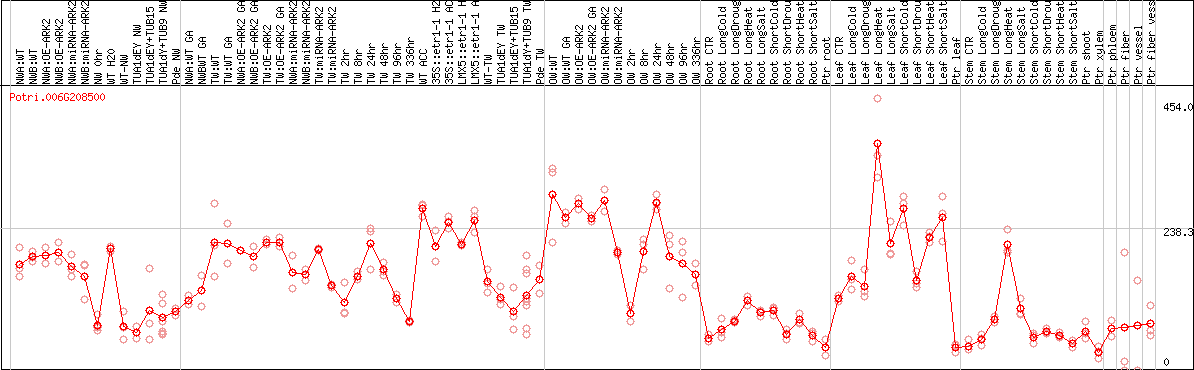
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G208500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.