gdcP1,Pt-GDCP.2 (Potri.006G229300) [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | gdcP1,Pt-GDCP.2 | ||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G229300.1 pacid=42769919 polypeptide=Potri.006G229300.1.p locus=Potri.006G229300 ID=Potri.006G229300.1.v4.1 annot-version=v4.1
ATGGAACGTGCAAGGAGGCTAGCTAACCGTGCAATCTTGAAAAGGCTGGTTAATGAATCAAAGCAAAGTCACAAGCAGGCGAGAAATGATTCATCTTTGT
TAAACTCTTCATCTCCTGTATCATACACACCTTCAAGGTATGTTTCCTCATTGTCATCATTTGGTTCTAGGAGTCCTAGATCAGGTCTATTGCCAGGAAC
CAAAAATATTGTATCCCATAATGTTCCTGCTGGTTCTTATGGCATTGGGTCACAGATAAGATCCATTTCTGTTGAATCTTTGAAACCAAGTGACACTTTC
CCTCGCCGCCATAACTCAGCCACCCCAGAAGAACAAACCAAAATGGCTGAGTTATGTGGCTTTGATACTCTTGATTCATTGATAGATGCAACTGTGCCTA
AATCTATCAGACTTGATTCTATGAAGTTCTCAAAGTTTGATGGAGGGTTAACCGAGAGCCAAATGATTGAACATATGAATTATCTGGCATCAAAGAATAA
GGTTTTTAAGTCCTATATTGGGATGGGATATTATAACACTCATGTTCCACCTGTGATTTTGAGGAATATAATGGAGAATCCGGCTTGGTATACCCAGTAC
ACACCGTACCAAGCTGAGATATCTCAGGGTAGATTGGAGTCTTTGCTCAATTACCAAACCATGATCACGGATCTTACTGGATTGCCTATGTCAAATGCTT
CATTGCTTGATGAAGGGACTGCTGCTGCAGAGGCAATGGCCATGTGTAATAACATTCAAAAGGGAAAGAAAAAGACTTTTATTATTGCTAATAATTGTCA
CCCTCAAACTATTGATATTTGTGATACTAGAGCCGGAGGTTTTGATCTTAAGGTAGTTACCGCAGATCTTAAGGACATTGATTATAAATCTGGGGATGTT
TGTGGGGTCTTAGTTCAGTATCCCGGGACTGAAGGTGAAGTTTTGGATTATGGAGAGTTTATAAAGAATGCTCATGCTCATGGAGTTAAGGTTGTTATGG
CATCGGATTTGTTGGCATTGACGATGTTGAAGCCTCCTGGTGAACTGGGTGCGGATATTGTTGTTGGTTCAGCTCAGAGGTTTGGTGTTCCAATGGGGTA
TGGTGGTCCTCATGCTGCGTTTTTGGCAACCTCACAAGAGTACAAGAGGATGATGCCTGGTAGAATTATTGGTGTTAGTGTTGATTCTTCAGGAAAGCCT
GCTCTGCGCATGGCGATGCAAACTAGGGAGCAGCATATTCGCAGGGACAAGGCTACCAGCAACATCTGTACTGCTCAAGCATTGCTTGCAAACATGGCTG
CTATGTATGCTGTCTATCATGGACCTGAAGGCCTTAAAACCATCGCCCAAAGGGTCCATGGTCTTGCTGGAGCATTCACAGTTGGACTGAAGAAACTTGG
GACAGTTGAAGTCCAGGGCCTTCCCTTCTTTGACACTGTGAAGGTTAAGTGTGCTGATGCACATGCAATTGCTGATGCTGCTTACAAAAGCGAGATAAAT
TTGAGAGTTGTAGATGCAAAAACCATTACTGTTTCTTTTGATGAAACAACCACCTTAGAGGATGTCGATAAACTTTTCAAAGTTTTTTCTGGTGGAAAGC
CTGTCCCTTTCACAGCTGCATCTCTGGCACCAGAGGTTCAGAATGTGATCCCATCTGGTCTAACAAGGGAGAGCCCATATCTGACCCATCCAATCTTTAA
CACGTACCATACCGAGCATGAGCTGCTCAGATACATGCATAGGCTGCAATCAAAAGATCTTTCTTTGTGTCACAGTATGATTCCACTGGGGTCTTGTACA
ATGAAATTGAATGCTACAAGTGAAATGATGCCAGTGACATTGCCTAATTTCACTGATATGCACCCTTTTGCTCCTACTGAACAGTCTCAAGGTTACCAGG
AAATGTTCGATGATTTGGGTGATCTATTGTGTACGATTACTGGGTTTGACTCTTTCTCATTCCAACCAAATGCTGGTGCTGCTGGAGAGTATGCTGGGCT
TATGGTTATTCGTGCATATCATAAGGCAAGAGGAGATCATCATCGAAATGTGTGCATCATACCTGTTTCAGCACATGGAACGAATCCTGCGAGTGCTGCC
ATGTGTGGAATGAAGATTGTTGCTGTTGGAACTGATGCCAAGGGAAACATCAATGTTGAAGAGCTAAGGAAGGCTGCTGAAGACAATAGGGACAACCTAT
CAGCACTTATGGTTACATATCCCTCAACTCACGGGGTCTATGAAGAAGGAATCGATGAGATATGTAAGATAATTCATGACAATGGTGGTCAAGTATACAT
GGATGGGGCTAACATGAATGCTCAGGTTGGTCTCACAAGCCCGGGTTTTATTGGAGCTGATGTTTGCCATCTCAATCTCCACAAAACATTTTGCATTCCG
CATGGAGGTGGTGGTCCTGGCATGGGTCCTATTGGTGTGCAGAAGCACTTGGCACCATATCTTCCTTCACATCCTGTGGTGCCTACAGGTGGTATTCCAG
CCCCTGACCAGTCTCAACCTCTTGGCACCATTTCAGCTGCGCCTTGGGGCTCTGCACTTATCTTGCCAATATCATACACGTACATAGCGATGATGGGTTC
GAAGGGGTTAACTGATGCTTCAAAGATAGCTATACTGAATGCAAACTACATGGCAAAGCGATTGGAGAATTACTATCCCATTCTTTTCCGTGGTGTCAAC
GGAACAGTTGCCCATGAATTCATTGTGGACTTGAGAGGCGTTAAGAACACCGCTGGAATTGAGCCTGAAGATGTTGCCAAGCGTCTTATGGACTATGGAT
TTCATGCACCTACAATGTCATGGCCAGTTCCTGGAACACTTATGATTGAACCCACTGAGAGTGAAAGCAAGGCAGAGTTAGACAGGTTCTGTGATGCGCT
TATCTCCATTAGGGAAGAAATTGCAGAGATTGAGAAAGGCAAAGCTGATATCCACAACAATGTCCTCAAGGGAGCTCCTCATCCACCATCTTTGCTTATG
GGAGATGCATGGACAAAACCATATTCTCGGGAATATGCAGCCTTCCCTGCTTCTTGGCTTCGTGTTGCCAAGTTCTGGCCTAGTACAGGTCGTGTCGACA
ATGTCTATGGTGACCGCAACCTCACCTGCACCCTTCTCTCAGTATCACAGGTTGTTGAAGAACAGGCTGCAGCTACTGCTTAA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.006G229300.1 pacid=42769919 polypeptide=Potri.006G229300.1.p locus=Potri.006G229300 ID=Potri.006G229300.1.v4.1 annot-version=v4.1
MERARRLANRAILKRLVNESKQSHKQARNDSSLLNSSSPVSYTPSRYVSSLSSFGSRSPRSGLLPGTKNIVSHNVPAGSYGIGSQIRSISVESLKPSDTF
PRRHNSATPEEQTKMAELCGFDTLDSLIDATVPKSIRLDSMKFSKFDGGLTESQMIEHMNYLASKNKVFKSYIGMGYYNTHVPPVILRNIMENPAWYTQY
TPYQAEISQGRLESLLNYQTMITDLTGLPMSNASLLDEGTAAAEAMAMCNNIQKGKKKTFIIANNCHPQTIDICDTRAGGFDLKVVTADLKDIDYKSGDV
CGVLVQYPGTEGEVLDYGEFIKNAHAHGVKVVMASDLLALTMLKPPGELGADIVVGSAQRFGVPMGYGGPHAAFLATSQEYKRMMPGRIIGVSVDSSGKP
ALRMAMQTREQHIRRDKATSNICTAQALLANMAAMYAVYHGPEGLKTIAQRVHGLAGAFTVGLKKLGTVEVQGLPFFDTVKVKCADAHAIADAAYKSEIN
LRVVDAKTITVSFDETTTLEDVDKLFKVFSGGKPVPFTAASLAPEVQNVIPSGLTRESPYLTHPIFNTYHTEHELLRYMHRLQSKDLSLCHSMIPLGSCT
MKLNATSEMMPVTLPNFTDMHPFAPTEQSQGYQEMFDDLGDLLCTITGFDSFSFQPNAGAAGEYAGLMVIRAYHKARGDHHRNVCIIPVSAHGTNPASAA
MCGMKIVAVGTDAKGNINVEELRKAAEDNRDNLSALMVTYPSTHGVYEEGIDEICKIIHDNGGQVYMDGANMNAQVGLTSPGFIGADVCHLNLHKTFCIP
HGGGGPGMGPIGVQKHLAPYLPSHPVVPTGGIPAPDQSQPLGTISAAPWGSALILPISYTYIAMMGSKGLTDASKIAILNANYMAKRLENYYPILFRGVN
GTVAHEFIVDLRGVKNTAGIEPEDVAKRLMDYGFHAPTMSWPVPGTLMIEPTESESKAELDRFCDALISIREEIAEIEKGKADIHNNVLKGAPHPPSLLM
GDAWTKPYSREYAAFPASWLRVAKFWPSTGRVDNVYGDRNLTCTLLSVSQVVEEQAAATA
|
DESeq2's median of ratios [POPLAR]
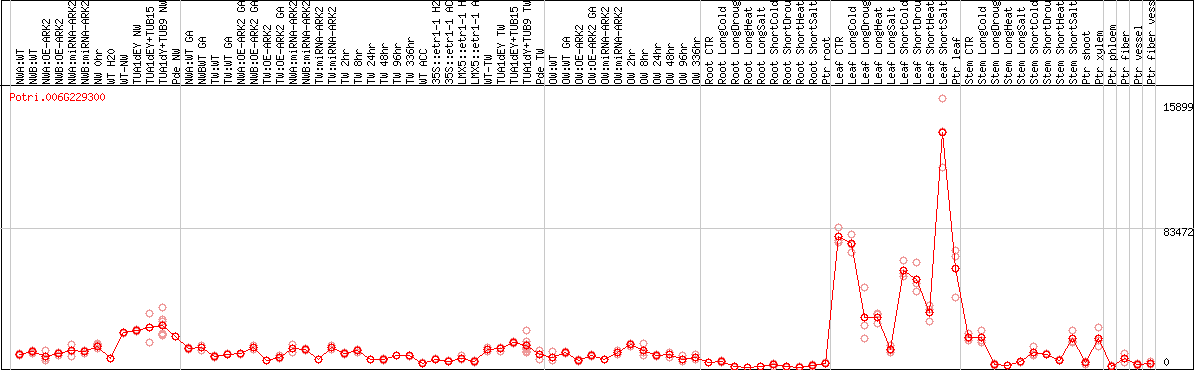
Coexpressed genes
Potri.006G229300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
