Potri.006G234500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G234500.1 pacid=42770177 polypeptide=Potri.006G234500.1.p locus=Potri.006G234500 ID=Potri.006G234500.1.v4.1 annot-version=v4.1
ATGGCTCGGTTTCAGTTCATTTTCACCTTTATACCCATTTTAATTACAATAATAACCCTAACCACAAACCTTAGAGTTCTCTGTTCGGGTTCGGATTCGG
ATTCGGAATCGGGTTCCTTTTCGGTAATTGATTTCGATTCCAATCTCCTCTTCCACCAAGACTACTCGCCACCAGCACCACCTCCACCGCCGCCGCATCC
ACCGTCGGCATCTTGCACTGACGATCTCGGAGGCATCGGTTCGATTGATACTGTATGTCAAATAGTTGCTGATGTGAACCTCACTCGTGACGTGTACATA
GAAGGGAAAGGAGATTTTAATATCCATCCTGGTGTGAGGTTTCATTGCCCTAATTTTGGATGTTCAATAACGATAAATGTCTCAGGGAATTTCAATTTAT
CAGTCAATTCGTCTATTGTGACTGGGACTTTTGAGCTTGTGGCGAATAACGCTAGTTTTTTTAATGGTTCGGTTGTGAATACGACGGGATTAGCTGGAGA
TCCTCCCCCGCAAACGAGTGGAACACCACAGGGCTTAGAAGGGGCGGGCGGAGGGCATGGAGGGAGAGGGGCATGTTGTTTGGTGGATAAAGAGAAGTTG
CCGGAGGATATTTGGGGAGGAGATGCGTATTCTTGGTCAAGTTTGCAGGATCCTTGGAGTTATGGGAGTAAAGGAGGGAGTACGAGTAAGGAAGTGGATT
ATGGAGGGGCAGGAGGTGGGAGAGTGAAAATGAAAGTGAAGGAGTATTTGGCTGTGGATGGAGCTATATTGGCTGATGGTGGTTATGGGGGTGTCAAGGG
AGGTGGTGGTTCGGGTGGGAGCATACTTTTGAAGGCTTATAAAATGACTGGAGGTGGCAGGATAAGCGCTTGTGGAGGCAATGGTTTTGCTGGTGGCGGT
GGTGGAAGAGTTTCAGTTGACATTTTCAGTAGGCATGACGACCCTCAAATTTTTGTGCATGGAGGGAATAGTTTCGGTTGTCCAGAAAATGCAGGTGGTG
CTGGAACTTTATATGATGCTGTTGCCCGGAGCCTTACTGTGAGCAATCACAATATGTCAACAGATACAGATACACTTATTTTGGAGTTTCCTTATCAGCC
TCTCTGGACAAATGTTTATGTTCGAAATCATGCCAGGGCTACTGTTCCATTGCTCTGGAGTCGTGTTCAGGTCCAAGGACAAATAAGTCTGTTATGCAGT
GGAGTCCTAAGCTTTGGGCTAGCTCATTATGCTTCGTCAGAGTTTGAATTATTTGCGGAAGAGCTATTAATGAGTGACTCTGTAATCAAGGTTTATGGGG
CTCTACGCATGAGTGTGAAAATGTTCTTAATGTGGAATTCCAAAATGATCATAGATGGCGGTGAAGATGTAACTGTTGCGACTTCTTTGCTTGAGGCTAG
CAATCTAGTTGTTCTTAAGGAATCATCTGTCATACATTCTAATGCCAATCTGGGGGTTCATGGTCAAGGTCTATTGAATTTGTCTGGATCAGGCAATTGG
ATTGAAGCTCAACGTTTGGTTCTATCACTTTTTTATAGTATTCATGTTGCACCTGGATCTGTGTTGCGTGGTCCTGTGGAGAATGCAACAAGTGATGCAA
TTACGCCAAGGCTCCATTGCCAACTTGAGGAGTGCCCTGCAGAACTTTTTCATCCTCCTGAAGATTGCAATGTGAACTCATCTCTTTCTTTTACCCTTCA
GATATGCCGAGTGGAGGATATTACAGTTGAGGGACTTATAGAAGGATCTGTTGTCCATTTTAATCAGGCAAGAGCAATATCTGTTCCATCTTCTGGAACA
ATAAGTGCATCTGGCATGGGCTGCACTGGTGGTGTGGGTAGAGGTAATGGTTTAAGCAATGGGATTGGCAGTGGTGGCGGCCATGGTGGCAAAGGTGGGT
CTGCATGTTACAATGATAATTGTGTTGATGGTGGTGTTTCATATGGTGATGCAGAGCTACCTTGTGAACTTGGTAGTGGAAGTGGACAGGAAAACTCTTC
TGGTTCTACTGCAGGTGGTGGCATTATAGTAATGGGTTCATTGGAGCATCCTTTGTCAAGTTTGTCTGTTGAAGGTTCAGTCAGAGTTGATGGTGAAAGT
TTTAAAGGAATCACTAGAGACCAACTGGTTGTCATGAAAGGTACTGCTGGAGGTCCTGGCGGTGGTTCTGGAGGAACTATTCTTCTGTTCTTGCATACAC
TAGATCTTGGGGAGCATGCTGTACTTTCAAGTGTTGGGGGATATGGTAGTCCAAAGGGGGGCGGTGGAGGTGGTGGTGGAAGAGTTCACTTCCATTGGTC
AGACATTCCAACTGGGGACATGTACCAGCCCATTGCTCGTGTGAATGGAAGCATCCACACTTGGGGAGGATTGGGAAGAGATGATGGTCATGCTGGGGAA
AACGGAACAGTGACGGGAAAAGCTTGCCCAAAGGGGCTGTATGGAATATTTTGTGAGGAATGTCCAGTTGGCACATATAAGAATGTAACTGGATCTAGTA
GAGTGCTTTGTCATTCGTGCCCTGCTGATGATCTTCCTCGTCGTGCTGCTTATATTGCTGTTCGAGGTGGTATTGCTGAAACTCCATGTCCTTACAAATG
TGTTTCGGAGAGATTTCACATGCCACATTGTTATACAGCCCTAGAAGAGTTGATTTACACGTTTGGTGGGCCATGGCTGTTTTGTCTTCTTCTTTTGGGT
CTCCTCATCCTCTTAGCGCTGGTGCTCAGCGTTGCTCGGATGAAATTTGTTGGGGTGGATGAATTACCTGGTCCAGCTCCCACTCAACATGGCTCTCAGA
TAGATCATTCTTTTCCCTTCCTAGAGTCATTAAATGAGGTTTTGGAAACAAATAGAGCTGAGGAGTCTCAGAGTCATGTGCACAGAATGTATTTTATGGG
CCGTAATACTTTCAGTGAGCCTTGGCATCTTCCCCATACCCCTCCTGAGCAAATAAAAGAGATTGTATATGAGGGTGCATTCAATACATTTGTGGATGAG
ATTAATGGTATAGCTGCTTATCAATGGTGGGAAGGTGCAATTTACATTTTAGTGTCTGTTCTTGCATACCCCCTTGCATGGTCATGGCAGCAATGGAGAC
GGAGAATTAAACTGCAAAGACTGCGAGAGTTTGTTCGATCGGAGTATGACCATGCTTGCTTGCGCTCTTGCCGTTCACGGGCTCTCTATGAAGGGCTTAA
GGTAGCTGCAACCTCTGATTTAATGCTCGGATACTTGGATTTCTACCTTGGTGGAGATGAGAAGAGAACTGATATTCCTGCTCGCCTACATCAAAGATTT
CCAATGTCTATACTTTTTGGAGGTGATGGAAGTTACATGGCTCCTTTCTCAATTCAAAGTGACAATATTCTTACCAGCCTCATGAGCCAGATGGTCCCTT
CTACCACTTGGTATCGAATAGCTGCTGGTTTGAATGCACAACTGCGCTTGGTTTGTCGTGGGCGGCTAATAGTAACTTTTCGACCTGTCCTTAGATGGCT
TGAAACGCATGCTAATCCAGCTTTGAGAAACCATGGAGTGCATGTTGATCTTGCGTGGTTTCAGGCTACTACTTCTGGTCATTGCCAGTATGGACTTTTG
GTTCATGCTGTTGAAGAAGAAAGTGAACATACATCTGTTGAAGGCATTGATGGTGCAAAACAAATTGAAGAAGACTCACGCCTCGTGAAGAACACAAACC
AAGAAAATCCATCTGGTCATTGGAGAGAAGAGGTGTTTGTATCCCAAGCCCACAGAAACAGGGACAACTACATGCGGCGGAAAAGGATTTATGGGGGGAT
TATAGATACTAACAGCTTAAGAATGCTGGAAGAGAAGAGGGATTTATTTTACCTCATCTCATTCATAGTTCATAACACTAAACCTGTTGGCCATCAGGAT
CTTGTTGGTTTGGTCATCTCTACTCTTCTTTTAGGAGATTTTAGCTTAGTGTTGCTTACTTTACTCCAGCTGTATTCAATTTCATTGGCGGGTGTCTTTC
TGGTTTTGTTTATTTTACCTCTTGGCATTCTTATGCCCTTTCCTGCGGGAATCAATGCTTTATTCAGTCATGGACCTAGGCGCTCTGCAGGCCTTGCACG
TATCTATGCTTTGTGGATTGTTACATCCTTGATTAATGTTGTGGTTGCTTTTATTTGTGGATATATTCACTATAACAGTCAATCATCTTCGAGTAAGAAG
TTCCCCTTTCAGACATGGAGTATTAGCATGGATGAAAGTGAATGGTGGATTTTTCCAGCTGGACTGGTTGTGTGTAAAATCTTACAGTCTCAACTTATAA
ATTGGCACGTTGCAAATCTGGAGATTCAAGACCGCTCACTGTATAGTAACGATTTTGAGCTATTCTGGCAGTCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G234500.1 pacid=42770177 polypeptide=Potri.006G234500.1.p locus=Potri.006G234500 ID=Potri.006G234500.1.v4.1 annot-version=v4.1
MARFQFIFTFIPILITIITLTTNLRVLCSGSDSDSESGSFSVIDFDSNLLFHQDYSPPAPPPPPPHPPSASCTDDLGGIGSIDTVCQIVADVNLTRDVYI
EGKGDFNIHPGVRFHCPNFGCSITINVSGNFNLSVNSSIVTGTFELVANNASFFNGSVVNTTGLAGDPPPQTSGTPQGLEGAGGGHGGRGACCLVDKEKL
PEDIWGGDAYSWSSLQDPWSYGSKGGSTSKEVDYGGAGGGRVKMKVKEYLAVDGAILADGGYGGVKGGGGSGGSILLKAYKMTGGGRISACGGNGFAGGG
GGRVSVDIFSRHDDPQIFVHGGNSFGCPENAGGAGTLYDAVARSLTVSNHNMSTDTDTLILEFPYQPLWTNVYVRNHARATVPLLWSRVQVQGQISLLCS
GVLSFGLAHYASSEFELFAEELLMSDSVIKVYGALRMSVKMFLMWNSKMIIDGGEDVTVATSLLEASNLVVLKESSVIHSNANLGVHGQGLLNLSGSGNW
IEAQRLVLSLFYSIHVAPGSVLRGPVENATSDAITPRLHCQLEECPAELFHPPEDCNVNSSLSFTLQICRVEDITVEGLIEGSVVHFNQARAISVPSSGT
ISASGMGCTGGVGRGNGLSNGIGSGGGHGGKGGSACYNDNCVDGGVSYGDAELPCELGSGSGQENSSGSTAGGGIIVMGSLEHPLSSLSVEGSVRVDGES
FKGITRDQLVVMKGTAGGPGGGSGGTILLFLHTLDLGEHAVLSSVGGYGSPKGGGGGGGGRVHFHWSDIPTGDMYQPIARVNGSIHTWGGLGRDDGHAGE
NGTVTGKACPKGLYGIFCEECPVGTYKNVTGSSRVLCHSCPADDLPRRAAYIAVRGGIAETPCPYKCVSERFHMPHCYTALEELIYTFGGPWLFCLLLLG
LLILLALVLSVARMKFVGVDELPGPAPTQHGSQIDHSFPFLESLNEVLETNRAEESQSHVHRMYFMGRNTFSEPWHLPHTPPEQIKEIVYEGAFNTFVDE
INGIAAYQWWEGAIYILVSVLAYPLAWSWQQWRRRIKLQRLREFVRSEYDHACLRSCRSRALYEGLKVAATSDLMLGYLDFYLGGDEKRTDIPARLHQRF
PMSILFGGDGSYMAPFSIQSDNILTSLMSQMVPSTTWYRIAAGLNAQLRLVCRGRLIVTFRPVLRWLETHANPALRNHGVHVDLAWFQATTSGHCQYGLL
VHAVEEESEHTSVEGIDGAKQIEEDSRLVKNTNQENPSGHWREEVFVSQAHRNRDNYMRRKRIYGGIIDTNSLRMLEEKRDLFYLISFIVHNTKPVGHQD
LVGLVISTLLLGDFSLVLLTLLQLYSISLAGVFLVLFILPLGILMPFPAGINALFSHGPRRSAGLARIYALWIVTSLINVVVAFICGYIHYNSQSSSSKK
FPFQTWSISMDESEWWIFPAGLVVCKILQSQLINWHVANLEIQDRSLYSNDFELFWQS
|
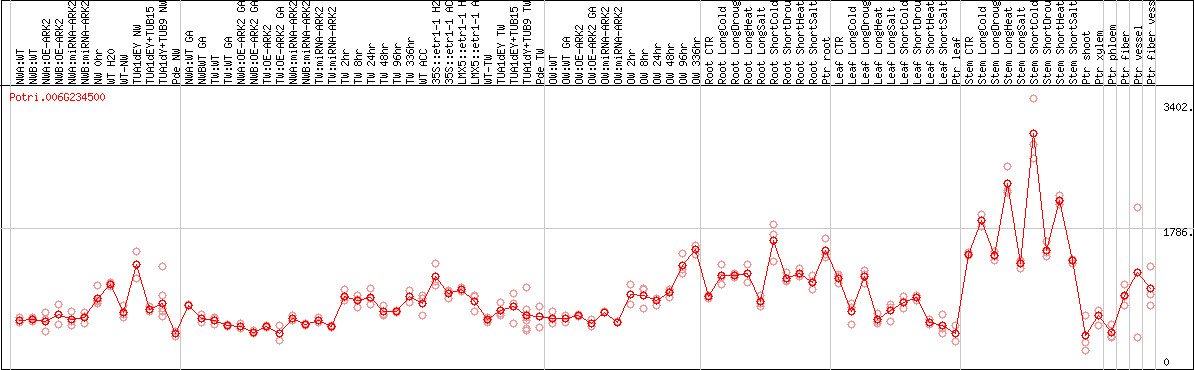
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G234500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
