Potri.006G240300 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G240300.1 pacid=42767413 polypeptide=Potri.006G240300.1.p locus=Potri.006G240300 ID=Potri.006G240300.1.v4.1 annot-version=v4.1
ATGGCCATGGCGATTAGGTCTTTGCTTATCTTTTTATTTATTCTATCACTTACTGTCCCTACCTTTTCTCTTCATGAAGATCAAGTTGGTCTCATGGACT
GGCATCAAAAGTACATAGGGAAAGTGAAGCACGCTGTGTTCCAAACACAAAAAACGGGTCGAAAGCGTGTTCTTGTTTCTACTGAAGAGAATGCTATTGC
TTCTCTTGATCTCCGGCATGGCGAGATTTTTTGGAGGCATGTTCTTGGGGCCAATGATGCTATCGATGGAATCGACATTGCCATGACAAAATATGCTATT
ACCCTTTCTTCAGGAGGAAGCATTCTACGAGCATGGAATCTTCCTGATGGGCAGATGGTGTGGGAATCTTTTCTTCAGGGACCAATCGACTCTAAGTCAT
TCTTGTTTGTGTCGACAAGCTCGAAAGTTGACAAGGACAACACAATCCTTGTTTTTGGTAAAGGAAGTCTCCATGCTGTTTCAAGCATACATGGTGAAAT
TGTTTGGAAGATAGATTTTCCTTCTGAAAGTTTTGAGGTTCAGGAGGTAATTCAGCACCATGATGGCAATACAATATACGTGGTGGGATTTGTTGGTTCT
TCCCAGTTTGATGTGTACCAAATCAATGCTAAGAATGGAGAGCTCCTAAAACATGACAGTGCAGCCGTTGATGGTGGGTTTTCAGGGGAAGTTTCTCTAG
TTTCAAGAGCCAAACTTGTGGTGCTGGATGCTGCTAGGTCAACTTTGTTAACAATAAGTTTTCAGAGCGGAGAAATCAGCTTTCAGAAAACATACATCTC
AGATCTAGTTGAGGATTTCTCAGGGATTGCAGTGATATTACCTTCAAAACTTACTGGTTTGTTTGCTGTGAAAACCAATACAGCTACAGCATTTATTAGT
GTGTCAAGTGAAGGTAAGTTGGAGGTGGTTGACAAAATCAAGCATGCAACAGTTATTAGTAATGTTCTTTCAATCTCAGAGGACCAACAGGCTTTTGCAC
TGGTTCAGCATGGGGGCAATGACATTCACCTCAATGTGAAGCAAGTTCATGACTGGAACAGTGATTTGCTAAAAGAGAGAATCAAGCTGGACAAGCAAAG
AGGACTTGTCCATAAGGTTTTCATTAACAATTATGTCCGGACTGATAAGTCTCATGGATTTAGGGCTTTGATTGTTATGGAAGATCATTCATTGTTGCTG
TTACAACAAGGCGAGGTTGTCTGGAGTAGGGAGGATGGCCTTGCCTCAATAATAGGTGTTACCACGTCAGAACTTCCTGTGGAAAGGGAAGGTGTATCTG
TGGCAAAAGTGGAGCAGAACCTCTTCGAATGGCTTAAGGGACACATGCTGAAGGTTAAGGGTACCTTAATGCTTGCAAGTGCTGAGGATGTAGCAGCCAT
TCAAGGCATGAGGTTGAAAAGCTCTGAGAAAAGTAAAATGATCAGGGACCATAATGGATTCCGAAAGCTGCTTATTGTGCTCACTAAATCCAGGAAACTT
TTCGCATTGCACACTGGAGATGGACGGATTGTCTGGTCTCTCTTATTGAACTCTCTACGTCAAACAGAAGCATGTGAAAATCCAACTGGGATTAATGTGT
ACCAGTGGCAAGTTCCTCATCATCATGCTATGGATGAAAATCCATCTGTCCTGGTTGTTGGCAGGTGCAGAACTGGTACTGATGCACCGGGAATATTTTC
CTACGTTGATACTTACACAGGGAAGGAACTTAAGTCTTTTGGTCTTGATCATTCTGTTGCACAAGTAATTCCACTCCCGCTTACAGATTCAACTGAACAA
CAGCTGCATCTACTAATAGATGCTAATGGACAAGCACATTTGTACCCAAGAGCACCTGAGGCTGCTGCCATTTTTCAACGTGAGTTTTCAAACATCTACT
GGTACTCGGTAGAAGCTGACAAGGGTGTTATTAAGGGACATGGATTGCAGAGCAATTGTGATGGTGAAGTTGCAGACAACTATAGTTTTGGTACCAGAGA
AATATGGTCCATTGTATTCCCCTCAGAATCAGAAAAGATCATTTCAACCGTAACAAGAAAATCAAATGAGGTTGTTCATACCCAAGCAAAGGTTATAGCT
GATCAAGATGTGATGTACAAGTATATATCAAAAAAGTTGCTTTTTGTGGCCACTGTTTCACCAAAAGCCAGTGGAGATATTGGATCAGCCACACCAGGAG
AATCACAGTTGGTTGTCTATGTTGTTGATACTGTCACTGGTCGTATATTACACCGAATGACTCATCATGGCTCACAGGGCCCTGTCCATGCAGTTTTTAG
TGAAAACTGGATTGTCTACCACTACTTCAATCTCAGAGCACACAGATACGAGATGACAGTTATTGAAATTTATGATCAGTCCCGGGCAGACAACAAAGAT
GTTTTAAAGCTTGTTCTTGGAAAGCACAATCTCACTTCACCTATTTCTTCGTATTCGCGACCAGAGGTCACAACAAAGTCACAGTCTTACTATTTCACCC
ATTCTATCAAAGCAATAACAGTTACATCAACTGCTAAGGGTATCACTTCGAAGCATCTTCTTATTGGGACAATTGGTGATCAGGTCTTGGCAATGGATAA
GCGTTTCTTTGACCCTCGTAGGTCAGTTAACCCCACACAATCAGAGAAGGAAGAAGGAATTTTACCTCTTACAGATTCATTACCTATCATTCCTCAGTCG
TATGTTACACACTCCCATAAAGTGGAGGGGCTACGAGGCATAGTAACAGTGCCTGCCAAGCTGGAGTCAAACACCCTTGTCTTTACATATGGTGTTGATC
TCTTTTTCACTCGCCTTGCACCCTCAAGGACCTATGATTCTCTTACAGAGGATTTTAGCTATGCCCTGCTGCTCATAACCATTGTTGCACTTGTCGTGGC
AATCTTCGTGACTTGGGTCCTGTCCGAGAAGAAAGATCTAAGTGATAAATGGAGGTGA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.006G240300.1 pacid=42767413 polypeptide=Potri.006G240300.1.p locus=Potri.006G240300 ID=Potri.006G240300.1.v4.1 annot-version=v4.1
MAMAIRSLLIFLFILSLTVPTFSLHEDQVGLMDWHQKYIGKVKHAVFQTQKTGRKRVLVSTEENAIASLDLRHGEIFWRHVLGANDAIDGIDIAMTKYAI
TLSSGGSILRAWNLPDGQMVWESFLQGPIDSKSFLFVSTSSKVDKDNTILVFGKGSLHAVSSIHGEIVWKIDFPSESFEVQEVIQHHDGNTIYVVGFVGS
SQFDVYQINAKNGELLKHDSAAVDGGFSGEVSLVSRAKLVVLDAARSTLLTISFQSGEISFQKTYISDLVEDFSGIAVILPSKLTGLFAVKTNTATAFIS
VSSEGKLEVVDKIKHATVISNVLSISEDQQAFALVQHGGNDIHLNVKQVHDWNSDLLKERIKLDKQRGLVHKVFINNYVRTDKSHGFRALIVMEDHSLLL
LQQGEVVWSREDGLASIIGVTTSELPVEREGVSVAKVEQNLFEWLKGHMLKVKGTLMLASAEDVAAIQGMRLKSSEKSKMIRDHNGFRKLLIVLTKSRKL
FALHTGDGRIVWSLLLNSLRQTEACENPTGINVYQWQVPHHHAMDENPSVLVVGRCRTGTDAPGIFSYVDTYTGKELKSFGLDHSVAQVIPLPLTDSTEQ
QLHLLIDANGQAHLYPRAPEAAAIFQREFSNIYWYSVEADKGVIKGHGLQSNCDGEVADNYSFGTREIWSIVFPSESEKIISTVTRKSNEVVHTQAKVIA
DQDVMYKYISKKLLFVATVSPKASGDIGSATPGESQLVVYVVDTVTGRILHRMTHHGSQGPVHAVFSENWIVYHYFNLRAHRYEMTVIEIYDQSRADNKD
VLKLVLGKHNLTSPISSYSRPEVTTKSQSYYFTHSIKAITVTSTAKGITSKHLLIGTIGDQVLAMDKRFFDPRRSVNPTQSEKEEGILPLTDSLPIIPQS
YVTHSHKVEGLRGIVTVPAKLESNTLVFTYGVDLFFTRLAPSRTYDSLTEDFSYALLLITIVALVVAIFVTWVLSEKKDLSDKWR
|
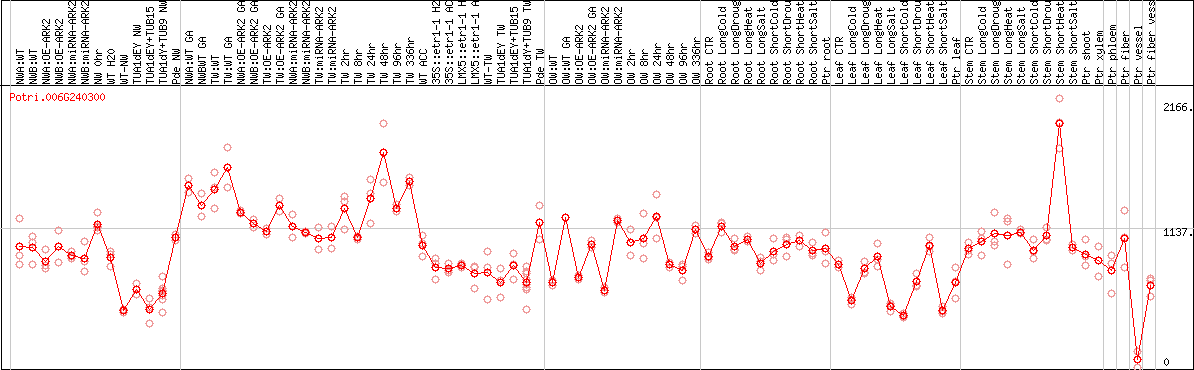
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G240300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
