External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25760 315 / 2e-112
PEX4, UBC21
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
AT2G16740 121 / 8e-36
UBC29
ubiquitin-conjugating enzyme 29 (.1)
AT5G53300 121 / 1e-35
UBC10
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
AT4G27960 120 / 2e-35
UBC9
ubiquitin conjugating enzyme 9 (.1.2)
AT1G16890 120 / 2e-35
UBC36 ,UBC13B
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
AT1G64230 120 / 3e-35
UBC28
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
AT1G78870 119 / 5e-35
UBC35 ,UBC13A
UBIQUITIN CONJUGATING ENZYME 13A, ubiquitin-conjugating enzyme 35 (.1.2.3)
AT3G08690 119 / 5e-35
ATUBC11, UBC11
ubiquitin-conjugating enzyme 11 (.1.2)
AT5G41700 118 / 2e-34
ATUBC8, UBC8
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
AT1G14400 117 / 4e-34
ATUBC1, UBC1
UBIQUITIN CONJUGATING ENZYME 1, ubiquitin carrier protein 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G039200
324 / 8e-116
AT5G25760 314 / 5e-112
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
Potri.001G392500
121 / 1e-35
AT1G78870 310 / 1e-110
UBIQUITIN CONJUGATING ENZYME 13A, ubiquitin-conjugating enzyme 35 (.1.2.3)
Potri.011G168200
119 / 4e-35
AT1G64230 295 / 1e-104
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Potri.011G111400
119 / 5e-35
AT1G78870 311 / 1e-110
UBIQUITIN CONJUGATING ENZYME 13A, ubiquitin-conjugating enzyme 35 (.1.2.3)
Potri.001G471200
119 / 8e-35
AT1G64230 292 / 2e-103
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Potri.004G175000
119 / 8e-35
AT5G53300 292 / 2e-103
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Potri.013G064400
117 / 4e-34
AT2G02760 313 / 1e-111
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Potri.006G110200
117 / 4e-34
AT5G41700 304 / 4e-108
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
Potri.019G131400
117 / 4e-34
AT5G41700 304 / 4e-108
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009495
277 / 2e-95
AT5G25760 273 / 1e-93
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
Lus10011695
256 / 7e-86
AT5G25760 253 / 3e-84
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
Lus10017654
123 / 2e-36
AT1G16890 311 / 5e-111
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
Lus10033611
123 / 2e-36
AT1G16890 311 / 5e-111
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
Lus10030786
118 / 2e-34
AT2G02760 305 / 2e-108
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10014187
118 / 2e-34
AT1G64230 303 / 7e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10022726
118 / 2e-34
AT1G64230 303 / 7e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10037203
118 / 2e-34
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10036727
118 / 2e-34
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10027846
118 / 2e-34
AT1G64230 289 / 3e-102
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.006G240900.2 pacid=42769249 polypeptide=Potri.006G240900.2.p locus=Potri.006G240900 ID=Potri.006G240900.2.v4.1 annot-version=v4.1
ATGCAGGCATCGAGAGCGAGGTTATTAAAGGAATACAAGGAGGTTCAGCGAGAGAAAGTAGCTGATCCGGATATTCAATTAGTTTGTGATGATTCTAACA
TTTTTAAATGGACTGCTCTGATCAAGGGACCGTCGGAGACTCCTTTTGAGGGTGGAGTTTTTCAGCTTGCGTTTTCTGTTCCTGAACAGTATCCTTTGCA
GCCTCCGCAAGTGCGGTTCTTGACGAAAATATTTCACCCAAATGTGCATTTTAAGACAGGAGAAATATGCCTTGATATATTGAAGAATGCCTGGAGCCCA
GCTTGGACTCTTCAGTCTGTTTGTAGGGCTATAATTGCTTTGATGGCCCACCCAGAACCTGATAGCCCTCTAAACTGTGATTCAGGCAATCTTCTACGTT
CTGGTGATGTTAGAGGTTATCAATCCATGGCAAGAATGTATACCAGACTTGCAGCCTCGCCAAAGAAAGGATGA
AA sequence
>Potri.006G240900.2 pacid=42769249 polypeptide=Potri.006G240900.2.p locus=Potri.006G240900 ID=Potri.006G240900.2.v4.1 annot-version=v4.1
MQASRARLLKEYKEVQREKVADPDIQLVCDDSNIFKWTALIKGPSETPFEGGVFQLAFSVPEQYPLQPPQVRFLTKIFHPNVHFKTGEICLDILKNAWSP
AWTLQSVCRAIIALMAHPEPDSPLNCDSGNLLRSGDVRGYQSMARMYTRLAASPKKG
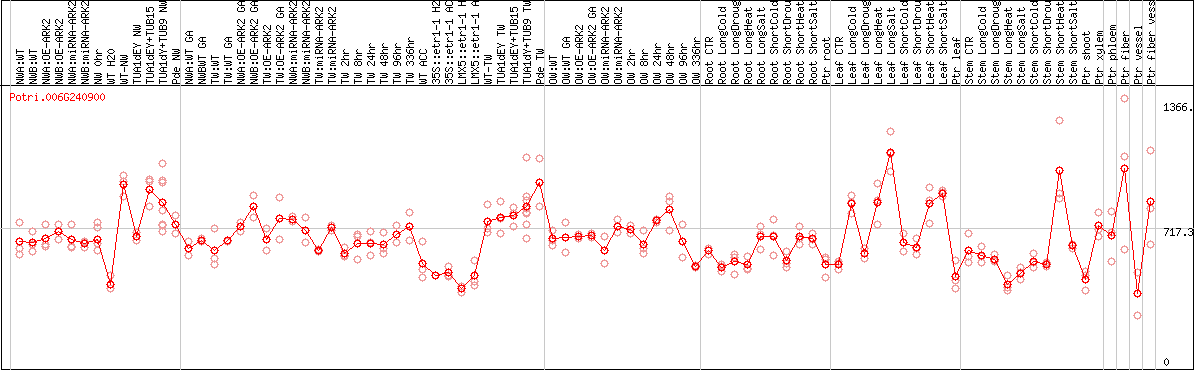
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G240900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.