External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G24570 214 / 3e-71
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
AT2G14860 57 / 3e-10
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
AT5G43140 54 / 4e-09
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
AT4G33905 52 / 3e-08
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
AT4G14305 42 / 5e-05
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
AT1G52870 42 / 0.0001
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
AT5G19750 41 / 0.0002
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G037100
272 / 5e-94
AT3G24570 289 / 6e-100
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Potri.018G081600
209 / 4e-69
AT3G24570 309 / 7e-108
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Potri.001G296400
58 / 2e-10
AT2G14860 300 / 6e-103
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Potri.009G090600
54 / 7e-09
AT4G33905 299 / 1e-102
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Potri.010G068900
48 / 5e-07
AT4G14305 252 / 1e-86
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Potri.001G404600
46 / 3e-06
AT1G52870 438 / 6e-154
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Potri.006G032500
45 / 9e-06
AT5G19750 279 / 8e-94
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Potri.011G123700
44 / 2e-05
AT1G52870 472 / 1e-167
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038626
225 / 2e-75
AT3G24570 273 / 1e-93
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10037900
214 / 1e-70
AT3G24570 246 / 3e-82
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10034922
205 / 2e-67
AT3G24570 330 / 4e-116
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10023648
203 / 1e-66
AT3G24570 330 / 6e-116
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10011679
185 / 8e-60
AT3G24570 249 / 2e-84
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10022130
180 / 9e-58
AT3G24570 256 / 6e-87
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10011424
59 / 2e-10
AT2G14860 288 / 3e-98
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Lus10002118
57 / 5e-10
AT2G14860 292 / 6e-100
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Lus10018700
49 / 5e-07
AT5G43140 253 / 2e-83
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
Lus10007761
44 / 3e-05
AT5G43140 223 / 1e-70
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04117
Mpv17_PMP22
Mpv17 / PMP22 family
Representative CDS sequence
>Potri.006G242700.1 pacid=42767963 polypeptide=Potri.006G242702.1.p locus=Potri.006G242700 ID=Potri.006G242700.1.v4.1 annot-version=v4.1
ATGTTGGAGTTGTGGAAATGGTACCAAAATTGCTTGGCTGTGCATCCAGTTAAGAAACAAGTGATTAGCTCAGGGTTTATTTGGGGATTTGGTGATATAG
CTGCAAAATCCATCGCTCACCATACAGCCAAGAAATATCATCAAATCAAGCACTTCACTGCTACTGTCAAAAGATTTTTAATTGATGGGACTGCATATAG
TATGAAGGGTTGCCGATTTATTAGATCACGACTTCTAATGCGTCCAAATTCCTTGCGGTTTGTTGGTGCTAAAGTAGCAATTGATGGGTTCCTCTTTGGA
CCACTGGATCTGCTTGTATTTTTCAGTTACATGGGTTTTGCCACTGGAAAGAGCGTTCCCCAAATCAGGAAAGATTTGAAGAGGGACTTGATACCAGCTT
TTGTTTTGGAAGGAGGTATACGGCCAATTGTTCGAGTTGCGAATTTCCGATTTGTACCTGTGAGGTATCAACTCTTGTATGTCAACTTCTTCTGCTTGTT
GGATAGTTGTTTTCTGTCATGGCTTGAGTAG
AA sequence
>Potri.006G242700.1 pacid=42767963 polypeptide=Potri.006G242702.1.p locus=Potri.006G242700 ID=Potri.006G242700.1.v4.1 annot-version=v4.1
MLELWKWYQNCLAVHPVKKQVISSGFIWGFGDIAAKSIAHHTAKKYHQIKHFTATVKRFLIDGTAYSMKGCRFIRSRLLMRPNSLRFVGAKVAIDGFLFG
PLDLLVFFSYMGFATGKSVPQIRKDLKRDLIPAFVLEGGIRPIVRVANFRFVPVRYQLLYVNFFCLLDSCFLSWLE
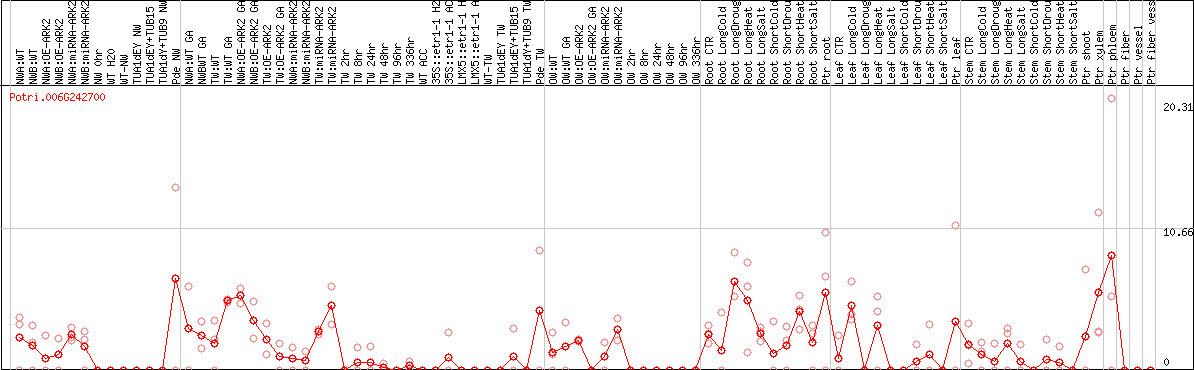
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G242700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.