External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G11480 422 / 1e-149
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G22870 300 / 2e-101
EMB2001
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G58370 102 / 2e-24
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
AT1G08410 45 / 6e-05
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G27200 43 / 0.0002
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G037000
496 / 6e-179
AT5G11480 421 / 7e-149
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.007G006300
299 / 4e-101
AT2G22870 434 / 2e-154
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.014G006501
140 / 2e-41
AT2G22870 199 / 2e-65
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.014G007450
100 / 8e-27
AT2G22870 131 / 3e-39
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.019G128000
106 / 1e-25
AT5G58370 461 / 1e-158
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Potri.009G017000
41 / 0.0007
AT1G08410 755 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011673
426 / 7e-151
AT5G11480 418 / 1e-147
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10022126
424 / 2e-150
AT5G11480 418 / 8e-148
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10011703
302 / 2e-102
AT2G22870 427 / 5e-152
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10039653
295 / 2e-99
AT2G22870 426 / 2e-151
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10009919
106 / 2e-25
AT5G58370 473 / 9e-165
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Lus10004103
42 / 0.0004
AT1G08410 756 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10013359
42 / 0.0004
AT1G08410 745 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10028574
41 / 0.0009
AT3G07050 743 / 0.0
nucleostemin-like 1, GTP-binding family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF01926
MMR_HSR1
50S ribosome-binding GTPase
Representative CDS sequence
>Potri.006G243200.1 pacid=42767912 polypeptide=Potri.006G243200.1.p locus=Potri.006G243200 ID=Potri.006G243200.1.v4.1 annot-version=v4.1
ATGGCTCTTTCTCAGCTACCAAAATTCACTTACTGCCCACCTTCTCTTTACAACACCCATATCAACCTCCCTCTCCTCGCCAAGCTCAAACTACCAACCC
TTACAAGACTCAAATCCACACTAACCACAACAGAACCTATACCCATCACCGAAGCCCACAATTTCTTGACACCCCAAGAAGAAACCCAAACCCAATTTTC
ACTTGACAAACTATTTATCCCACCAGACACTGAGGTTTCAATCAATGAGAATAGTGGTCTAAGTGCTAGAATTTTGAAAGGGTCTAATATTGTGTTGAGT
AAGTATGCAAGGGATGCACAGGTTGTTCAAGCTGAGTTTATTAAAAGTAGTGTCAGAACTGAAGATTGTCCTTCTGATGGTTTGCCTGAGTTTGCTCTTG
TTGGAAGGTCTAATGTTGGAAAATCTTCTCTTCTTAATTCTCTTGTTAGAAGAAAGAAACTTGCACTTACCTCCAAAAAACCTGGGAAGACGCAGTGCAT
CAATCATTTTAAGGTCAATGATAGCTGGTACTTGGTGGATTTGCCTGGATATGGGTATGCATCTGCACCTCAGGAACTTCGGACAGATTGGAATAAGTTT
ACTAAAGATTATTTTCTCAATCGCTCAACCTTGGTCTCAGTTTTCCTTCTTATTGATGCTAGTATTCCTGCAAAGAAAATTGATCTTGAATATGCCAGCT
GGCTGGGTCAGAATCAGGTTCCCATGACATTCATTTTCACCAAATGCGACAAGCGGAAGAAGAAAAGGAATGGTGGGAAAAGGCCTGAAGAAAATGTGAA
TGAATTTCAGGAGCTCATACGCGGCTTCTTTGAGACAGCACCTCCTTGGATTATGACCAGCGGTGTTACCAATCAAGGTCGTGATGAGATGCTACTGCAC
ATGGCTCAATTGCGGAACTACTGGCTCAAACACTGA
AA sequence
>Potri.006G243200.1 pacid=42767912 polypeptide=Potri.006G243200.1.p locus=Potri.006G243200 ID=Potri.006G243200.1.v4.1 annot-version=v4.1
MALSQLPKFTYCPPSLYNTHINLPLLAKLKLPTLTRLKSTLTTTEPIPITEAHNFLTPQEETQTQFSLDKLFIPPDTEVSINENSGLSARILKGSNIVLS
KYARDAQVVQAEFIKSSVRTEDCPSDGLPEFALVGRSNVGKSSLLNSLVRRKKLALTSKKPGKTQCINHFKVNDSWYLVDLPGYGYASAPQELRTDWNKF
TKDYFLNRSTLVSVFLLIDASIPAKKIDLEYASWLGQNQVPMTFIFTKCDKRKKKRNGGKRPEENVNEFQELIRGFFETAPPWIMTSGVTNQGRDEMLLH
MAQLRNYWLKH
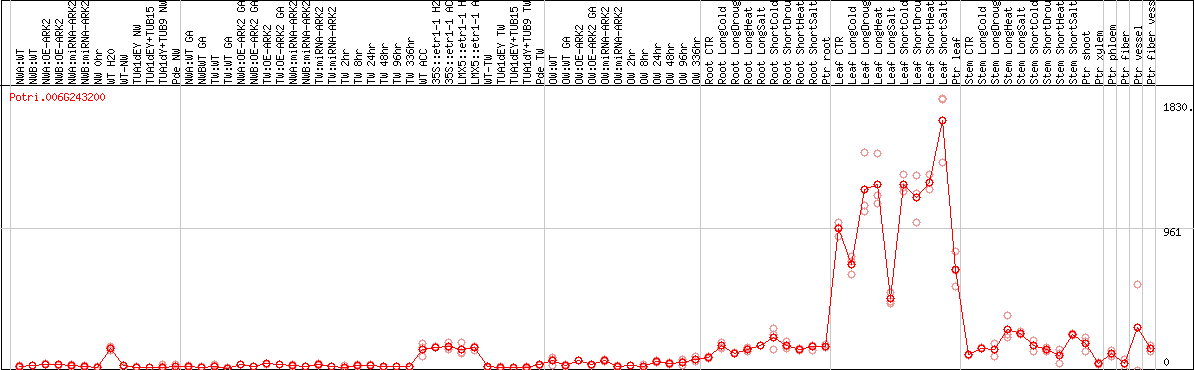
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G243200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.