Potri.006G246200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G246200.2 pacid=42770578 polypeptide=Potri.006G246200.2.p locus=Potri.006G246200 ID=Potri.006G246200.2.v4.1 annot-version=v4.1
ATGGAAAATAGAGTAGGAAAGTCTCATGGAGTGGGAATTCCTAAGAAGTCGAGATCCTTGGATCTCAAGAGTCTCTATGAAACTAAAAATTCAAAATGGT
ATCAAAATAGTAATAATTTGAAGCGGAAGGGTGGTGGTATAGGTGATGATGAGAAGGGACATAAAAACAAGAAGAGTAGAAAGGAGGTTTGCATTAGTAG
TTTTAAGAATGTTAACAGTAGCTATAGTAAGAGTTTAAAGGAAGTGTATAATGGGAGTTTGAGTTCAGGTTTGAAGGACCCGAGGACTGGGTTGATTCAG
AGACTGGCTGATAGTAATGGGTTTAGTGGTGCTTCGTTACCCTTGGAAGACGGGGCTGTTAAAATTCCTAGGCGAAAACGGGGTTTTGTGGGGCGAAGGA
AGGTGGACAATGGTAGTGAAGGGGGCAAGCTGGCAAGAGGGTTTGGCAGGGAAGTGGGCAATGCTGACCAGGCTGATAAGTTAACCGGTGAAGATGAGGG
TAAAGGGGTTGAGAATGGCAGTCAGGAGTCGAAGGCAGTGGTGATTTTGGTTAGTGTAGTGGGTGATGTGGATCAGGCTAGCAAGTTAACTGGTGAAGGT
AAAGCTAAGCAGGTTGAGCATTCAAAGGCTAAACAGAAGAAAGGTTCTGATGATTTGAAAGAAAATAGAAATGGTGAATTGGATGCAAGTAGGCATTTGA
AGGAAGAAGACAGGCATGATGATCATTCAGTTGCTACAAAAAGAGATTCATTGTTGAAGAAGTCTGACAATTGCCCTTTGGTTGTTAATAATGGGGATTC
ATCTCTGAAAAAGTCACTGAGGAAGCGGAGTAGGAAAAAAAAGGATATGGTGTCTAACAAGAAAAGAACCAAGGAGGCTGACCCATCAGTTGATGCTTCT
ATAAAGATCAGTGATGTTTTGCACGATGAGGATGAGGAGAATCTTGAAGAGAATGCAGCAATGATGCTATCATCACGATTTGATCCAAGCTGCACGGGTT
TTTCTTCAAACAGCAAAGCCTCAGCCTCACCATCTAAAGATGGTTTCCAAGAATTTGCTGCTCGTGAGTCTAGCTATGTCTCTGGATCTGAATCTTCATC
TGTTGATACTGATGGTAGAGTATTACGACCTAGGAAACAGAACAAAGAGAAGGGTAACACGAGGAAACGGCGCCATTATTATGAAATTTTCTCTGGGGAC
TTGGATGCACATTGGGTTCTCAACCGGAGAATAAAGGTCTTTTGGCCTTTGGACCAGAGTTGGTATCATGGTCTTGTTGGTGACTATGACAAAGATAGGA
AGCTTCATCATGTAAAATATGATGACCGGGATGAGGAATGGATTAATCTCCAAAATGAAAGATTTAAACTCTTGATGCTCCCTTGTGAAGTTCCAGCTAA
GACCAGAAGGAAAAGGTCAGTGACAAGAAACAAGTGCTCCAATGGGGGAAAGGAAAAACTGATGTCTAGAAAAGAAAAGAGGGACCTGATGACAGAGGAT
GATAGCTATGAAGGGGCTTATATGGACTCAGAGCCTATCATCTCGTGGTTGGCTCGATCTACTCATAGAGTCAAATCTTCTCCTCTTTGTGCCTTGAAGA
AACAGAAAACTTCTTATCTATCTTCTACTAGGACACCACTGTCTTCCCTAAACAGGGATAGAGGCAAGTTATGTAGTAATTCTGCTTCATCAGAGAGTGT
CGCCACTGATGGAAGGAGTGGTTTGCCTGTGATGGAGAAGCCTGTTTACCCCAAAGGTAGTAAGCTCCCTATTGTTTATTACAGGAAGCGGTTCCGCGAG
ACAAGCAATGTGTTGTGTCATGAATCCAAGGGTGTTCATATTTCTGCTAGTGTAGCAGAATCTGTTAGATCTCTGGTGCATCATACTGTTAATTCTGGGG
CTTTGGAAGAGCATGATACTTCTCTTGGAAGATTGAATCCTGATGAGGATTTGGATAGGTTGGATGCTTTTGATCCTTTATGGTCAACTAATAAAGCAGG
GTTGCTGAGATTAAATATTTCTGCCATAGAACCAAGATGGTTCAGGTTCAAGTTAAGCTTCCTATTGCCTTCTGTCCCCCGTCACTACTCATTTGGCTCA
GAAATTGTTTGGTTGATCCATGCTATGGCACTGCTTCAGTATGGTATGCTTATGACTACATGGCCAAGGATTCATTTAGAGATGCTTTTTGTCGATAATG
GGGTTGGATTGAGGTTTCTCCTGTTTGAAGGTTGCTTGAAAGAGGCTGTAGCTTTTGTTTTCCTGGTCTTGACAATATTTTATCAACCTAATGAGCAGCA
GGGGAAATGTGCTGACTTCCAGTTGCCAATAACTTCAATCAGGTTCAAATTCTCTTGCATTCAAGATTTCAGAAAGCAGTTTGCCTTTGCATTCCACAAC
TTCTCTGAAGTAGAGAATTCAAAGTGGATATACCTGGACCATAAGCTCAAAAAGCATTGCTTGCTTTCTAGGCAATTACCTCTGTCTGAATGCACTTATG
ATAATGTGAAGGCCTTGCAATGTGGAATGAATCAGCTCCTTAGTCCTTGGGCCTGCAGTGATGCCACCTTAAATAAGGTCTCGCATAGGAGGTCTAGAGA
GAGCATCGGTCTCGTGGGGTTCTCCAGAGAATCTACTTGTGTTAATGCTAATCTGTCTTCCTCTAAGTCTGATAAAAATCACAGATATTTACCTTCATTT
GCGCTTTCTTTTACTGCTGCTCCTACTTTCTTTTTGGGTTTGCATCTGAAGATGCTTATGGAACATAGTATGATGCACATTAACTTTCTGGATCACGATT
CAATAGAGCATCCAGAAAAATCCAGTGGTTTGCTGGCTGACAGCTGTTCTAGTGTAGAGGACTGCTCCAAAGAATATTTAGATGGTACTCCTGGTAATGA
TTTTAAAGCTTTGTCGATGGGTGCTGACTTTGATGGATGCATATCTCGTGCTAAACCAGAGTCACAAACAGTTGATGGGACTGATCCTGGTTCTCGTACT
CTTCTGAAGGGTATTACGGTTGAAATTCCATCAGTTAATTTGAACCAGCATGTTAATAAGGAATTACACAGTGTTCAACGGTCCTCTGATCTGTCATGGA
ATATGAATGGTGGCATTATTCCCAGCCCAAACCCTACTGCTCGCAGAAGTACCTGGTATCGAAATAGAAGTAGTTCTGCATCTTTTGGGTGGTCTGATGG
AAGGACTGACTTTCTTCAAAACAATTTTGGTAATGGACCTAAGAAGCCACGGACACATGTTTCTTATACATTGCCTTTAGGAGGTTTTGATTACAGCCCG
AGGAATAGAGGACAACAGCAGAAAGGGTTTTCACATAAACGAATCAGGACAGCCACTGAGAAGAGGACTTCAGACATTTCCAGGGGTTCTGAAAGAAACT
TAGAGTTGTTGTCTTGTGATGCAAATGTGTTGATTACAAATGGTGACAAAGGATGGAGAGAATGTGGGGTGCAGGTTGTACTAGAGCTTTTCGATCATAA
TGAGTGGAGGCTTGGTATTAAGCTTTCAGGAACTACAAAGTATTCATACAAGGCACATCAGTTTCTGCAAACTGGGTCAACAAATCGGTTTACACATGCT
ATGATGTGGAAAGGAGGAAAAGAATGGACATTGGAATTTCCTGATAGGAGTCAGTGGGTTCTTTTTAAAGAGATGCATGAAGAATGCTACAATCGGAACA
TGCGAGCTGCATCAGTTAAAAACATTCCTATTCCTGGGGTCTGCCTGATAGAGGAAAATGATGATAATGGAATAGAAGCACCATTTTTCCGGGGTTTTAA
GTACTTTCAGCAGCTGGAAACAGATGTTGAGTTGGCTTTGAATCCATCAAGGGTGTTATATGATATGGATAGTGATGATGAAAAGTGGATGCTGAAAAAT
CGAAGTTCTCCAGAAGTCAACAGTAGTTCACGGCAGATTTCAGAAGAAATGTTTGAGAAGGCGATGGACATGTTTGAGAAGGCTGCATATTCTCAACAGC
GGGACCAGTTTACATCTGATGAAATAATGAAACTTATGGCTGGAATAGGGCCCACAGGAGCCATCAAAATCATCCATGAATATTGGCAGCATAAGAGGCA
GAGGAAGAGAATGCCTCTGATCCGTCATCTCCAGCCACCACTCTGGGAAAGGTATCAGCAGCAATTGAGGGAGTGGGAGCAAGCAATGGAAAGAAGCAGC
ACAAGCCTCCCCAGTGGATGCCATGGGAAGGTTGCACTCGAAGACAAGCCACCCATGTATGCTTTCTGTTTGAAACCTCGGGGTTTGGAAGTGCCTAACA
AGGGATCAAAACAAAGGTCACACAGGAAATTTTCAGTTGCTGGAAAAAGCAATTCTTTTGCAGGAGATCATGATGGCTTTCATCCTTATGGAAGAAGAAT
AAATGGCTTTGCTTCTGGAGATGAAAAGACTATATATCCAATACATAACAATGAATCTTTTGATGATTCCCCATTGCCTCGGATATCACCCAGGTTCTTC
TCACCACAAGATGCTTGTGCCCCTAGATATTTCTCAATGACAGGTGATAGGTCTGATAGAAATCATCTCCAGAAACTTCGCAGGACCAAGTCAAAGAAAC
TTGGGACCTGTGTGTCTCCCTATGGTACTCAGATGGCTGCTTTATATAACCAGAGAATGATGGATCAGGGAAATGGATTTCATCGGTGGAATGCAAGTTT
CTCTGACTGGCCTAGTCAGCAGCATCACCAGATAGATTTTAATGTGAGACATGGCCTTGAACAGTTAAATGGTTCAGATCTTGATGAATTCAGGTTGCGT
GATGCTTCTGGTGCAGCTAAGCATGCACTCAATATGGCTAATATCAAGAGAGAAAGGGCACAAAGATTGCTTTACAGGGCAGATCTTGCAATTCACAAGG
CTGTGGTTGCTCTAATGAATGCTGAGGCGATAAAAGCTTCTTCTGAGGACTTAAATGGTGATGGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G246200.2 pacid=42770578 polypeptide=Potri.006G246200.2.p locus=Potri.006G246200 ID=Potri.006G246200.2.v4.1 annot-version=v4.1
MENRVGKSHGVGIPKKSRSLDLKSLYETKNSKWYQNSNNLKRKGGGIGDDEKGHKNKKSRKEVCISSFKNVNSSYSKSLKEVYNGSLSSGLKDPRTGLIQ
RLADSNGFSGASLPLEDGAVKIPRRKRGFVGRRKVDNGSEGGKLARGFGREVGNADQADKLTGEDEGKGVENGSQESKAVVILVSVVGDVDQASKLTGEG
KAKQVEHSKAKQKKGSDDLKENRNGELDASRHLKEEDRHDDHSVATKRDSLLKKSDNCPLVVNNGDSSLKKSLRKRSRKKKDMVSNKKRTKEADPSVDAS
IKISDVLHDEDEENLEENAAMMLSSRFDPSCTGFSSNSKASASPSKDGFQEFAARESSYVSGSESSSVDTDGRVLRPRKQNKEKGNTRKRRHYYEIFSGD
LDAHWVLNRRIKVFWPLDQSWYHGLVGDYDKDRKLHHVKYDDRDEEWINLQNERFKLLMLPCEVPAKTRRKRSVTRNKCSNGGKEKLMSRKEKRDLMTED
DSYEGAYMDSEPIISWLARSTHRVKSSPLCALKKQKTSYLSSTRTPLSSLNRDRGKLCSNSASSESVATDGRSGLPVMEKPVYPKGSKLPIVYYRKRFRE
TSNVLCHESKGVHISASVAESVRSLVHHTVNSGALEEHDTSLGRLNPDEDLDRLDAFDPLWSTNKAGLLRLNISAIEPRWFRFKLSFLLPSVPRHYSFGS
EIVWLIHAMALLQYGMLMTTWPRIHLEMLFVDNGVGLRFLLFEGCLKEAVAFVFLVLTIFYQPNEQQGKCADFQLPITSIRFKFSCIQDFRKQFAFAFHN
FSEVENSKWIYLDHKLKKHCLLSRQLPLSECTYDNVKALQCGMNQLLSPWACSDATLNKVSHRRSRESIGLVGFSRESTCVNANLSSSKSDKNHRYLPSF
ALSFTAAPTFFLGLHLKMLMEHSMMHINFLDHDSIEHPEKSSGLLADSCSSVEDCSKEYLDGTPGNDFKALSMGADFDGCISRAKPESQTVDGTDPGSRT
LLKGITVEIPSVNLNQHVNKELHSVQRSSDLSWNMNGGIIPSPNPTARRSTWYRNRSSSASFGWSDGRTDFLQNNFGNGPKKPRTHVSYTLPLGGFDYSP
RNRGQQQKGFSHKRIRTATEKRTSDISRGSERNLELLSCDANVLITNGDKGWRECGVQVVLELFDHNEWRLGIKLSGTTKYSYKAHQFLQTGSTNRFTHA
MMWKGGKEWTLEFPDRSQWVLFKEMHEECYNRNMRAASVKNIPIPGVCLIEENDDNGIEAPFFRGFKYFQQLETDVELALNPSRVLYDMDSDDEKWMLKN
RSSPEVNSSSRQISEEMFEKAMDMFEKAAYSQQRDQFTSDEIMKLMAGIGPTGAIKIIHEYWQHKRQRKRMPLIRHLQPPLWERYQQQLREWEQAMERSS
TSLPSGCHGKVALEDKPPMYAFCLKPRGLEVPNKGSKQRSHRKFSVAGKSNSFAGDHDGFHPYGRRINGFASGDEKTIYPIHNNESFDDSPLPRISPRFF
SPQDACAPRYFSMTGDRSDRNHLQKLRRTKSKKLGTCVSPYGTQMAALYNQRMMDQGNGFHRWNASFSDWPSQQHHQIDFNVRHGLEQLNGSDLDEFRLR
DASGAAKHALNMANIKRERAQRLLYRADLAIHKAVVALMNAEAIKASSEDLNGDG
|
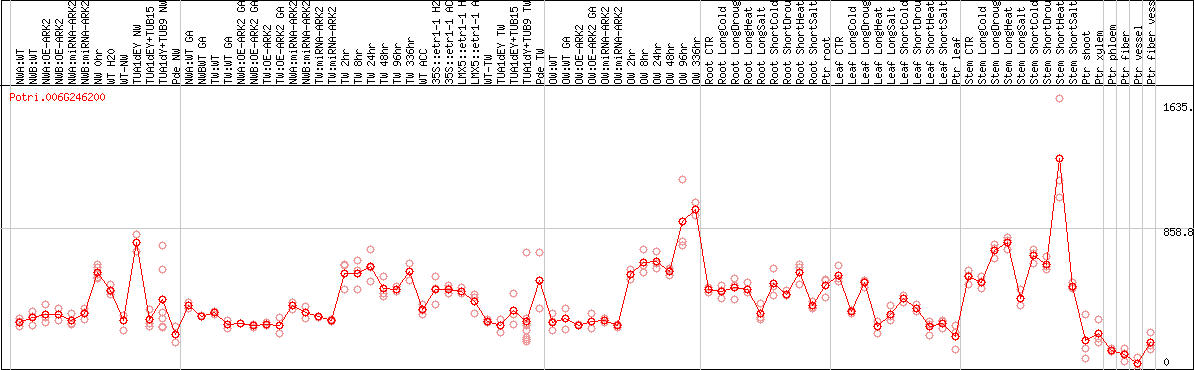
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G246200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
