External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G32340 157 / 2e-47
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G80130 139 / 1e-39
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G20190 135 / 1e-38
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G17940 118 / 4e-32
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G04530 89 / 2e-20
TPR4
tetratricopeptide repeat 4, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G07280 69 / 3e-13
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
AT2G29670 67 / 2e-12
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G47080 56 / 1e-08
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G068200
155 / 8e-46
AT5G20190 167 / 6e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G130100
151 / 3e-44
AT5G20190 180 / 5e-55
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G130300
150 / 4e-44
AT5G20190 166 / 2e-49
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G140500
121 / 5e-33
AT4G17940 196 / 9e-62
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G256900
106 / 2e-26
AT4G17940 125 / 2e-33
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.008G172200
90 / 2e-20
AT1G04530 137 / 1e-36
tetratricopeptide repeat 4, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G065400
82 / 6e-18
AT1G04530 160 / 5e-46
tetratricopeptide repeat 4, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.009G044200
71 / 6e-14
AT2G29670 484 / 5e-166
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G250100
71 / 8e-14
AT2G29670 499 / 5e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002013
166 / 5e-50
AT4G32340 152 / 5e-45
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10030103
153 / 6e-45
AT5G20190 179 / 1e-54
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002166
149 / 1e-43
AT5G20190 192 / 8e-60
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10043266
124 / 3e-34
AT5G20190 162 / 1e-48
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019409
122 / 1e-33
AT5G20190 164 / 1e-49
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10024949
105 / 7e-26
AT4G17940 125 / 8e-33
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10022877
101 / 6e-25
AT4G17940 121 / 5e-32
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014468
97 / 6e-23
AT1G04530 177 / 1e-51
tetratricopeptide repeat 4, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10023721
97 / 8e-23
AT1G04530 171 / 4e-49
tetratricopeptide repeat 4, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002914
94 / 1e-22
AT4G32340 82 / 2e-18
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Representative CDS sequence
>Potri.006G254300.1 pacid=42770127 polypeptide=Potri.006G254300.1.p locus=Potri.006G254300 ID=Potri.006G254300.1.v4.1 annot-version=v4.1
ATGCTTCTTAGAAGTACTTCTACACCGGTATTAAGGACACTTGTCTGCCAGTCATCAACAAGCCGGCCAGTGTCCATGTGCCTTCAAAGAACAGCATCAG
AGGCTGATATTAAGCCACTCTACTTGACCAGAGAGAGAATGTTCTCTAAACGTTCTTTCATGTCACCAGTCCTTAAGGAAAAAGAAGAAATGAGCGTTTG
CATAGAAGCAGTCGAGGAGGAGGAGATGGTCTGTGCAGGGGGTGGAAGTGGTGGTATTTGTGGCAGTGGTGGTGGTGGTGGTGGGAGCTGGGACTCGGGT
CATCAGCCTTATGAAAGTGATCATGAGAGCATGAATCTTTATTACCAGAACATGATCAAGGCCTATCCTGGTGATGCTCTTCTTCTTGCAAACTACGCTA
AATTCTTGAAAGAGGTACGAGGTGATGTTGTGAAGGCAGAGGAGTTCTGTGAGAAAGCAATTCTGGCAAATGGAAGAGATGATGGGAATGTTTTGTCAAT
GTACGGAGATCTAATATGGAATAACCACAAGGACAGTAATCGTGCTCAGGCCTACTTTGATCAAGCTGTCAAGTCCTCTCCCGATGACTGTTATGTGCTT
GCTTCTTATGCTCACTTCCTCTGGGATGCTGGTGAAGAAGATGGTGATGAAGAAGAAGAGACCAAGCAGAATGAAATTCAATGTGATAGTTTGCCAACAT
ATAAACAGATATACAATCTTCCTCAAGGATTCCCTCCTTTGGCTGCTGCTTCTTAA
AA sequence
>Potri.006G254300.1 pacid=42770127 polypeptide=Potri.006G254300.1.p locus=Potri.006G254300 ID=Potri.006G254300.1.v4.1 annot-version=v4.1
MLLRSTSTPVLRTLVCQSSTSRPVSMCLQRTASEADIKPLYLTRERMFSKRSFMSPVLKEKEEMSVCIEAVEEEEMVCAGGGSGGICGSGGGGGGSWDSG
HQPYESDHESMNLYYQNMIKAYPGDALLLANYAKFLKEVRGDVVKAEEFCEKAILANGRDDGNVLSMYGDLIWNNHKDSNRAQAYFDQAVKSSPDDCYVL
ASYAHFLWDAGEEDGDEEEETKQNEIQCDSLPTYKQIYNLPQGFPPLAAAS
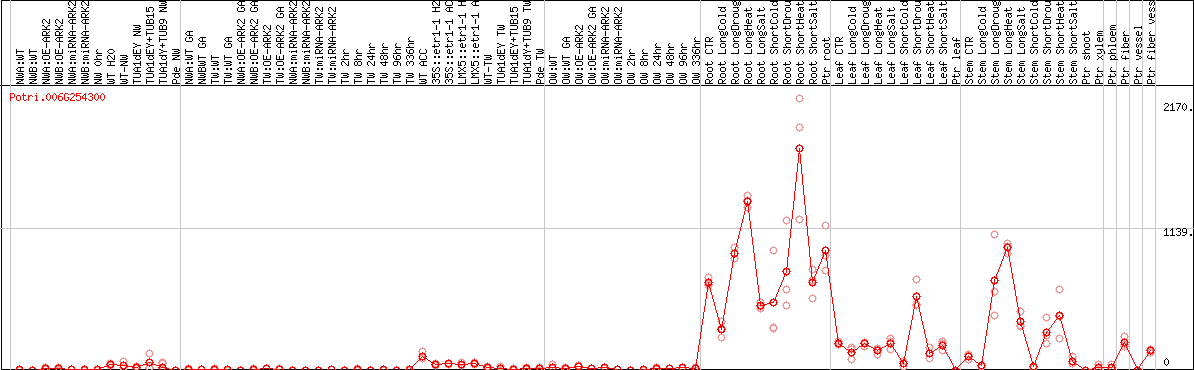
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G254300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.